Impact Factor : 0.548
- NLM ID: 101723284
- OCoLC: 999826537
- LCCN: 2017202541
Qiuyan Wang1#, Chengcheng Tang2#, Shulong Li2, Yangyang Zhang2, Junjie Li2, Shimeng Zhang2, Zhiyou Chen2, Zhirong Tan2, Yan Zhuang2 and Yanting Jiang2*
Received: August 07, 2023; Published: September 15, 2023
*Corresponding author: Yanting Jiang, Shenzhen Qianhai Shekou free Trade Zone hospital, Shenzhen, 518067, China
DOI: 10.26717/BJSTR.2023.52.008322
GRIA3 belongs to a class of an alpha-amino-3-hydroxy-5-methyl-4-isoxazole propionate (AMPA)-sensitive glutamate receptor that operates as a ligand-gatedion channel in the central nervous system and has an essential role in excitatory synaptic transmission, localized on the X chromosome, is related to syndromic X-linked intellectual disability 94(MRX94). MRX94, characterized by brain anomalies, namely cerebellar hypoplasia, specific facial features, and intellectual disability, is produced by different mutations in the GRIA3 gene. The exon 5-12 deletion in GRIA3 gene causes MRX94 in a Chinese family, which provides a reliable theoretical basis for prenatal diagnosis and genetic counseling. Amniotic fluid cell genetic testing was performed retrospectively in one fetus with a family history of schizophrenia, and the results of genetic testing were analyzed. The resulting fetus had exon 5-12 deletion in GRIA3 gene, the deletion originating from the maternal grandmother of the fetus. The mother and maternal grandmother of the fetus carried the exon 5-12 deletion in GRIA3 gene. Conclusion genetic diagnosis of a fetus and family with MRX94 caused by exon 5-12 deletion in GRIA3 gene, prenatal genetic testing can avoid the birth of MRX94 fetus, and the occurrence of exon 5-12 deletion in GRIA3 gene causing MRX94 is reported for the first time.
Keywords: Syndromic X-Linked Intellectual Disability 94; GRIA3; Deletion; Exon5-12; Prenatal Diagnosis.
Syndromic X-linked intellectual disability 94 (MRX94) is a rare, inherited, syndrome characterized by moderate mental retardation, with additional features including constitutional weakness, macrocephaly, autism, seizures, and myoclonic seizures [1]. Mutations in GRIA3 have been reported to be associated with MRX94 [2-6]. GRIA3 belongs to a class of an alpha-amino-3-hydroxy-5-methyl-4-isoxazole propionate (AMPA)-sensitive glutamate receptor, localized on the Xq25, is a coding protein with 16 exons, consisting of 2948bp, 894aa. These receptors contain three functional domains–transmembrane, ligand binding, and receptor channel core [7-8]. The genetic variation of AMPAR is associated with different neurological characteristics in humans, such as sleep patterns, migraine, psychoactive drug addiction, intellectual disability and autism [9-16]. In the present family with patient MRX94, in which DNA was examined using gene sequencing to screen for large GRIA3 exon5-12 deletions, and prenatal diagnosis was performed on the fetus, it was found that the proband and this family had the same GRIA3 exon5-12 deletion locus, and that the deletion of the large GRIA3 exon5-12 fragment was previously unreported and is a novel pathogenic GRIA3 mutant.
Materials
A pregnant woman came to hospital for an intrapartum examination at 16+ weeks, the propositus’s maternal uncle, the proband (maternal brother) was mentally retarded and had schizophrenia, brain atrophy, seizures, episodic irritability. Whole exome and epilepsy omnibus genetic testing of the proband revealed a hemizygous exon 5-12 deletion in GRIA3 gene, and the pregnant woman (the proband’s sister) and her mother were heterozygous for exons 5-12 of GRIA3 gene. Whole exome and comprehensive genetic testing of epilepsy for the fetus and other family members should be requested from the family, 4 ml of venous blood (EDTA-K2) was collected, and 10 ml of amniotic fluid was collected and then used for DNA extraction. The detailed pedigree diagram is shown in Figure 1. The father (32 years) and mother (27 years) of the fetus, not recently married, were phenotypically normal. The work in this study was approved by pregnant women and family members and signed informed consent.
