Impact Factor : 0.548
- NLM ID: 101723284
- OCoLC: 999826537
- LCCN: 2017202541
Aibing Rao
Received: April 06, 2026; Published: April 29, 2026
*Corresponding author: Aibing Rao, Shenzhen Luwei (Biomanifold) Biotechnology Limited, Shenzhen, PR China
DOI: 10.26717/BJSTR.2026.65.010193
Background: Early studies have shown that the kidney organ can also breathe through olfactory receptors, but
the roles that the olfactory system may play are unclear. This study was aimed to analyze the gene expression
profiles in the olfactory system as to various kidney diseases.
Methods: A public data set of eight types of kidney diseases and healthy controls (HC) was used for gene expression
comparison in the olfactory system. Pairwise t-tests were performed and receiver operating characteristic
(ROC) curves were plotted. The summary statistics concerning statistic significance were collected.
Results: Most of the olfactory receptor (OR) genes were down-regulated universally and four genes were down
with statistic significance. Seven of ten adenylate cyclase genes were down- regulated while ADCY7 and ADCY3
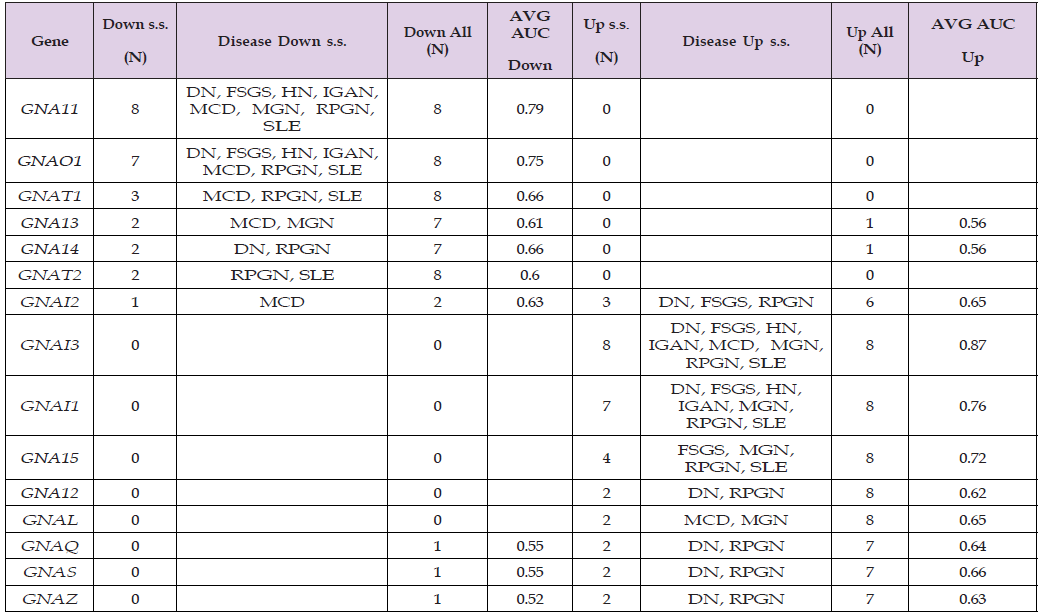
were up-regulated for most disease types. In the selected G-protein α units, GNA11, GNAO1 were down-regulated
while GNAI3 and GNAI1 were up- regulated. Moreover, OR1F2P, OR2W1 and OR7C1 were down mainly in
systemic lupus erythematosus and OR2B2, OR2B6 and OR6A2 were down in rapidly progressive glomerulonephritis.
OR12D3, OR3A2, OR3A3 and OR7C2 were down- regulated for both SLE and RPGN. The linear regression
models gave rise to AUCs in between 0.75 and 1 with average 0.88.
Conclusion: The genes of the olfactory system showed very different expression profiles in kidney diseases
compared to HC. Most genes were uniformly down expressed. Some genes were more related to specific diseases
such as SLE and RPGN. The olfactory system had the expression abnormality in the diseases.
