Abstract
Background: Breast cancer is one of the most common malignancies among women in the world. Defects in DNA repair or the response to DNA damage (e.g. homologous recombination repair system) can result in increased rates of genetic mutations, in turn, leading to the development of breast cancer. Endonuclease/exonuclease/phosphatase family domain containing 1 (EEPD1) and crossover junction endonuclease MUS81 (MUS81) are two crucial enzymes in homologous recombination repair.
Methods: This study aimed to evaluate the expression of these genes in BC samples. This case-control study was included 40 paired tumor and adjacent non-tumor tissues from female BC patients. Three cancer cell lines (MCF-7, MDA-MB-231, and SKBR3) were also used. The total RNA was isolated and the expression levels of EEPD1 and MUS81 were assessed using quantitative real-time PCR (qRT-PCR). Potential correlations between gene expression and clinicopathological characteristics were also analyzed. The survival analysis of the candidate genes was applied using in silico data.
Results: The expression of EEPD1 was significantly decreased in BC tumor tissues compared to the adjacent non-tumor tissues (P-value < 0.05). The expression of MUS81 was higher in Her-2 positive BC patients than Her-2 negative individuals. Low expression of EEPD1 was detected in estrogen receptor (ER)-positive cells (P-value < 0.05). Using in silico data, we showed that low expression of EEPD1 and MUS81 was associated with poor survival of BC patients.
Conclusion: This study showed that EEPD1 was significantly downregulated in BC, particularly in ER-positive tumors.
Keywords: Breast Cancer; EEPD1; Gene Expression; MUS81; Prognosis; In Silico Analysis
Abbreviations: BC: Breast cancer; HR: Homologous Recombination; DSB: DNA double- Strand Breaks; EME1: Essential Meiotic Structure-Specific Endonuclease-1; TNM: Tumor–Node–Metastasis; HER‐2: Human Epidermal Growth Factor Receptor 2; ER: Estrogen Receptor; PR: Progesterone Receptor
Introduction
Breast cancer is one of the chief causes of death and also the most common type of cancer among women worldwide [1]. During the last 20 years, the incidence of BC has increased in developed countries [2,3]. BC neoplasms are intimately related to DNA damage repair defects or defects in cell-cycle checkpoints which allow mutated DNA to remain unrepaired in doughter cells [4,5]. Despite advances in diagnosis and also treatment, the survival rate of BC patients remains low [6,7]. Thus, there is an urgent need for introducing the gene expression profiles to identify possible treatment targets or even prognosis markers in BC. It has been identified that deficient DNA repair seems to be a predisposing factor in both familial and sporadic BC cases [8]. Homologous recombination (HR) is one of the important DNA repair systems that maintain the stability of genome [9]. Tumors with BRCA1/2 mutations typically carry a deficient HR system, vital for DNA repair from an insult (reviewed in [10]). It is guesstimated that defects in this system account for up to 40% of familial or spontaneous BC [10]; this high percentage indicates the importance of the HR system in BC etiology [11]. HR contains a complex signaling pathway that plays an important role in repairing stopped replication forks and DNA double-strand breaks (DSB) [12,13].
DBS is the most harmful form of DNA damage and tied closely
to cancer progression. This form can be repaired by the HR system,
as the error-free pathway, in the S and G2 phases of the cell cycle
[12,14,15]. Various enzymes contribute to HR repair, among those,
Endonuclease/exonuclease/phosphatase family domain containing
1 (EEPD1) plays an undeniable role [16]. Particularly, in the
DSB repair, EEPD1 via its 5ˊ exonuclease activity forms 3ˊ single
strands that, in turn, attack to the homologous sequences into the
sister chromatid lead to DSB repair by replication [17]. It seems
that this protein takes a center stage as a gatekeeper and functions
through cleaving the fork by endonuclease activity allowing HR to
be embarked upon [18]. Despite the important functions of EEPD1,
its role in BC is shrouded in mystery. Crossover junction endonuclease
MUS81 is also involved in the HR system, a structure-specific
DNA endonuclease from the conserved xeroderma pigmentosum
group F family that, in turn, cleavage ‘Holliday structures’ in the
complex with Essential Meiotic Structure-Specific Endonuclease-1
(EME1) or EME2 [19,20]. MUS81 also takes part in DNA repair process
whereby the repair and restart of stalled replication forks and
interstrand cross-links proceed [21]. Wu et al. demonstrated that
Mus81 was downregulated in gastric cancer which points to the
way that it might be a tumor suppressor and even a potential marker
for the malignancy of gastric cancer [22]. However, there is little
information showing its contribution to BC.