Methods
DNA Extraction: Genomic DNA was extracted from the peripheral blood using a blood DNA extraction kit (QIAamp DNA blood Mini Kit; Qiagen, Germany), respectively, according to the Manufactuer’s instructions DNA concentration was measured by nanodrop2000 (Thermo Fisher Scientific, Waltham, MA, USA). The isolated genomic DNA was stored at −20 °C.
Test Method: The whole-exome sequencing and epilepsy panel V2 were used to detect SNV, InDel and exon5-12 deletion of GRIA3 gene in the fetus and its family members. Sequence alignment and variant calling were performed against the reference human genome (UCSC Hg19). Sequencing data analysis was conducted by the Verita Trekker® mutantion site detection system and enliven® mutation site annotation interpretation system Functional prediction of genetic mutations. The analysis filtered out the variants with mutation frequencies greater than 1‰ in the human exon database (ExAC), the 1000 Genomes Project, and the Genome Aggregation Database (gnomAD), and filtered the nonfunctional variation site. The pathogenicity prediction was performed using multiple software packages including SIFT, Polyphen2, and CADD. The potential pathogenic variant was determined along with the related disease database and relevant clinical reports.
Results of Genetic Testing
Whole exome sequencing identified no pathogenic SNV and indel variants associated with the phenotype of case MRX94, according to the genetic mutation type of the proband of this family, a totipotent version of the genetic test for epilepsy was performed. Found to have the same mutation type as the proband, deletion of exon5 -12 in GRIA3. The mutation was inherited from the fetal grandmother. Both the mother and grandmother of the fetus were heterozygous deletion of exon5-12 of GRIA3 gene. The fetus’ father, sister, great aunt, second aunt, and maternal grandmother’s sister were wild type (see Figure 2).
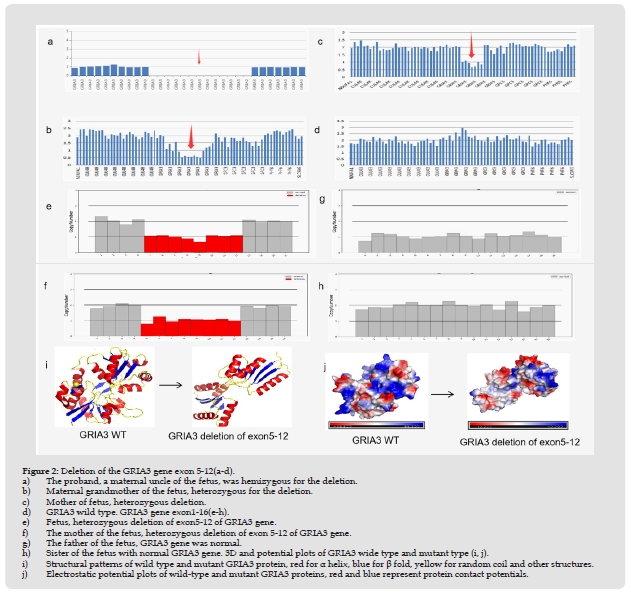
Figure 2 Deletion of the GRIA3 gene exon 5-12(a-d). a) The proband, a maternal uncle of the fetus, was hemizygous for the deletion. b) Maternal grandmother of the fetus, heterozygous for the deletion. c) Mother of fetus, heterozygous deletion. d) GRIA3 wild type. GRIA3 gene exon1-16(e-h). e) Fetus, heterozygous deletion of exon5-12 of GRIA3 gene. f) The mother of the fetus, heterozygous deletion of exon 5-12 of GRIA3 gene. g) The father of the fetus, GRIA3 gene was normal. h) Sister of the fetus with normal GRIA3 gene. 3D and potential plots of GRIA3 wide type and mutant type (i, j). i) Structural patterns of wild type and mutant GRIA3 protein, red for α helix, blue for β fold, yellow for random coil and other structures. j) Electrostatic potential plots of wild-type and mutant GRIA3 proteins, red and blue represent protein contact potentials.