Abbreviations: ROC: Receiver Operating Characteristic; OR: Olfactory Receptor; HC: Healthy Controls; GPCRs: G
Protein-Coupled Receptors; HC: Healthy Controls; SLE: Systemic Lupus Erythematosus; IGAN: IgA Nephropathy;
RPGN: Rapidly Progressive Glomerulonephritis; HN: Hypertensive Nephropathy; MGN: Membranous Glomerulonephritis;
DN: Diabetic Nephropathy; MCD: Minimal Change Disease; FSGS: Focal Segmental Glomerulosclerosis
Olfactory receptors (ORs) are sensory G protein-coupled receptors (GPCRs) and have emerged as key contributors to kidney physiology. ORs have largely been understudied for their roles in kidney [1] and they were expressed in different areas of the nephron, regulating blood pressure, fibrosis, and filtration [2,3]. In the ordorant sensory signaling cascading, first ordorants bind with ORs, and then the trimeric olfactory G protein Golf dissociates into active α and βγ subunits where the α subunit triggers adenylate cyclase activation [3]. The cascading in kidney might be involved with pathogenesis. For example, loss of AC3 in mouse kidney resulted in abnormal glomerular filtration rate and plasma renin [4]. OR51E2, activated by short-chain fatty acids produced by gut microbiota, modulates renin release and blood pressure [5] and the intestinal response to colitis [6]. OR51E1 modulates sex-dependent renin alternation, vascular reactivity and arterial stiffness [7]. OR2Y1 participates kidney glucose handling via SGLT1[8] and contributes to the progression of type 2 diabetes [9]. The expressions of OR10AA1P, OR10A2, OR2Y1, OR1F1, and OR52P1 changed along with the progression of kidney fibrosis [10]. In addition, ectopic ORs have biological effects in cardiovascular cells [2]. In this research, a systemic analysis was performed for the gene expression of selected genes in the olfactory system.
Microarray Data Preparation and Description
A microarray data set, GSE104954 [11], was downloaded from GEO. It included 195 kidney biopsy tissues from ten different kidney diseases and healthy controls (HC). Eight kidney disease groups with at least 13 samples were selected for the analysis: systemic lupus erythematosus (SLE), IgA nephropathy (IGAN), rapidly progressive glomerulonephritis (RPGN), hypertensive nephropathy (HN), membranous glomerulonephritis (MGN), diabetic nephropathy (DN), minimal change disease (MCD), and focal segmental glomerulosclerosis (FSGS). The group sizes range from 13 to 25. The data was normalized by mapping the 25th and the 75th percentiles to 0 and 1 in both sample and gene dimensions. It included 37 OR genes, 10 ADCY genes, and 15 GNA genes listed in Table 1. The validation sets only contained IGAN, HN and HC by combining GSE37460 and GSE93798.
T-tests and Receiver Operating Characteristic Concerning Eight Dis- Eases with HC
For each gene in Table 1, pairwise t-tests were performed for a given disease and HC. The test results included HC mean μ0, disease mean μ1, and p value. A gene is up-regulated if μ0 < μ1 and down-regulated otherwise. It is statistically significant (denoted as s.s.) if p ≤ 0.05. Furthermore, a receiver operating characteristic (ROC) curve was plotted and the area under the curve (AUC) was collected [12] for each disease and HC.
Statistic Summary
Each gene was summarized by first counting the number of ups or downs across the diseases and then only counting for cases with p ≤ 0.05. The average AUC was also collected for up or down groups. Linear Regression Models and Associated Rocs Genes which were s.s. in either direction for at least five disease were selected to build linear regression models and ROC curves were plotted.
Data Analysis and Software
The statistics and the plots were implemented in R scripts developed in house. The student t-test used t.test in R package stats. The ROC used R package ROCR.