By and large, according to the importance of these genes in the
HR pathway, it sounds that the alteration of their expression can
impose the cellular damages on cells, leading to genomic instability
and neoplastic changes. Up to now, no studies have been done on
the expression of these genes in BC, so this study was designed to
evaluate the expression of EEPD1 and MUS81 genes in BC tissues.
Materials and Methods
Tissue Collection
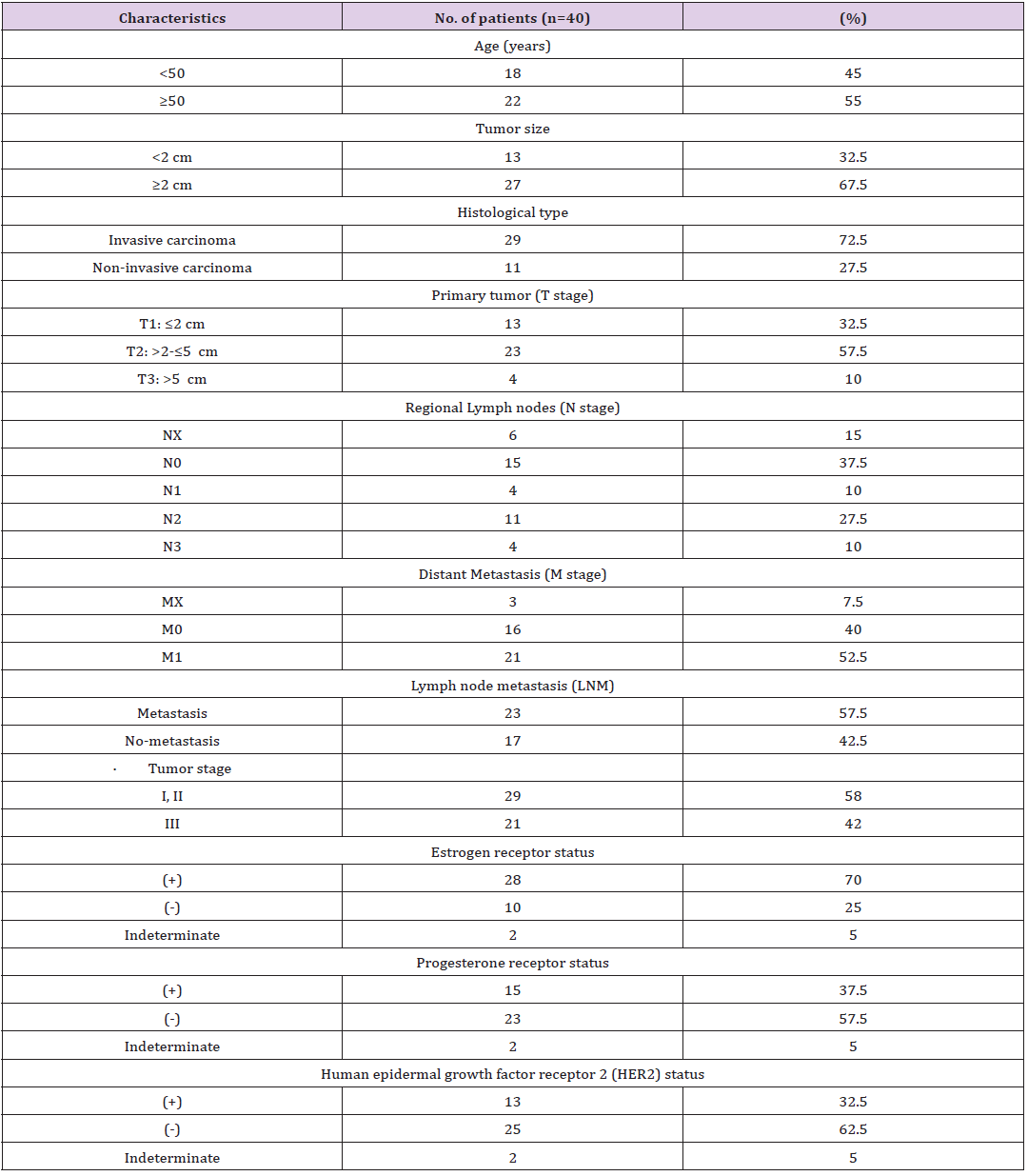
In this case-control study, 40 pairs of BC and adjacent non-tumor tissues were obtained from the Rasoul Akram Hospital, Tehran, Iran. No patient received preoperative chemotherapy or radiotherapy before resection. The study protocol was approved by the ethics committee of Iran University of Medical Sciences, Tehran, Iran, under the ethics code of “IR.IUMS.FMD.REC.1397.160”, and written informed consent was taken from all participants. All tissues (from female patients) were stored in liquid nitrogen immediately upon resection. The BC conditions were diagnosed by pathologists in participating centers based on the American Society of Clinical Oncology guidelines [23]. Cellular differentiation was also graded based on the World Health Organization grading system [24] using the histologic grade, the tumor–node–metastasis (TNM) system for stage classification, and also human epidermal growth factor receptor 2 (HER‐2) status. The clinicopathological parameters, e.g. tumor stage, lymph node metastasis, and TNM stages are listed in (Table 1). According to the data, the estrogen receptor (ER), progesterone receptor (PR), and HER-2 status of each patient were indicated as positive, negative, or indeterminate.
