Impact Factor : 0.548
- NLM ID: 101723284
- OCoLC: 999826537
- LCCN: 2017202541
Manosha Lakmali Perera1* and Irosha Rukmali Perera2
Received: December 04, 2022; Published: December 13, 2022
*Corresponding author: Manosha Perera, School of Dentistry and Oral Health, Griffith University, Queensland, Australia (Alumni) and 14/2, 3rd Lane, Kadalana, Moratuwa, Sri Lanka
DOI: 10.26717/BJSTR.2022.47.007527
There are trillions of bacterial cells co- habit with human cells. These procaryotes use efflux pumps and enzymes to prevent cellular intoxication of ions and compounds respectively. There is promising evidence on the role of the gut microbiome and its enzymes in metabolizing xenobiotics. The genetic potential of oral bacteria in drug and xenobiotic metabolism is yet to be unveiled. This study aimed to characterize the bacteriome associated with oral fibroepithelial polyps (FEP) and to predict the genetic potential. A representative sub-sample of 22 clinically diagnosed oral FEP (the control group) was selected from a main case-control study. Amplification of nucleotides of extracted DNA from frozen tissues was performed for the V1 to V3 region and sequencing of the amplicon with Illumina’s 2X 300–bp chemistry. Classification of high-quality nonchimeric merged reads was done to the species level with a prioritized BLASTN-based algorithm. Downstream compositional analysis was performed with QIIME (Quantitative Insights into Microbial Ecology).
Functional prediction of bacteriome was obtained by PICRUSt (Phylogenetic Investigation of Communities by Reconstruction of Unobserved States). Rothia mucilaginosa, Streptococcus mitis, Gamella haemolysans, Streptococcus sp. oral taxon 431, and Rothia dentocariosa accounted for the top five taxa among 810 bacterial species according to the percentage of average relative abundance. Rothia mucilaginosa was elevated statistically significantly (p < 0.05). The genetic potential of xenobiotics and drug metabolism catalyzed by the P450 enzymes was observed for the first time as an attribute of bacteriome associated with oral FEP tissues dominated by R. mucilaginosa. This finding needs further investigation.
Keywords: Xenobiotic; Drug; Oral Fibro Epithelial Polyps; Rothia Mucilaginosa; Bacteriome
Abbreviations: NGS: Next-Generation Sequencing Technologies; FEP: Fibroepithelial Polyps; OSCC: Oral Squamous Cell Carcinoma ; OMF: Oral and Maxillo- Facial; LDA: Linear Discriminant Analysis
Microbial genome surpases human genome more than 100 folds [1,2]. Hence, the human body hosts trillions of microorganisms as a “super habitat” or “incredible niche” according to human microbiome studies, that revolutionized the landscape of personalized medicine by probiotic microbial interventions [3]. The oral cavity contains the second most diverse meta community of microbes no less than 700 bacterial species in humans [1,4] and the entryway of the human body, which connects essentially with systems namely the respiratory tract, gastrointestinal tract, and sinuses. When considering the anatomy of the oral cavity, it divides into several components: soft- tissue (gingival sulcus, attached gingiva, tongue, cheek, lip, and soft palate), hard tissue (hard palate and teeth), and saliva [1,5]. From a microbial ecology point of view, each component can be considered a well-defined niche or microhabitat. Numerous factors, such as attachment ligands, nutrient supply, pH, availability of oxygen, bacteriocins, toxic compounds, and immune elements govern the proportions and functions of inhabitants [6,7].
