Impact Factor : 0.548
- NLM ID: 101723284
- OCoLC: 999826537
- LCCN: 2017202541
Kenji Ohe*1, Yoshihiro Harada1, Yuta Horita1, Hiroyoshi Harada1, Hiroki Terai1, Masayoshi Mori1, Yusuke Murata1, Kenji Ashida2, Munechika Enjoji1
Received: May 31, 2018; Published: November 16, 2018
*Corresponding author: Kenji Ohe, Department of Pharmacotherapeutics, Faculty of Pharmaceutical Sciences, Japan
DOI: 10.26717/BJSTR.2018.11.002058
The hERα gene is known for its major role in estrogen dependent growth of breast cancer. ERα positivity is a hallmark in treating these cancers. Though there are many isoforms reported for hERα, the splicing regulation, especially for the 5’ non-coding exons is largely unknown. We have recently reported that the oncogenic product, high-mobility-group protein A1a (HMGA1a) can induce the expression of hERα46 through regulating alternative splicing of the hERα gene. An RNA decoy of HMGA1a can reverse this splicing event as well as alter estrogen responsiveness of MCF-7 breast cancer cells. Here, we will review on the emerging issue of HMGA1a as a sequence-specific RNA-binding protein and show experimental evidence of the HMGA1a RNA-binding site found in the seed sequence of a microRNA and possible feedback loop.
Keywords: ERα Gene; Hmga1a; Non Coding Exon; Splicing Regulation; Microrna; Tamoxifen Resistance
Abbreviations: HERα: Human Estrogen Receptor Alpha; HGRα: Human Glucocorticoid Receptor Alpha; HMGA1a: High Mobility Group Protein A1a; HPS2: Human Presenilin-2; PR: Progesterone Receptor; FOS: FBJ Murine Osteosarcoma Viral Oncogene Homolog; TFF1: Trefoil Factor 1; ERBB2: v-erb-b2 Avian Erythroblastic Leukemia Viral Oncogene Homolog 2; Her2, Human Epidermal Growth Factor Receptor 2; miR-16: MicroRNA 16; mir21: microRNA 21; RNU-6: U6 Small Nuclear RNA
MCF-7 tumors were removed from transplanted nude mice and quickly frozen in liquid nitrogen. The tumors were crushed into small pieces and put into an Eppendorf tube with TRIzol reagent (Invitrogen). The tissue was grinded with a disposable grinder and total RNA, protein was purified following the manufacturer’s instruction of TRIzol reagent. In brief, after passing the extract through a 25G needle five times, the sample was centrifuged.
For RT-PCR, the aqueous phase was precipitated by isopropanol treated by RQ1 DNase (Promega, M6101) and total RNA was precipitated. One μg of total RNA was subjected to reverse transcription by Rever Tra Ace (TRT-101) and 0.5μl of the reverse transcribed product was subjected to subquantitative PCR (25 to 30 cycles) by TaKaRa Ex Taq (TaKaRa Bio, RR001A). For immunoblot assays, protein was purified from the interphase and organic phase by precipitating with 6V of a solution for precipitating protein (50% Ethanol, 24.5% Acetone, 24.5% Methanol, 1% distilled water). Twenty micrograms of protein were boiled in sample buffer and separated by 10% sodium dodecyl sulfate-PAGE (SDSPAGE), transferred to a PVDF membrane and analyzed by primary antibodies: ERα antibody (HC-20) (Santa-Cruz, sc-543) or HMGA1a antibody (Santa Cruz, FL95) and appropriated secondary antibodies and revealed by Chemi-Lumi One L (Nacalai Tesque, 07880-54).
For micro-RNA expression analyses, total RNA was reverse transcribed by miScript II RT Kit (QIAGEN, 218160) and miR-16, miR-21, RNU-6 was amplified by miScript Primer Assay (QIAGEN, 218300).