Cell Culture
Human BC cell lines including MCF-7 (ER+ Luminal A-like BC cell model; ATCC® HTB-22™), MDA-MB-231 (Basal-like BC cell model; ATCC® HTB-26™), and SKBR3 (Her2 positive BC cell model; ATCC® HTB-30™) were purchased from the Pasture Institute of Iran. Cells were cultured in RPMI-1640 medium (Nissui Pharmaceutical, Tokyo, Japan) supplemented with 10% heat-inactivated fetal bovine serum (FBS), 2 mM L-glutamine, 100 U per ml penicillin, and 100 μg per ml streptomycin (all from Gibco BRL, Rockville, MD, USA). The cells were incubated at 37 °C in a humidified atmosphere with 5% CO2.
RNA extraction and Complementary DNA (cDNA) Synthesis
Total RNA was extracted from the samples using the TRIzol reagent (Invitrogen, CA, USA) according to the manufacturer’s instruction. The extracted RNA was solubilized in RNase‐free dH2O and its concentration was measured using Nano Drop 2000c (Thermo Fisher Scientific, NY, USA) and the quality of the samples was analyzed by 2% agarose gel electrophoresis. To cut down on any possible DNA contaminants, the samples were also treated with RNase-free DNase I. Around 2 μg RNA was reverse‐transcribed into cDNA using the TAKARA kit (Primer Script TM RT reagent Kit, Japan) according to the manufacturer’s protocol.
Quantitative Real-Time PCR
Quantitative real-time PCR (qRT-PCR) was performed on ABI Step One Sequence Detection System (Applied Biosystems, USA) using an SYBR Green-based PCR Kit. The reaction was performed using a single cycle in 95 °C for 5 min followed by continual 40 cycles (95 °C for 10 sec, 60 °C for 20 sec, and 72 °C for 30 sec). The experiments were applied independently in triplicate and the expression level of each gene was presented as a relative value to hypoxanthine phosphoribosyl transferase (HPRT) gene as an internal control. The primers were designed using Primer 3.0 plus online tool [25] as following: HPRT (Forward: 5 ́-CTGGCGTCGTGATTAGTGATG-3 ́; Reverse: 5 ́-AGACGTTCAGTCCTGTCCATAA‑3 ́), EEPD1 (Forward: 5 ́-AGGCTCGAAGTCTCTGGACA-3 ́; Reverse: 5 ́-GCATCCTTCTTGCTCCAGTC- 3 ́), MUS81 (Forward: 5 ́-AGGCTCTGGCAGCTACTGG-3 ́; Reverse: 5 ́-TTCTGACTCGGCCAACTTCTG-3 ́).
GOBO Analysis
GOBO (http://co.bmc.lu.se/gobo/), an online resource with mRNA microarray profiling data from 51 breast-derived cell lines [26], was used to validate the mRNA expression levels of MUS81 gene, although the expression of EEPD1 was absent in this database. UALCAN (http://ualcan.path.uab.edu/index.html), an interactive web resource that contains a large amount of cancer transcriptome data [27] was also exploited to explore the mRNA expression levels of the candidate genes, EEPD1 and MUS81.
