Impact Factor : 0.548
- NLM ID: 101723284
- OCoLC: 999826537
- LCCN: 2017202541
NAKAYAMA Tomohiro1*, ETO Kaoru2, Mitsuyo Machida3, Yotaro Takagaki3, TACHIKAWA Emiko2, SHIRATO Yuri2 and NAKANO Kazutoshi2,4
Received: March 16, 2023; Published: April 03, 2023
*Corresponding author: NAKAYAMA Tomohiro, Department of Pediatrics, Ibaraki Prefectural University of Health Sciences, 4733 Ami, Ami-town, Inashiki-gun, Ibaraki, 300-0331, Japan
DOI: 10.26717/BJSTR.2023.49.007818
Nakano first discovered the “Mitochondrial cells” (Mito Cells), which can survive for long term despite lacking a nucleus. However, there was no clear evidence to indicate whether MitoCells were alive. To resolve this question, we constructed a GFP-actin induced Mito Cell (GFP-actin Mito Cell). Herein, we report the molecular analysis of both MitoCells and GFP-actin MitoCells. Our results suggest that MitoCells have the minimum amounts of proteins and DNA needed to sustain life.
Keywords: “Mitochondrial Cell”; Nuclear DNA; Mitochondrial DNA
Some years ago, human cell lines without mitochondrial DNA (mtDNA), designated as Rho0 cells, were isolated [1]. After this achievement, cell lines obtained by fusing Rho0 cells with enucleated cells from patients with a variety of mitochondrial diseases have contributed to the understanding of several mitochondrial disorders [2]. Additionally, Nakano used cybrids, obtained by fusing mitochondrial-less HeLa cells and platelets from patients with Leigh syndrome-a subtype of mitochondrial encephalopathy-to isolate stable cell lines designated as Mito Cells [3]. Mito Cells lack nuclei but maintain mitochondrial activity. Moreover, MitoCells grow continuously in culture, despite the absence of nuclei [4] and TCA cycle enzymes [5]. We analyzed MitoCells at the molecular level and found that they had cytoskeletal proteins encoded by nuclear DNA (nDNA) and mitochondrial respiratory enzymes encoded by both nDNA and mtDNA. Moreover, we successfully constructed a GFP-actin Mito Cell line, which indicated that MitoCells were alive. These results suggest that MitoCells possess the ability of protein synthesis and DNA components needed to sustain life. Herein, we report the results from Mito Cell molecular analysis.
Materials
MitoCells were cultured in Roswell Park Memorial Institute medium (RPMI 1640; GIBCO Invitrogen; Carlsbad, CA, USA) supplemented with 10% fetal calf serum, 50 U/mL penicillin, and 50 μg/mL streptomycin at 37oC under a humidified gas mixture containing 5% CO2, as previously described [3]. Anti-tubulin monoclonal and anti-calreticulin polyclonal antibodies were purchased from Neo Markers (Fremont, CA, USA) and Affinity Bioreagents (Golden, CO, USA), respectively. Monoclonal antibodies against complex IV subunits I, II, III, IV, and VI, SURF-1, and phalloidin (anti-actin) were purchased from Molecular Probes (Eugene, OR, USA). As the secondary antibody to be used after each primary antibody, we purchased the Alexa 594-conjugated anti-rabbit IgG from Molecular Probes. The Vectashield (Vector Laboratories; Burlingame, CA, USA) mounting medium was used before cover slipping cell samples. For constructing GFP-actin Mito Cells, we used an EGFP-actin plasmid (Clontech Laboratories, Inc.; Mountain View, CA, USA), Lipofectamine 2000, and Plus Reagent (both from Invitrogen). For live cell imaging, ER-Tracker Green and MitoTracker Red CMXRos (Molecular Probes) were used. For sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDSPAGE) staining, we utilized SYPRO Ruby Protein Gel Stain (Molecular Probes).