Thus, each microhabitat provides a setting to establish homeostasis with delicate equilibrium in normal physiology [8], stimulating the innate immune mechanisms of the host [9]. Human exposes to environmental pollutants due to the excessive use of agrochemicals [10] that accumulate in the last trophic level of humans via food chains and food webs, in addition to the usage of prescribed drugs or pharmaceuticals in disease conditions. Hence, endogenous microbiota struggle for cellular survival against xenobiotics of synthetic origin and heavy metal ions [11]. Evolutionary, discrete bacterial genetic systems have developed to synthesize a plethora of enzymes [10] and amazing efflux pumps [11,12] to metabolize xenobiotic compounds and extrude poisonous ions respectively. Chronic irritation to the oral mucosa due to oral risk habits may cause inflammation. Alarmingly, these risk habits comprising smoking [13] heavy alcohol consumption, as well as betel quid chewing with smokeless tobacco, areca nut, and slaked lime, are well documented as major risk factors of oral carcinogenesis [14].
Mucosal benign oral lesions also account for these oral risk habits [15]. Hyperplastic parakeratinized stratified squamous epithelium with arcading pattern and mixed inflammatory cell infiltrate lymphocytes predominantly and plasma cells usually describe anatomical pathology of oral fibroepithelial polyps (FEP) [16]. In the 4th industrial revolution era, next-generation sequencing technologies (NGS) have provided artificial intelligence of bioinformatics algorithms viewing microbial communities in unprecedented breadth and depth and predicting their genetic potential [8]. In metagenomic studies prokaryotic 16S rRNA gene manipulate as the house keeping or universal gene [15,17]. This supergene consisted of constant regions for amplification, and variable regions for identification by nucleotide sequencing [18]. One of the bacteria, Rothia mucilaginosa is a member of the commensal oral flora and respiratory tract [19]. The objective of this study was to find out the compositional profile and predict the genetic potential of bacteriome in a group of Sri Lankan male oral FEP patients.
Ethical Approval
Ethical approval for present study was obtained from the Faculty Research Committee, Faculty of Dental Sciences, University of Peradeniya, Sri Lanka (FRC/ FDS/UOP/E/2014/32) and Griffith University Human Research Ethics Committee, Australia (DOH/18/14/ HREC).
Study Design, Sample Size Calculation, Setting, and Subjects
This epidemiology study was based on a multicenter field study [15]. Hence, a representative sub-sample of 22 clinically diagnosed FEP (the control group) was selected from the main unmatched case-control study of 134 histologically confirmed oral squamous cell carcinoma (OSCC) cases and clinically diagnosed 134 benign mucosal lesion (BML) controls. The sample size calculation for the unmatched case-control study was based on Kelsey, et al. (Appendix 3.1). Selected Oral and Maxillo- Facial (OMF) Units across Sri Lanka were visited representing six provinces for profiling of bacteriome comprised Sinhala, ≥40-year-old with a clinical diagnosis of FEP also involving the buccal mucosa or tongue. Written informed consent was obtained from each participant [7,15].
Risk Habit Profile
Pre tested interviewer administered questioner was used to collect the details of oral risk habits and other sociodemographic information of study subjects as described previously [8,15].
Tissue Sampling and DNA Extraction
Deep tissue pieces (~3 mm3) from excisional FEP biopsies were taken. Subsequently, DNA extraction from frozen samples (=800C) was performed according to (solid tissue protocol) as described previously [8,15].
Amplicon Library Preparation and Nucleotide Sequencing
Prokaryotic primers 27FYM (5′-AGAGTTTGATCMTGGCTCAG- 3′) and 519R (5′-GW ATTACCGCGGCKGCTG3′), were used to amplify V1-V3 region of the 16S rRNA gene. Amplicon library preparation, indexing, and sequencing were performed at the Australian Centre for Ecogenomics (University of Queensland, Australia. Specifically, 2 × 300–bp chemistry was used on a MiSeq platform (Illumina) for sequencing [15].
Data Processing
Preprocessing of data was performed as detailed previously for prokaryotic raw sequencing reads [7,20]. The high-quality non-chimeric merged reads were classified upto species level using BLASTN searched against 4 databases of 16S rRNA prokaryotic gene reference sequences at alignment coverage and percentage identity ≥ 98% as described previously [7,21].