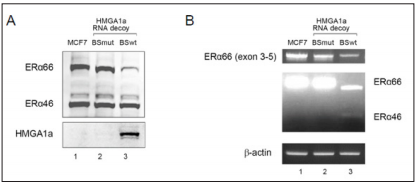
Multiple non-coding exons are frequently found in the 5’ region of hormone receptor genes. Most typical are hERα and hGRα (Figure 1). The hERα gene is encoded of eight exons with five 5’ non-coding exons, exons F to B from 5’ to 3’, that form F to B hERα mRNA in addition to A hERα mRNA that is transcribed immediately upstream of exon 1 [1]. More non-coding exons were found with other numerous variants for hERα [2-4] (Figure 1A). Likewise, the hGRα gene is encoded of nine exons with eight 5’ non-coding exons. Non coding exon 1A is far upstream and activated by glucocorticoids [5]. The rest are expressed in a tissue specific manner with exons 1-D, 1-E, 1-B, 1-F, 1-C, 1-H from 5’ to 3’. The ABCs are the order they were discovered [6] (Figure 1B). The regulation of multiple non-coding exons in the 5’ region of genes involves alternative splicing as well alternative promoter selection. We have found a factor, HMGA1a, previously known as a DNA-binding protein and oncogenic product that binds RNA in a sequence-specific manner and regulates alternative splicing of the ERα gene [7,8]. The 5’ splice site of the hGRα variant exon 1-D has a HMGA1a RNA-binding site adjacently upstream the 5’ splice site (5’-UGCCGCACAAGgu-3’). This hGRα HMGA1a RNA-binding site resembles the original HMGA1a RNAbinding site adjacently upstream the 5’ splice site of hPS2 exon 5 (5’-GCUGCUACAAGgu-3’) [9,10] and the one adjacently upstream the pseudo-5’ splice site of hERα exon 1 (5’-GCGGCUACACGgu-3’) [7,8] with a GC-rich stretch followed by common nucleotides shown in bold and underlined (Figure 2A). It has recently been reported that HMGA1a binds stems of 7SK RNA (5’---CCG---loop---CGG---3’; the HMGA1a binding site is underlined) [11] (Figure 2B). As for hGRα exon 1-D, a CGG trinucleotide can be found 17 nt downstream the CCG trinucleotide found in the hGRα exon 1-D HMGA1a RNAbinding site. However, this kind of HMGA1a RNA-binding site could not be found for the HMGA1a RNA-binding site in PS2 exon 5 or hERα exon 1.
Figure 1: Multiple 5’ non-coding exons of the hERα and hGRα genes and HMGA1a RNA-binding site.
A. Structure of the hER gene showing tissue distribution of each isoform.
B. Structure of the hGR gene showing tissue distribution of each isoform.
C. Comparison of HMGA1a RNA-binding sites in hPS2 exon 5 and hERα exon 1 with the one yet to be confirmed in hGRα exon 1-D.
D. The HMGA1a RNA-binding site (red letters) found in 7SK RNA.
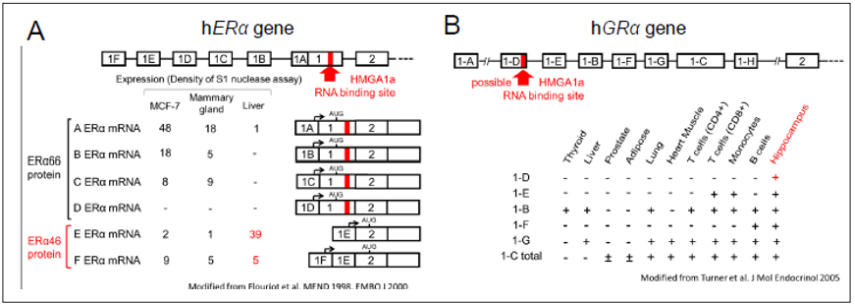
Figure 2: HMGA1a RNA-binding sites.
A. Comparison of HMGA1a RNA-binding sites in hPS2 exon 5, hERα exon 1, and the one yet to be defined in hGRα exon 1-D.
B. The HMGA1a RNA-binding site (red letters) in 7SK RNA
This HMGA1a RNA-binding site was originally discovered as a factor binding adjacently upstream the 5’ splice site of PS2 exon 5 [9]. PS2 as well as Presenilin-2 and amyloid-precursor-protein are known to harbor mutations in familial Alzheimer’s disease [12,13]. The challenge was to seek for alternative spliced variants of these genes in the brain of sporadic Alzheimer’s disease patients. A relentless search for aberrantsplicing variants found one for PS2 exon 5 [14]. The patients who harbor these aberrantly spliced PS2 exon 5 isoform showed vulnerability to endoplasmic reticulum stress and increased production of beta-amyloid [15]. In a pursuit to identify the factor responsible of this exon-skipping, HMGA1a was found to bind a region adjacently upstream the 5’ splice site of PS2 exon 5. HMGA1a interacted with U1-70K, a component of the U1 snRNP that is important for 5’ splice site recognition by the spliceosome (splicing complex). HMGA1a was expressed in splicing speckles of the cell and showed increased expression in the CA1 region of the hippocampus [9] just as the resulting spliced product, PS2V did (70% of sporadic Alzheimer’s disease patients’ brains compared to 17.6% of control brains [14].