T-test and ROC Summary
In Table 2, AUC averages were taken for up and down respectively. Most of OR genes have a ”Down” count of 8 or 7 (Column 4) with OR11A1, OR2J3 and OR2S2 as exceptions. OR10H3, OR7A5, OR1D2, and OR2F1 were s.s. for most diseases (Column 2) and their AUCs were 0.79, 0.75, 0.77 and 0.74. There are 13 genes (from OR10H1 to OR7A10) which were down s.s. mainly for RPGN and SLE. Moreover, OR1F2P, OR2W1 and OR7C1 were down s.s. only for SLE and OR2B2, OR2B6 and OR6A2 were down s.s. only for RPGN. OR12D3, OR3A2, OR3A3 and OR7C2 were down s.s. for both SLE and RPGN. OR11A1 was the only up-regulated OR gene for most diseases but none was s.s.. In Table 3, most genes were universally down, but ADCY7 and ADCY3 were universally up and ADCY8 was in between. ADCY6, ADCYAP1, ADCY1 and ADCY7 were s.s. In Table 4, the top 6 genes were down (Column 4) and the bottom 9 genes were up (Column 8) universally. GNA11, GNAO1, GNAI3 and GNAI1 were s.s. with average AUCs greater than 0.74. GNA12, GNAQ, GNAS and GNAZ were up s.s. while GNA14 was down s.s. only for DN and RPGN. GNAT2 was down-regulated s.s. solely for SLE.
Linear Regression Models and ROC
Based on the above t-test results, OR10H3, OR7A5, OR1D2, and OR2F1 in OR, and ADCY6, ADCYAP1, ADCY1 and ADCY7 in ADCY, and GNA11, GNAO1, GNAI3 and GNAI1 in GNA, were s.s. universally and were selected for linear models. Supplemental Tables 1,2 & 3 in the supplemental list the model results. Based on the linear models, ROCs were plotted and the model performance is listed in Table 4. It showed that AUCs ranged from 0.75 to 1 with average 0.88, FPRs ranged from 0.0% to 24% with average 12%, and TPRs ranged from 60% to 100% with average 80%, indicating that each model separated the disease from HC very well.
Validation
Limited validation due to data availability was performed only for IGAN and HN. Most of the genes showed the same expression status except occasional exceptions which might be due to the data capture variations as shown in Supplemental Tables 2,3 & 4.
Summary
In summary, 4 genes in each group, namely OR10H3, OR7A5, OR1D2, OR2F1; ADCY6, AD- CYAP1, ADCY1, ADCY7; GNA11, GNAO1, GNAI3 and GNAI1 were universally down or up across all kidney diseases, hence they might be related to the common pathogenesis of most kidney diseases. Only 3 of them were up: ADCY7, GNAI3 and GNAI1. For disease specific genes, OR1F2P, OR2W1, and OR7C1 might be related to SLE, while OR2B2, OR2B6 and OR6A2 might be related to RPGN. For disease-pair specific genes, OR12D3, OR3A2, OR3A3, OR7C2 and GNAT2 might be specifically related to SLE and RPGN, while GNA12, GNAQ, GNAS and GNAZ might be specifically related to DN and RPGN.
Limitations
These results need to be further investigated and validated due to the group size. Since there are more than 400 human ORs, a lot of other ORs, ADCYs and GNAs were not considered here due to data availability
The genes in the olfactory system have been understudied especially for kidney diseases but there were some results in general. OR7A5 was shown to be up in glioma and inhibit lipid metabolism and proliferation for glioma cells [13], so down-regulation of OR7A5 in kidney tissue might disturb lipid metabolism. The activation of OR1D2 by specific agonists such as bourgeonal triggered Ca2+ increases in human airway smooth muscle cells via a cAMP-dependent signal transduction cascade and induced the secretion of IL-8 and CSF2 for airway inflammation [14]. Since undecanal is a potent antagonist to OR1D2 [15] and it is widely used in cosmetics and food industry, we now suspect that undecanal might be a risk factor by affecting OR1D2 expression. ADCY7 and ADCY6 were elevated under hypoxia, leading to the elevation of cAMP levels and enhanced PKA activity [16]. In chronic kidney diseases, hypoxia is associated with renal inflammation and fibrosis [17] and hence ADCY7 in the context of kidney pathogenesis should be investigated. GNA11 participates calcium signaling and mutation analysis demonstrated its involvement in hypercalcemia or hypocalcemia [18] and uveal melanoma [19], and GNA11/GNAQ mosaicism was demonstrated to induce hyperactivated calcium signaling [20], hence, GNA11 down-regulation in kidneys might reduce intracellular calcium influx to disturb multiple signaling pathways.