Compositional Analysis and Functional Prediction
Compositional analysis and functional prediction were performed as previously described [15,20]. The microbial metagenome was imputed from the 16S rRNA data with PICRUSt (Phylogenetic Investigation of Communities by Reconstruction of Unobserved States) based on KEGG Orthology genes and pathways [15].
Statistical Analysis
Fisher’s exact test to compare percentages of relative abundances (cell counts < 5) of R. mucilaginosa with R. dontocariosa and linear discriminant analysis (LDA) effect size analysis were used to prediction of fnctional pathway levels of oral bacteria sought with between the cases and controls as described previously [15,22].
The result is presented in 3 parts to provide scientific evidence to achieve the objective.
Oral Risk Habits Profile of Study Group
According to Table 1, of oral FEP subjects, 45.4 %, 22,7% and 13.6% were current betel quid chewers (on daily basis), current smokers (on daily basis) and current drinkers (on weekly basis) respectively. However, 4% was never betel quid chewers, 7% was never smokers and 13.6% was never drinkers.
The Composition of Oral Microbiome in 22 FEP Subjects
A total of 810 bacterial species were identified in FEP subjects, with inter-individual variations ranging from 23 to 229. Of them, Rothia mucilaginosa dominated, followed by Streptococcus mitis, Gamella haemolysans, Streptococcus sp. oral taxon 431, and Rothia dentocariosa present in more than 50% of individuals depending on their relative abundance.
Top 5 Bacterial Species Detected in FEP Subjects
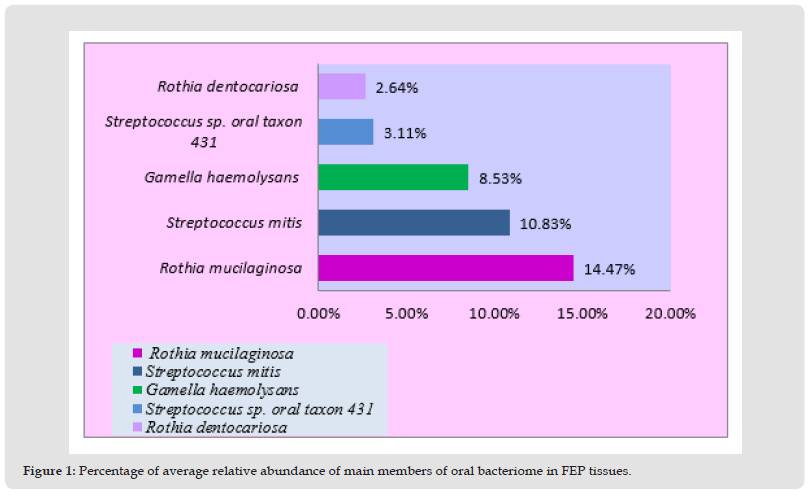
Rothia mucilaginosa is elevated by sevenfold compared with another member of the same genus name. Rothia dentocariosa in oral FEP tissues (Figure 1). Streptococcus mitis was the second most abundant oral taxon, increased by nearly four-fold when it comes to the relative abundance of Streptococcus sp. oral taxon 431 . Moreover, Gamella haemolysans was the 3rd most abundant oral bacterial species found in FEP tissues, followed by Streptococcus sp. oral taxon 431 andRothia dentocariosa 4th and 5th respectively (Table 2).
Table 2: Comparison of percentage of relative abundance of dominated R. mucilaginosa with other four over presented oral bacterial species in FEP tissues.
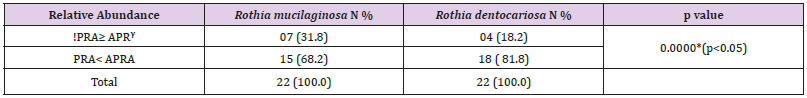
Note: !Percentage of Relative Abundance (PRA)
ᵞPercentage of Average Relative Abundance (PARA)
*Fisher’s exact test to compare groups (cell counts <5), p= 0.000*, (p< 0.05)
Figure 1 Percentage of average relative abundance of main members of oral bacteriome in FEP tissues.