However, it was puzzling because when HMGA1a was added to a purified U1 snRNP/ 5’ splice site RNA reaction, the recognition was improved. Carefully looking over the splicing reaction, it crossed our minds that U1 snRNP needed to dissociate to carry on the spicing reaction. Indeed, it was the trapping of U1 snRNP by HMA1a that made HMGA1a inhibit splicing and induce exon-skipping. Remarkably, this effect was only found when the HMGA1a RNA binding site was adjacent the 5’ splice site [10]. In a search for other targets for HMGA1a, the human estrogen receptor alpha (hERα) gene came to our minds, because a vast list of literature had been reporting the relation of hERα and HMGA1a in breast cancer cells. First, we made a search (using EMBL-EBI Pairwise Sequence Alignment tool (EMBOSS Needle: https://www.ebi.ac.uk/Tools/psa/emboss_ needle/nucleotide.html)) for the PS2 HMGA1a RNAbinding site with a gu at the 3’ terminal end (5’-GCUGCUACAAGgu-3’) in the hERα gene. We found a candidate in hERα exon1, 33 nt upstream the authentic 5’ splice site. Exon-skipping of hERα exon1 would result in a shorter isoform lacking the AF-1 domain of hERα called hERα46. A transcript consisted of 5’ non-coding exons E and/ or F (E/F ERα mRNA) also skips exon 1 [1]. Overexpression of HMGA1a induced exon-skipping of the endogenous hERα gene We transiently expressed a decoy of the HMGA1a RNA binding site (bold) with a microRNA nucleus localization signal (underlined) at its 3’ end (5’-GCUGCUACAAGAGUGUU [16]. This HMGA1a RNA decoy efficiently blocked HMGA1a-induced skipping of hERα exon1 [10]. We also pursued the mechanism of this exon-skipping. Unlike PS2 exon 5, the 5’ splice site adjacent the HMGA1a RNA binding site is 33 nt upstream the authentic 5’ splice site. Using psoralen UV crosslinking combined with RNA-EMSA, we were able to show that HMGA1a trapped U1 snRNP at the upstream but not the authentic 5’ splice site [8]. We believe the 5’ splice site of the hGRα variant exon 1-D will work in a similar manner. It is known well that exon 1-D is expressed exclusively in the hippocampus along with all other hGRα exon 1s, and that exon 1-D is important for neuron-specific expression of hGRα [17].
The expression of exon 1-D is possibly regulated by the Ying Yang 1(YY1) transcription factor [6]. YY1 is known to enhance the expression of APBB1 (amyloid beta precursor protein binding family B member 1), a factor whose expression level correlates with the risk of developing, and severity of late-onset Alzheimer’s disease [18,19]. Thus, the target of HMGA1a may also be exon 1-D of hGRα. Indeed, in senescence accelerated mouse (SAM) [20] the expression of GRα was found to be attenuated [21]. The importance of the GRα signaling pathway can be found in Addison’s disease patients where impaired episodic memory is a frequent symptom [22].
One important target of HMGA1a RNA binding site is microRNA. In a search of a seed sequence of microRNA that may resemble the HMGA1a RNA binding site, we found miR-16 as a candidate. In theory, HMGA1a would bind to the antisense strand of miR-16 in its pre-primary miR (Figure 3A) (boxed) and impair its maturation. This would result in further enhanced HMGA1a expression because HMGA1a is one of miR-16 targets [23]. A decoy RNA of HMGA1a RNA-binding site (Figure 3A) (blue letters) would bind the seed sequence of miR-16 and would also inhibit the function of miR-16. The primary miR-16 in the nucleus, as a stem loop, has no 5’-CCG-3’ and complementary 5’-CGG-3’ sequence as 7SK RNA (Figures 3B & 2B).
Figure 3: Predicted HMGA1a RNA-binding site in miR-16.
A. HMGA1a RNA-binding site (black box) in human pre-primary-miR-16 (mature miR-16 in red letters) (hsa-miR-16-5p). HMGA1a RNA-binding site shown as reversed sequence (blue letters) and complementarity to seed sequence of miR-16.