Pathogenic Bioinformatics Analysis
The deletion of exon5-12 of GRIA3 gene was not included in HGMD database, PubMed search for GRIA3 gene exon5-12 deletion pathogenicity has not been reported. Due to the large exon 5-12 deletion in GRIA3 gene, its involved protein functions include transmembrane, ligand binding and receptor pathways (see Figure 2). The altered amino acid charge is reduced and would interfere with ionic interactions with other transmembrane helices, thus interfering with the proper folding of the protein. Regarding the relationship between GRIA3 mutations and MRX94, seven mutations have been identified, as detailed in Table 1.
Notes: TM represents the transmembrane domain; LBD represents the ligand-binding domain; S represents the polypeptide chain that makes up the LBD.
The maternal uncle of the fetus had mental retardation, schizophrenia, macrocephaly, seizures, and occasional mania. Studies have shown that GRIA3 gene mutations are related to the occurrence of nervous system diseases such as intellectual disability, especially in schizophrenia, epilepsy and autism [6]. The clinical manifestations and genetic sequencing results of the proband involved in this family were consistent with the characteristics of MRX94, the mutation was detected in four generations of the family. Therefore, the comprehensive analysis suggested that the deletion of exon5-12 in GRIA3 gene caused MRX94. In 2020, Piard j et al [17] reported that the GRIA3 c.2477g > a variant receptor caused reduced current responses and reduced cell surface expression in response to agonist induction, leading to pathogenicity through a loss of function mechanism. The deletion of exon5-12 in GRIA3 gene in this paper leads to the reduction of translated amino acids, which leads to the reduction of charge expressed on the cell surface. At the same time, the structure of glutamate receptor is also changed accordingly. It has been shown that the severity and phenotype of mental retardation vary greatly among patients due to different types of GRIA3 mutations [1]. In patients with GRIA3 gene mutation, p. Gly630arg and p. Ala653thr mutations lead to severe mental retardation and behavioral changes [1,4]; p. Arg631ser, p. Phe655ser, p. Met706thr, p. Gly833arg, and 0.4-MB DEL were associated with moderate mental retardation and behavioral changes [3,5]. Pathogenic variants in this gene have all been previously reported in neurodevelopmental disorders, mainly in male patients and rarely found in female patients. In 2020, Trivisano [18] reported a case of GRIA3 gene p. Glu787Lys mutation in a male child, which led to mental retardation and epilepsy symptoms. But in 2022, some scholars found that the same mutation site caused intellectual developmental disorder and epileptic encephalopathy in an affected female child by genetic testing [19]. In 2021, Chinese scholars conducted genetic testing on a girl with severe epilepsy and generalized developmental disorder and found p.R660T mutation in GRIA3 gene [20]. It is important for us to understand more about the genetic variation of GRIA3 in women, and not always believe that the mutation of this gene can only cause disease in men, especially for prenatal diagnosis. In summary, we retrospectively analyzed the etiology and prenatal diagnosis of a family with MRX94 caused by GRIA3 gene exon 5-12 deletion. The fetus was heterozygous for the deletion of exon 5-12 of GRIA3, which is pathogenic. This mutation has not been reported before and is a de novo mutation. This finding enriches the mutation spectrum of GRIA3 and provides a theoretical basis for further research on the pathogenesis, genetic counseling, and prenatal diagnosis of MRX94.
We thank colleagues at the Prenatal Diagnosis Center of Hainan Women and Children’s Medical Center for their prenatal services.
The authors declare that they have no competing interests.
Data and materials are available upon request.