In silico Identification of Gene Expression Associated with Prognosis in BC
Because of this fact the patients follow-up were impossible in our cases, to further validate, we used the Kaplan-Meier plotter database [28] and pan-cancer tool to analyze the relationship between gene expression of EEPD1 and MUS81 and overall survival of 3955 BC patients from the aforementioned database. Hazard ratio, 95% confidence intervals (CI) and log-rank P-values were calculated. P-value < 0.05 was considered statistically significant.
Statistical Analysis
All statistical analyses were performed using SPSS v.25 software (SPSS Inc., Chicago, IL, USA) and GraphPad Prism v.7.0 (GraphPad Prism version 5.0, San Diego, USA). Descriptive statistical analysis including an independent t-test was used to evaluate the expression of EEPD1 and MUS81 in the samples. All experiments were repeated at least three times and P- values < 0.05 were considered to be statistically significant.
Results
Down-Regulation of EEPD1 in BC Tissues
The expression of EEPD1 was significantly decreased in tumor tissues in comparison with adjacent non-tumor tissues (P-value < 0.001). On the other hand, the expression of MUS81 did not show any significant differences in the specimens. Also, a significant downregulation of EEPD1 was found in 70% of breast tumors, while MUS81 down-regulation was remarkable in 54% of BC patients but it did not result in a significant difference (Figures 1a & 1b).
EEPD1 and MUS81 Expression vs. Clinicopathological Characteristics
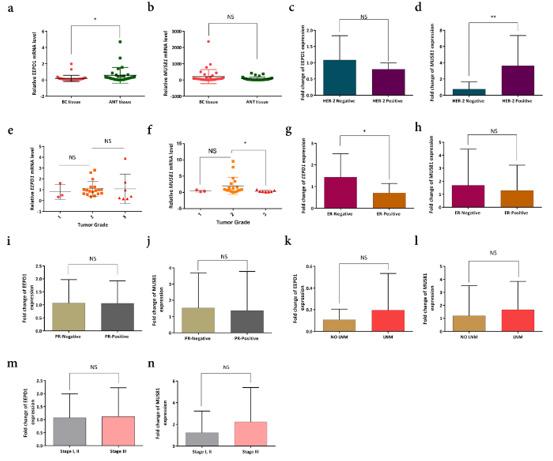
Figure 1:
a. The expression level of the EEPD1 gene in breast cancer [36] tissues compared to the adjacent non-tumor tissues (ANT).
b. the expression level of MUS81 was not significantly different between the samples.
c. and d) expression of MUS81 and EEPD1 in Her-2 positive and negative specimens. MUS81 expression was significantly
associated with Her-2 positive status (** P-value <0.01).
e and f. corellation between gene expressions and tumor grades. Low expression of MUS81 was evident in grade 3, i.e.
expression of this gene increased in tumors with grade 2 and decreased again in grade 3.
g and h. EEPD1 expression was higher in estrogen receptor (ER)-negative BC tissues than positive ones (*P-value < 0.05), while
MUS81 did not show any significant correlation.
i and j. The expression level of candidate genes was not in association with progesterone-receptor (PR) status. k-n. the expression
of MUS81 and EEPD1 did not show any significant correlation with Lymph node metastasis (LNM) or tumor node metastasis
(TNM).
MUS81 expression was significantly higher in Her-2 positive BC patients than negative individuals (P-value < 0.01) (Figures 1c & 1d). The expression of EEPD1 was not correlated with tumor grades, whereas it seemed that MUS81 expression increased in tumors with grade 2 and decreased in grade 3 (Figures 1e & 1f). Besides, the expression of the MUS81 gene did not show any significant difference regarding PR and ER statuses, although low expression of EEPD1 was significantly detected in ER-positive BC samples (Figures 1g-1j). On the other side, we found no association of EEPD1 and MUS81 expression with TNM stages and Lymph node metastasis (LNM) status (Figures 1k-1n).
The Expression of EEPD1 and MUS81 in BC Cell Lines
The experiment was expanded into MCF-7, MDA-MB-231, and SKBR3 cell models. The results showed that the expression levels of EEPD1 and MUS81 increased in all Her-2 positive cell models (Figure 2a).