Immunocytochemistry Assays
Cells on glass slides were fixed with formalin for 5 min at 37 ℃ by directly adding a concentrated solution to the culture medium, to a final concentration of 4%. After washing twice with Hank’s balanced salt solution (HBSS), samples were permeabilized with 0.2% Triton X-100 in HBSS at room temperature for 5 min, washed again twice with HBSS, and incubated with HBSS supplemented with 2% goat serum and 0.2% Triton X-100 at 4 ºC overnight. Cell samples were then incubated with individual primary antibodies (anti-tubulin, 1:500; anti-calreticulin, 1:800; complex subunits, 1:500; and phalloidin, 1:500) in HBSS containing 2% goat serum and 0.2% Triton X-100 for 2 h at room temperature. After washing the cells with HBSS, they were incubated with Alexa-594 conjugated anti-rabbit IgG for 1 h at 4 ºC and washed three times with HBSS. Then, samples on glass slides were mounted in Vectashield and covered with coverslips.
PCR
To amplify ß-actin, primer pairs were designed according to the gene’s cDNA sequence; the forward primer included nucleotides 558-577, whereas the reverse primer included nucleotides 960-941.
Similarly, to amplify mtDNA coding Cambridge Reference Sequence stretches 3010-3420, 8191-8364, and 8646-9199, primers pairs were designed; forward primers corresponded to sequence numbers 3010-3031, 8191-8210, and 8646-8665, whereas reverse primers included nucleotides 3420-3400, 8364-8345, and 9199-9180. Amplification by PCR was performed as follows: for mtDNA 8191- 8364, 35 cycles including nucleic acids denaturation at 94 oC for 1 min, primer annealing at 45 oC for 2 min, and extension at 72 oC for 2 min. All other fragments were amplified according to the following protocol: 35 cycles of denaturation at 94 oC for 30 s, primer annealing at 55 OC for 30s, and extension at 72 oC for 30s.
GFP-Actin Transfected MitoCells
To construct GFP-actin MitoCells, 1.0 μg of pEGFP-actin was mixed with 1.0 μL of Plus Reagent, followed by 3 μL of Lipofectamine 2000; the mix was incubated for 30 min at room temperature, and directly added to 500 μL of a MitoCell culture.
Time-Lapse Imaging of MitoCells
MitoCells and GFP-actin MitoCells were incubated for 8 weeks. In the meantime, photographs of the culture dishes were acquired every 1 to 2 weeks using a 4× lens. Same dish photographs were acquired in about the same position each time.
Live Cell Imaging of MitoCells
Mito Tracker Red CMXRos was directly added to GFP-actin MitoCells at a final concentration of 50nM. Besides, ER Tracker Green and Mito Tracker Red CMXRos were both directly added to Mito Cells at a final concentration of 100 nM and 50nM, respectively. Cells were incubated for 5 min in an incubator at 37 ℃ and observed under the microscope.
SDS-PAGE
Mixedells and GFP-actin MitoCells were centrifuged at 12,000 rpm for 30 min. Pellets were collected, mixed with SDS, bromophenol blue, and 2-mercaptoethanol, and then boiled for 5 min. The samples were directly added into 7% polyacrylamide gel wells, and the electrophoresis was run at 15 mA for about 1 h. Finally, the gel was stained with SYPRO Ruby Protein Gel Stain. Whole HeLa cells and Hela cell culture medium supernatant were used as controls.
Immunocytochemistry
Positive staining was observed for tubulin, actin, calreticulin, complex IV subunits I, II, III, IV, and VI, and SURF-1 (Figure 1).
PCR
All ß-actin and mtDNA fragments were successfully amplified (Figure 2). The results are summarized in Table 1.
Time-Lapse Images of MitoCells
Time-lapse images of GFP-actin Mito Cell cultures showed a gradual increase in cell number (Figure 3). Therefore, Mito Cells were not only able to survive after 2 months but also proliferated in the dish (data not shown).
Live Cell Imaging of GFP-actin MitoCells and MitoCells
Live cell images of GFP-actin MitoCells are shown in Figure 4. Mitochondria appear in red and GFP-actin in green. Given that MitoTracker Red CMXRos accumulation depends on mitochondrial membrane potentials [6], the results suggest that, besides being able to incorporate GFP-actin, Mito Cells membrane potentials are still functional. In Figure 5, mitochondria are shown in red and the endoplasmic reticulum in green. Noteworthy, the main structure of the endoplasmic reticulum appears scattered throughout the cytosol in Mito Cells.