Ethical Approval
Not applicable.
Informed Consent to Participate
Not applicable.
Competing Interests
A. Rao is a co-founder and full-time employee of Shenzhen Luwei (Biomanifold) Biotechnology Limited.
Data Availability Statement
The original data sets GSE104954, GSE37460 and GSE93798 are available via GEO data portal.
Funding Acknowledgement
Research reported in this paper was supported by President Foundation of the Third Affiliated Hospital of Southern Medical University under award number YQ202205.
(Tables 1-4) (Supplemental Text).
Table 1: Lists of OR, ADCY, and GNA genes contained in GSE104954. OR: olfactory receptor; ADCY: adenylate cyclase; GNA: G-protein α unit. Note that ADCYAP1 and ADCYAP1R1 are not ADCY enzymes but included in the ADCY group for convenience.
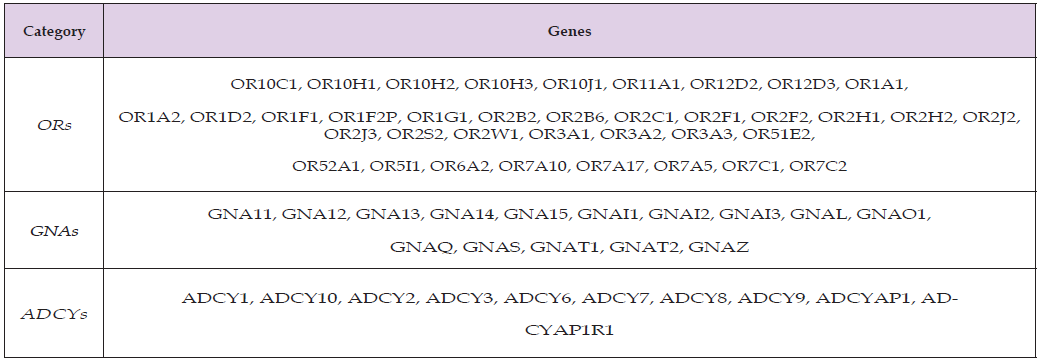
Table 2: T-test and AUC summary for ORs. N: count; s.s.: statistically significant; Up/Down: expression up/down-regulated for a disease type; AVG: average; AUC: area under the curve. Empty cells: not applicable.
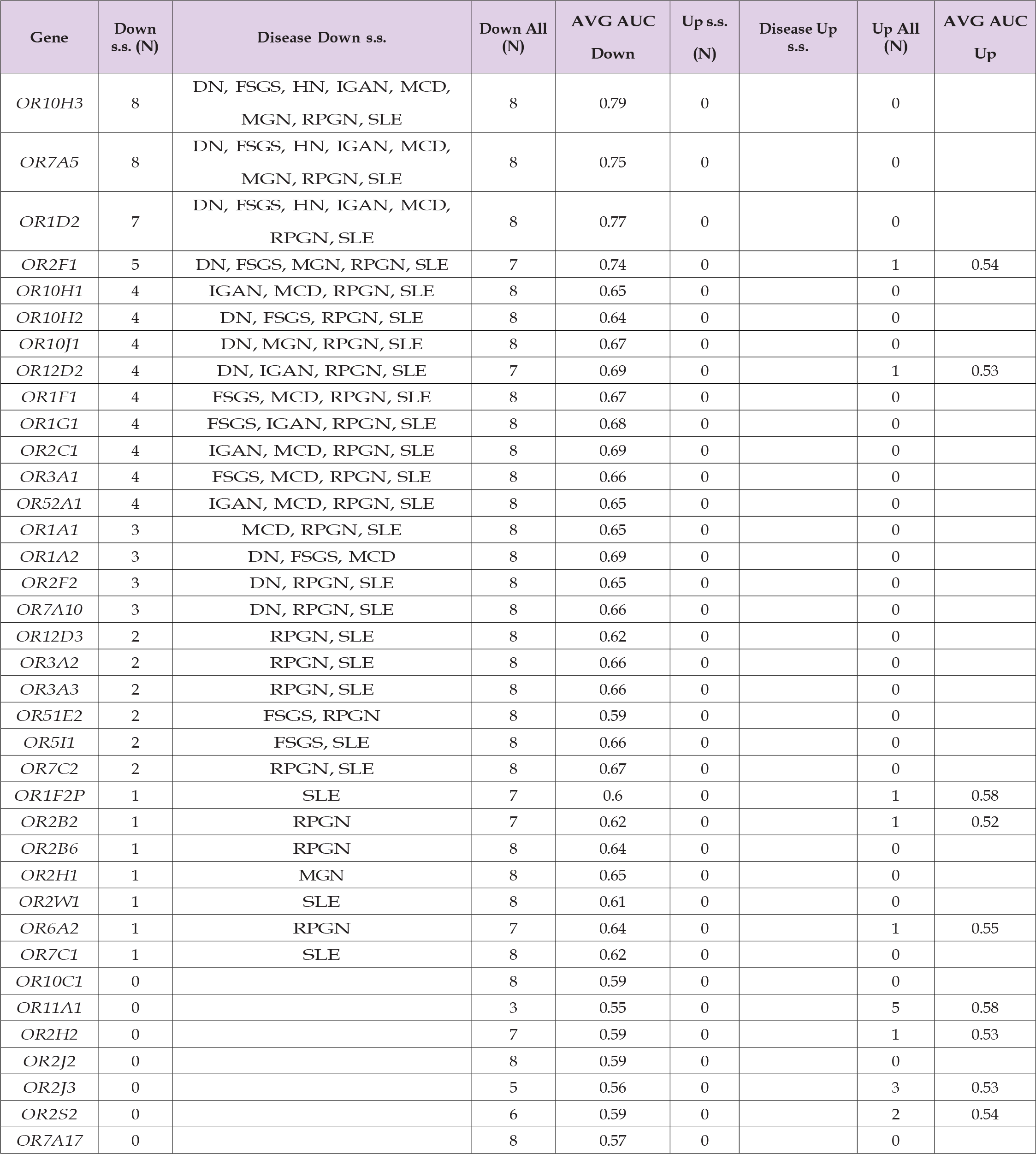
Table 3: T-test and AUC summary for ADCYs. N: count; s.s.: statistically significant; Up/Down: expression up/down-regulated for a disease type; AVG: average; AUC: area under the curve. Empty cells: not applicable.
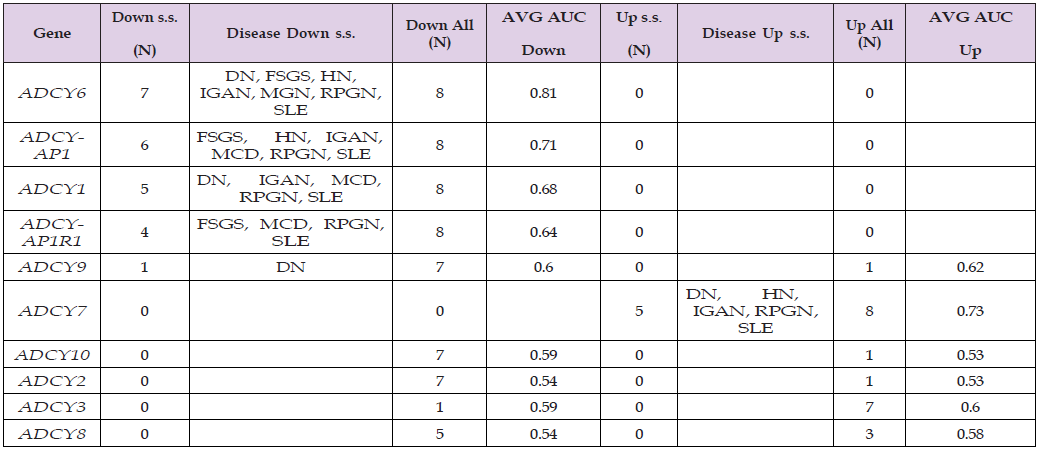
Table 4: T-test and AUC summary for GNAs. N: count; s.s.: statistically significant; Up/Down: expression up/down-regulated for a disease type; AVG: average; AUC: area under the curve. Empty cells: not applicable.