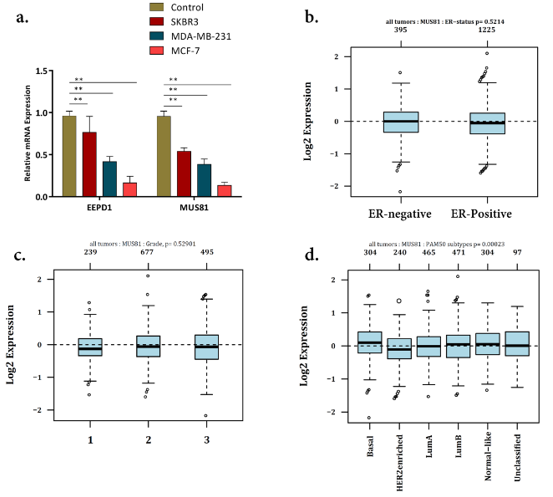
Figure 2:
a) expression level of MUS81 and EEPD1 in breast cancer [36] cell lines.
b) data mining showed no significant differences between MUS81 expression and Estrogen-receptor (ER) status.
c) the GOBO analysis also revealed no significant correlation between MUS81 expression and tumor–node–metastasis
(TNM) staging.
d) the expression of MUS81 in basal-like BC was significantly higher than that in luminal-like based on PAM50 gene
classifiers of BC.
GOBO and UALCAN Analysis for the Candidate Genes
GOBO analysis showed that MUS81 expression levels were inconsistent across the cell lines; however, there was no information in GOBO for EEPD1. The expression of MUS81 showed no considerable differences between ER-positive and ER-negative BC tissues and also in different TNM stages (Figures 2b & 2c). All in all, using GOBO analysis, MUS81 expression was shown to decrease in Her- 2 enriched BC tumors (Figure 2d). The UALCAN analysis showed that while the expression level of MUS81 increased in BC primary tumors (Figure 3a), EEPD1 had a lower expression level in BC compared with normal cases (Figure 3b). The expression of EEPD1 and MUS81 was also shown to have fluctuation in different tumors (Figure 3c & 3d).
Figure 3:
a) expression validation in the UALCAN database for upregulated-MUS81 and downregulated- EEPD1 genes in primary
tumors (Data from The Cancer Genome Atlas (TCGA) database [37]). (** P-value <0.01).
b and c) TCGA data analysis of the mRNA level of MUS81 and EEPD1 in different types of cancer. Cancer types with significantly
decreased EEPD1 mRNA levels were highlighted with a red circle.
The expression of EEPD1 and MUS81 Were Associated with Poor Overall Survival in BC Patients
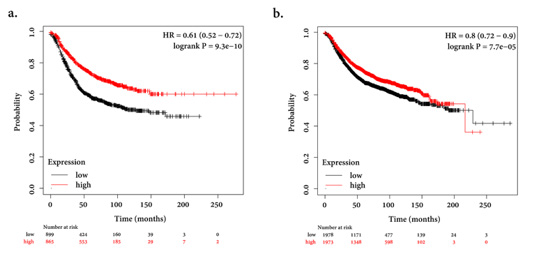
Figure 4:
a) Kaplan-Meier (https://kmplot.com/analysis/) survival curve showing the prognostic value of EEPD1 in predicting BC
using the Pan-cancer analysis which is a part of this database to analyze the prognostic relevance of mRNA in different types
of cancers. Low expression of mRNA indicated poor overall survival.
b) Low expression of MUS81 showed poor survival (HRs; 95 % confidence interval in parentheses).
Using the Kaplan-Meier plotter database, we determined that the overall survival of 3955 BC patients (from the Kaplan-Meier plotter database) is low in cases with decreased expression of EEPD1 and MUS81. The follow-up time was determined in 250 months. The plots showed that low expression of EEPD1 and MUS81 was associated with poor overall survival (Figures 4a & 4b).
Discussion
The majority of BC subtypes are considered to be defective in
the HR process which encompasses important pathways, e.g. DNA
damages repair and DNA interstrand crosslinks, and also plays a
vital role in maintaining the genome stability and integrity. Hence,
in this study, the expression level of EEPD1 and MUS81 as essential
encoding genes for the enzymes involved in HR was analyzed
in breast tumor and adjacent non-tumor tissues. Moreover, the
gene expression was analyzed in cell models. Potential correlations
between the EEPD1 and MUS81 expression levels and clinicopathological
characteristics were also evaluated. EEPD1 is a DNA
repair enzyme plays an important role in repairing large DSBs via
its nuclease activity [17]. Wu et al. demonstrated that EEPD1 (in
complex with EXO1 and BLM) had 5ˊ-overhang nuclease function
[17]. EEPD1 allows the initiation of HR repair and the resumption
of stopped replication fork by cutting the stalled fork, so it plays as a
gatekeeper [18]. Chun et al. confirmed the role of the EEPD1 gene in
the integrity of replication forks and stability of the genome during
cell division in embryonic zebrafish development.