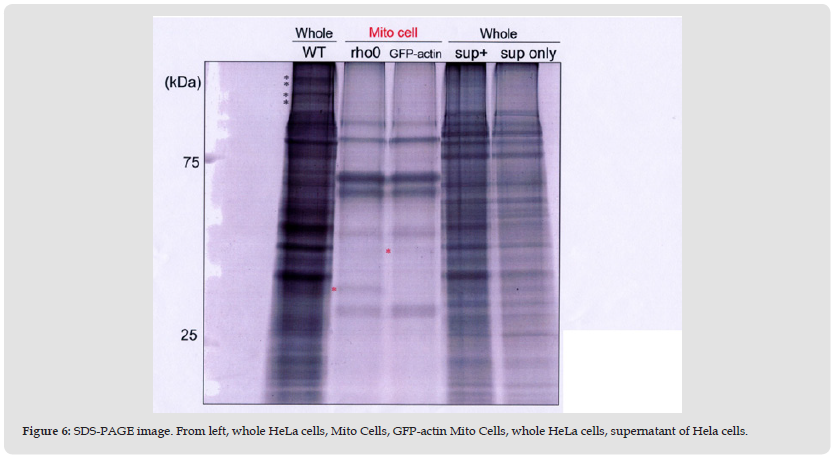
Figure 6 SDS-PAGE image. From left, whole HeLa cells, Mito Cells, GFP-actin Mito Cells, whole HeLa cells, supernatant of Hela cells.
SDS-PAGE
As shown in Figure 6, the protein patterns of HeLa cells (used as control), Mito Cells, and GFP-actin MitoCells, were comparatively different. For Mito Cells, only 6 major bands were recognized, and they were not consistent with those from control cells. Among Mito Cell and GFP-actin Mito Cell profiles, only two protein bands were different, most probably due to the differences between actin and GFP-tagged actin (Figure 6*).
Mito Cells Have Proteins Encoded by Both Nuclear and mtDNA
The results of the immunocytochemistry and PCR analyses indicated that MitoCells have cytoskeletal and endoplasmic proteins encoded by nDNA as well as mitochondrial respiratory enzyme proteins encoded by both nDNA and mtDNA (Table 1).
MitoCells are Alive and Sustain Life
The fact that a GFP-actin Mito Cell line could be successfully established indicated that MitoCells had components of the protein synthesis apparatus. Moreover, the introduction of extra DNA did not modify cell survival or mitochondrial membrane potentials (Figures 3 & 4, respectively). It is well-known that the rough endoplasmic reticulum plays a role in protein synthesis in mammalian cells; interestingly, endoplasmic reticulum proteins were present in MitoCells, but the corresponding membrane components were lost (Figure 5). Consequently, Mito Cell structure somehow resembled that of prokaryotic cells. Noteworthy, the time-lapse study showed that MitoCells were still alive after 2 months. These data suggest that Mito Cells lack nuclei but possess enough components of the protein synthesis apparatus to sustain life.
The Protein Profile of MitoCells was Different from that of HeLa Cells
An SDS-PAGE analysis showed that MitoCells and HeLa cells (which originated Mito Cells) had different protein profiles. The new profile must be due to a massive loss of proteins while MitoCells were being formed from cybrid cells. Either of the following hypotheses could explain the protein change observed in Mito Cells:
1. There was an active change to a prokaryotic cell-like structure as the result of an evolutionary throwback or
2. There was a passive mechanism of loss of those proteins that were not indispensable to sustain life.
The fact that there are only a few protein bands in Mito Cell profiles suggests that MitoCells have the minimum number of proteins needed for sustaining life. Therefore, by studying such a protein profile, we might be able to understand the minimum protein requirements to sustain life.
We have discovered a remarkable “mitochondrial cell” that lacks nuclei but survives for a long time. This cell has both nDNA and mtDNA, and its protein profile is significantly different from that of a human cell. There might be a possibility to understand the minimum protein content needed to sustain life by further studying this cell.
We thank Professor Shigeo Ota, Professor Ikuroh Ohsawa (Dept. of Biochemistry and Cell Biology, Institute of Gerontology, Nippon Medical School) for their advice. This work was supported in part by T. Satake scholarship foundation and in part by grant #14-Syouni-006, Clinical Research for Evidence Based Medicine, from the Ministry of Health, Labor and Welfare, Japan. This work was also supported by Grants-in-Aid of the Research on Intractable Diseases (Mitochondrial Disorder) from the Ministry of Health, Labour and Welfare of Japan.
None.