Functional Prediction of Pathway Levels of Oral Bacteria in FEP
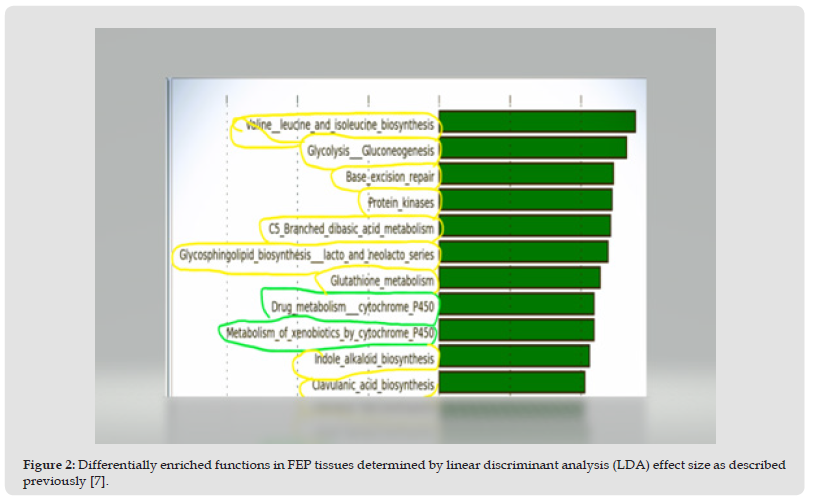
According to Figure 2, biosynthesis of amino acids, synthesis of cellular constituents, gene repair as well as a breakdown of carbohydrate substrates by glycolysis (circled in yellow color) were identified in the pathway-level analysis. Most importantly, the metabolism of xenobiotics and drugs was predicted (circled in green) on the functional metagenome associated with the FEP microenvironment.
Figure 2 Differentially enriched functions in FEP tissues determined by linear discriminant analysis (LDA) effect size as described previously [7].
The first and foremost observation of oral microbes way back in the 1600s. Antonie van Leeuwenhoek observed tiny moving objects or “animalcules” from dental plaque with a single lens microscope. Subsequently, this Dutch merchant was honored as the father of microbiology for his pioneering work as a microscopist or an oral microbiologist. From then onwards oral microbiology evolved from conventional culture techniques to adventurous modern metagenomic revelations. Unveiling the meta genome or total prokaryotic genome in a clinical or environmental sample performs efficiently by nucleotide revealing of next-generation sequencing (NGS), enabling snapshot viewing of microbiota in their natural habitat [1]. To the authors' knowledge, this is the first study to uncover the bacteriome of oral FEP, one of the common types of benign oral mucosal lesions always associated with inflammation.
The published information is not available on the same topic to compare the results of the present study. As a result, transdisciplinary findings are used to defend the unique discoveries of the present study. In the present metagenomic study, there was a statistically significant association (p< 0.05) between the elevation of R. mucilaginosa more than the other member of the same genus, R. dentocariosa in FEP tissues. This bacterium is a catalase-variable, oxidase-negative, and Gram-positive coccus that can appear in pairs, tetrads, or irregular clusters [19]. Hence, Rothia mucilaginosa was formerly known as Staphylococcus salivarius, Micrococcus mucilaginous, and Stomatococcus mucilaginous [19]. This commensal in the oral cavity and throat is capable of causing opportunistic infections usually in immunocompromised. Alarmingly, this opportunistic pathogen is also emerging as a nosocomial pathogen commonly associated with prosthetic valve endocarditis and prosthetic device infections.