They showed that the depletion of EEPD1, e.g. in BRCA1 deficiency,
results in nuclear defects along with the formation of
anaphase bridges and micronucleus in stressed zebrafish embryos.
As a result, EEPD1 stabilizes the genome during embryonic
replication stress [29]. Therefore, it seems that EEPD1 deficiency
drives cells to genome instability and carcinogenesis. In this study,
the EEPD1 expression was significantly decreased in tumor tissues
(confirmed by cell models and data from UALCAN) compared to
the control group. These findings are consistent with the tumor
suppressor’s function of EEPD1. Using Kaplan–Meier plotter database
[30] and a larger cohort of BC patients from this database, we
showed that patients with low expression of EEPD1 have a lower
chance of survival. We also showed higher expression of EEPD1
in ER-negative BC patients, also conformed in cell models using
SKBR3 as the ER-negative cell line. Although, we could not find any
significant association between EEPD1 expression and other clinicopathological
features, e.g. TNM and LNM.
The MUS81 gene encodes an endonuclease enzyme that repairs
the DSB of the DNA molecule. This gene preserves the integrity of
the genome and is regarded as a tumor suppressor. Many studies
introduced MUS81 as a new diagnostic marker for colorectal cancer
[22,31]. The results of an experiment showed that Mus81-/- and
Mus81+/- mice were susceptible to a variety of cancers including
BC. Therefore, it concluded that the biallelic expression of this gene
in mammalian is necessary to maintain genome integrity and suppress
cancer [32]. Various studies have been performed on the expression
of this gene in different cancers, e.g. Xie et al. showed that
expression of the MUS81 gene in serous ovarian cancer [33] tissue
increased at both transcript and protein levels. In other words, they
found that if they suppress gene expression of MUS81, it led to inhibited
serous ovarian cancer growth and development [34]. As
a contradiction in term, a comprehensive study compared the expression
of the MUS81 gene in hepatocellular carcinoma, colorectal
cancer, and gastric cancer tissues to normal tissue samples from the
same organs and the data indicated that MUS81 gene expression
was significantly reduced in tumor tissues [22].
Down-regulation of MUS81 in colorectal cancer was reported
in correlation with tumor metastasis and progression [35-37]. In
the present study, we could not find a significant difference regarding
the expression between BC and adjacent non-tumor tissues,
even though using UALCAN, we demonstrated that the expression
of this gene is increased in tumor cells. Herein, we elucidated that
MUS81 expression increased in Her-2 positive patients than negative
ones. GOBO analysis showed that Her-2 enriched BC samples
had lower MUS81 expression in comparison with other subtypes
of BC. It was consistent with the findings in cell lines, showing that
the expression of this gene increases in SKBR3, confirmed by GOBO
database. GOBO analysis also showed that MUS81 had a constant
expression in different tumor grades, while in the present study
we demonstrated that MUS81 expression increased in grade 2 and
decreased again in grade 3. Using Kaplan–Meier plotter database,
we evidenced that MUS81 expression in patients with poor survival
decreased.
Conclusion
In a nutshell, this study proposes EEPD1 as a prognostic marker for BC, in which, its expression decreases. However, further studies are still needed to determine how and why EEPD1 is downregulated in BC. Regarding MUS81, we could not find a significant difference among BC and adjacent non-tumor tissues; we believe that future studies should aim to shed light on results in a larger cohort of BC patients. However, our study was only conducted at the mRNA level and requires further analysis at the protein level as well. Notwithstanding, future large-scale prospective studies are still required to confirm the data.
Acknowledgement
This study is financially supported by Iran University of Medical Sciences, Tehran, Iran, in 2018. All parts of the work were carried out in the ‘Genetic laboratory of Ali Asghar Children’s’ and Rasoul Akram Hospitals, Tehran, Iran. The authors of the current study would like to express their deep thanks to all staff of these hospitals.
Declarations
Ethics Approval and Consent to Participate
The study protocol was approved by the ethics committee of Iran University of Medical Sciences, Tehran, Iran, under the ethics code of “IR.IUMS.FMD.REC.1397.160”, and written informed consent was taken from all participants.
Consent for Publication
Not applicable.
Availability of Data and Materials
Not applicable.
Competing Interests
The authors declare that they have no conflicts of interest.
Funding
Not applicable.
Authors’ Contributions
MM, ES, MKh, ER, and AZ collected the clinical data, wrote the paper and drew the figures; HA, ER and MM analyzed the clinical data and drew the table; ER searched papers; and NA provided guidance for article writing. All authors read and approved the final manuscript.
References
- Bachawal S, Bean G R, Krings G, Wilson K E (2020) Evaluation of ductal carcinoma in situ grade via triple-modal molecular imaging of B7-H3 expression. NPJ Breast Cancer 6(1): 1-9.
- Dai X, Fagerholm R, Khan S, Blomqvist C, Nevanlinna H (2015) INPP4B and RAD50 have an interactive effect on survival after breast cancer. Breast cancer research and treatment 149(2): 363-371.
- Wang J, Ding Q, Fujimori H, Motegi A, Miki Y, et al. (2016) Loss of CtIP disturbs homologous recombination repair and sensitizes breast cancer cells to PARP inhibitors. Oncotarget. 7(7): 7701-7714.
- Li X, Heyer WD (2008) Homologous recombination in DNA repair and DNA damage tolerance. Cell research 18(1): 99-113.
- Bindra RS, Schaffer PJ, Meng A, Woo J, Måseide K, et al. (2004) Down-regulation of Rad51 and decreased homologous recombination in hypoxic cancer cells. Molecular and cellular biology 24(19): 8504-8518.
- Chodosh L A (2011) Breast cancer: current state and future promise. Breast Cancer Res. 13(6): 113.
- Yalcin B (2013) Staging, risk assessment and screening of breast cancer. Exp Oncol 35(4): 238-245.
- Parshad R, Price FM, Bohr VA, Cowans KH, Zujewski JA, et al.(1996) Deficient DNA repair capacity, a predisposing factor in breast cancer. British Journal of Cancer 74(1): 1-5.
- Moynahan ME, Jasin M, (2010) Mitotic homologous recombination maintains genomic stability and suppresses tumorigenesis. Nature reviews Molecular cell biology 11(3):196-207.
- Belli C, Duso BA, Ferraro E, Curigliano G (2019) Homologous recombination deficiency in triple negative breast cancer. The Breast 45: 15-21.
- Den Brok WD, Schrader KA, Sun S, Tinker AV, Zhao EY, et al. (2017) Homologous recombination deficiency in breast cancer: A clinical review. JCO Precision Oncology 1: 1-13.
- Peng G, Lin CCJ, Mo W, Dai H, Park YY, et al.(2014) Genome-wide transcriptome profiling of homologous recombination DNA repair. Nature communications 5:3361.
- Carr AM, Lambert S (2013) Replication stress-induced genome instability: the dark side of replication maintenance by homologous recombination. Journal of molecular biology 425(23): 4733-4744.
- Hopfner K P (2014) ATP puts the brake on DNA double‐strand break repair: A new study shows that ATP switches the Mre11‐Rad50‐Nbs1 repair factor between signaling and processing of DNA ends. Bioessays36(12): 1170-1178.
- Barton O, Naumann SC, DiemerBiehs R, Künzel J, Steinlage M, et al. (2014) Polo-like kinase 3 regulates CtIP during DNA double-strand break repair in G1. J Cell Biol. 206(7): 877-894.
- Liu Y (2016) EEPD1: Breaking and Rescuing the Replication Fork. PLoS genetics 12(2): e1005742.
- Wu Y, Lee SH, Williamson EA, Reinert BL, Cho JH, et al. (2015) EEPD1 rescues stressed replication forks and maintains genome stability by promoting end resection and homologous recombination repair. PLoS genetics 11(12): e1005675.
- Kim HS, Nickoloff JA, Wu Y, Williamson EA, Sidhu GS, et al. (2017) Endonuclease EEPD1 is a gatekeeper for repair of stressed replication forks. Journal of Biological Chemistry. 292(7): 2795-2804.
- Amangyeld T, Shin YK, Lee M, Kwon B, Seo YS (2014) Human MUS81-EME2 can cleave a variety of DNA structures including intact Holliday junction and nicked duplex. Nucleic acids research. 42(9): 5846-5862.