FEP belongs to the spectrum of benign mucosal lesions of the oral cavity [15,23]. The vast majority of this study cohort was with oral risk habits which may cause hyperplastic oral FEP due to irritation to oral mucosa initiated by inflammogenic, immunogenic compounds in masticatory substances. Moreover, these chemicals may induce environmental stress [24] on oral microbial physiology. The destruction of susceptible bacteria by bactericidal compounds in masticatory ingredients might be possible. Thus in-vitro experiments have shown anti microbial activities of Areca catechu [25], Piper beetle [26], and Nicotiana tabacum [27]. Anti-bacterial phyto chemicals in masticatory substances may modify the composition and function of micro flora in a particular microhabitat of the oral cavity. The soaring of R. mucilaginosa might be due to the lack of competition for the host’s resources by competing neighbors [1,7] and possible immune evasion by this opportunistic pathogen. These microbial shifts and their metabolites in hosts may act as environmental risk modifiers or co-factors in the initiation and/or progression of oral carcinogenesis as per molecular epidemiology of oral cancer [28].
Oral bacteria equipped with the genetic potential of metabolizing xenobiotics and drugs catalyzed by Cytochrome P450. The finding of drug metabolism in this study is consistent with the previous finding of the ability of drug metabolism of the oral microbiome associated with healthy Saudi Arabian individuals [20]. The catalytic role of bacterial P450 enzymes is already known [29]. The gut microbiome engaged in xenobiotic metabolism was identified [10]. Pathway-level glycolysis, glutathione metabolism, and C5_branched_dibasic_acid metabolism were also observed in this study. More or less similar findings were obtained on oral microbiomes associated with healthy controls and oral cancer cases [15,20,25]. These heterotrophic microbes are essentially dependent on the [30] host carbonic substrates for their energy production.
In the present study, the genetic potential of oral bacteria extended up to xenobiotic metabolism for the first time. This prediction is limited to the oral bacteria associated with a benign inflammatory condition, common among masticatory substance abusers. The elevation of R. mucilaginosa may indicate a progressing dysbiotic condition due to overburdened immune functions at the FEP tumor microenvironment. Removal of noxious chemical stimuli by innate immune mechanisms is essential in inflammatory conditions without letting them progress into a premalignant or malignant condition. Hence, environmental stress factors may govern the resilience development and the cellular survival of uni-cellulars. A snapshot view of the genetic potential of unicellulars against cellular intoxication in the present study is noteworthy. The smaller sample size was a limitation of this study.
This preliminary study insights the genetic potential of bacteriome in a group of Sri Lankan oral fibro epithelial polyp male patients dominated by Rothia mucilaginosa. It needs to confirm this finding using metatranscriptomics in a large study cohort. Further cross-sectional studies with large sample sizes are recommended to cognate effectors of oral bacteria in healthy individuals.
Future work on identifying oral microbes’ fortune with xenobiotic- metabolizing enzymes and a comprehensive understanding of their genetic systems by whole genome sequencing, identifying mutations by plasmid cloning, and identifying mutations by transposon- directed insertion site sequencing, seems multidisciplinary yet fortae of biotechnology.
M. Perera, contributed to conception, design, data acquisition and interpretation, scientifically improved the manuscript; I. Perera, contributed to design, data acquisition,interpretation and statistical analysis.
First and foremost, I would like to sincerely thank Prof. Newell W.Johnson for allowing me to work under him as the Principal Supervisor for my Ph.D., establishing metagenomic studies at the School of Dentistry and Oral Health, Gold Coast Campus, Griffith University, Australia. I express my gratitude to Prof. G.C. Ulett for his overall supervision in the DNA extraction phase of my Ph.D. under the guidance of Dr. D. Ipe in his lab. Special thanks go to Dr. D.J. Speicher for his contribution to dispatching extracted DNA samples for amplicon sequencing. I am indebted to Dr. T.Chen for spending long hours of metagenomic data processing. Above all, I am overwhelmed by my deepest appreciation to Prof. Al-hebshi for his valuable guidance from nucleotide sequencing to three papers based on my Ph.D. thesis.
The authors declare no potential conflicts of interest with respect to the authorship and/or publication of this article.