- El Ghamrasni S, Cardoso R, Halaby M, Zeegers D, Harding S, et al. (2015) Cooperation of Blm and Mus81 in development, fertility, genomic integrity, and cancer suppression. Oncogene. 34(14): 1780-1789.
- Doe CL, Ahn JS, Dixon J, Whitby MC (2002) Mus81-Eme1 and Rqh1 involvement in processing stalled and collapsed replication forks. Journal of Biological Chemistry 277(36): 32753-32759.
- Wu F, Shirahata A, Sakuraba K, Kitamura Y, Goto T, et al. (2010) Down-regulation of Mus81 as a potential marker for the malignancy of gastric cancer. Anticancer research 30(12): 5011-5014.
- Kris MG, Hesketh PJ, Somerfield MR, Feyer P, ClarkSnow R, et al. (2006) American Society of Clinical Oncology guideline for antiemetics in oncology: Update 2006. Journal of clinical oncology. 24(18): 2932-2947.
- NHSBSP R (2005) Pathological Reporting of Breast Disease: A Joint Document Incorporating the Third Edition of the NHS Breast Screening Programmes Guidelines for Pathology Reporting in Breast Cancer Screening and the Second Edition of the Royal College of Pathologists' Minimum Dataset for Breast Cancer Histopathology. Sheffield: NHSBSP.
- Untergasser A, Nijveen H, Rao X, Bisseling T, Geurts R, et al. (2007) Primer3Plus, an enhanced web interface to Primer3. Nucleic acids research 35(suppl_2): W71-W74.
- Ringnér M, Fredlund E, Häkkinen J, Borg Å, Staaf J(2011) GOBO: Gene expression-based outcome for breast cancer online. PloS one 6(3).
- Chandrashekar DS, Bashel B, Balasubramanya SAH, Creighton CJ, PonceRodriguez I, et al. (2017) UALCAN: A portal for facilitating tumor subgroup gene expression and survival analyses. Neoplasia 19(8): 649-658.
- Hou GX, Liu P, Yang J, Wen S (2017) Mining expression and prognosis of topoisomerase isoforms in non-small-cell lung cancer by using Oncomine and Kaplan–Meier plotter. PloS one 12(3): e0174515.
- Chun C, Wu Y, Lee SH, Williamson EA, Reinert BL, et al. (2016) The homologous recombination component EEPD1 is required for genome stability in response to developmental stress of vertebrate embryogenesis. Cell Cycle 15(7): 957-962.
- Györffy B, Lanczky A, Eklund AC, Denkert C, Budczies J, et al. (2010) An online survival analysis tool to rapidly assess the effect of 22,277 genes on breast cancer prognosis using microarray data of 1,809 patients. Breast cancer research and treatment 123(3): 725-731.
- Wu F, Shirahata A, Sakuraba K, Kitamura Y, Goto T, et al. (2011) Downregulation of Mus81 as a novel prognostic biomarker for patients with colorectal carcinoma. Cancer science 102(2): 472-477.
- McPherson JP, Lemmers B, Chahwan R, Pamidi A, Migon E, et al. (2004) Involvement of mammalian Mus81 in genome integrity and tumor suppression. Science 304(5678): 1822-1826.
- ToralLópez J, GonzalezHuerta LM, Sosa B, Orozco S, González HP, et al. (2011) Familial pycnodysostosis: Identification of a novel mutation in the CTSK gene (cathepsin K). Journal of Investigative Medicine 59(2): 277-280.
- Xie S, Zheng H, Wen X, Sun J, Wang Y, et al. (2016) MUS81 is associated with cell proliferation and cisplatin sensitivity in serous ovarian cancer. Biochemical and biophysical research communications 476(4): 493-500.
- Wu F, Chen W, Tan G, Liu J (2014) Expression of Mus81 gene in Chinese Han patients with colorectal cancer and its clinical significance. Nan fang yike da xuexue bao 34(12): 1776-1779.
- Tripp RA, Haynes LM, Moore D, Anderson B, Tamin A, et al. (2005) Monoclonal antibodies to SARS-associated coronavirus (SARS-CoV): Identification of neutralizing and antibodies reactive to S, N, M and E viral proteins. Journal of virological methods 128(1-2): 21-28.
- Tomczak K, Czerwińska P, Wiznerowicz M (2015) The Cancer Genome Atlas (TCGA): An immeasurable source of knowledge. Contemporary oncology 19(1A): A68-A77.

 Research Article
Research Article