B. hsa-miR-16-5p as pre micro-RNA.

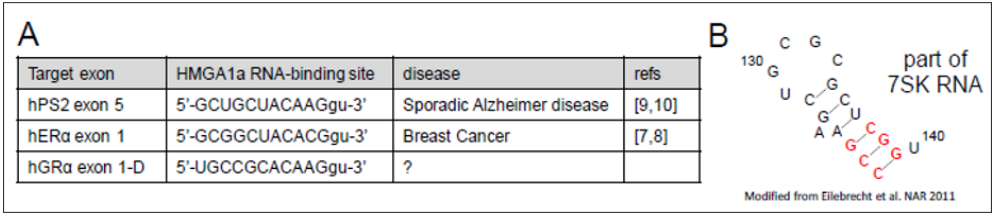
To test the relation of miR-16 and HMGA1a RNA-binding function, we checked its expression in the tumors of stably expressed MCF-7 cells of the HMGA1a RNA decoy transplanted into nude mice [7] as well as well as full length C type isoform of ERα expressed in mammary tissue and ERα target genes. To our surprise, the expression of the C type full length ERα was slightly lower in HMGA1a RNA decoy-expressed cells (Figure 4A). Thus, we also checked the expression of ERα target genes: PGR (progesterone receptor); FOS (FBJ murine osteosarcoma viral oncogene homolog); TFF1 (trefoil factor); and ERBB2 (v-erb-b2 avian erythroblastic leukemia viral oncogene homolog 2, also known as Her2 (human epidermal growth factor receptor 2)) but could find no differential expression with the control decoy RNA (Figure 4B) (lane 2 & 3). We though that the decreased expression of full length ERα may have increased its expression to compensate the low expression of type C full length ERα to maintain the expression of ERα target genes. We also checked the expression of micro RNAs, especially HMGA1a targeting miR16, in these HMGA1a RNA decoy stably expressed MCF-7 tumors. As expected, the expression of miR16 decreased (Figure 4C) probably because of the HMGA1a RNA decoy. However, the HMGA1a RNA decoy would also bind HMGA1a to prevent its binding to the antisense strand of primary miR-16. This makes the function of HMGA1a RNA decoy complicated due to its biphasic effect on miR-16 expression; binding to the seed sequence of miR-16; and binding to HMGA1a to prevent HMGA1a RNA-binding to the antisense strand of primary miR-16. In this respect, the expression of ERα46/ERα66 mRNA ratio may not be the same as prior transplantation. Thus, we checked expression ERα46 and ERα66 protein in stably expressed HMGA1a RNA decoy of MCF-7 tumors, we found the expression reversed (Figure 5A) (upper panel) compared to prior transplantation [7].
Figure 4: Expression of ERα and its target as well as miR-16 in stably expressed HMGA1a RNA decoy of MCF-7 tumors.
A. Expression of full length ERα66 mRNA,
B. Expression of ERα-target genes: PR, FOS, TFF1, and ERBB2.
C. Expression of miR-16, miR-21, and control: RNU-6. The control of A and B are also referred to RNU-6.
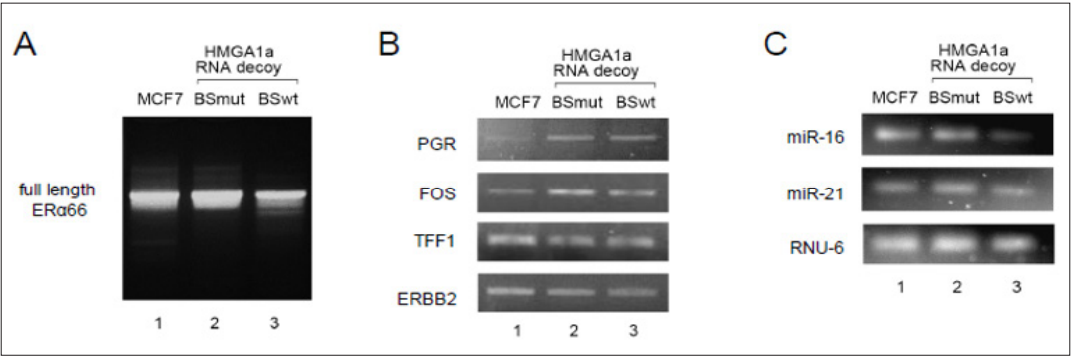
Briefly, HMGA1a RNA decoy is known to decrease the HMGA1ainduced aberrantly spliced ERα46 mRNA and increase the type D/F full length ERα66 mRNA and concomitant protein level in MCF-7 cells as we have reported [7]. We checked whether HMGA1a protein was re-expressed due to the decreased miR-16 expression by HMGA1a RNA decoy, and this was indeed the case (Figure 5A) (lower panel). The expression of ERα46 and ERα66 mRNA showed the same results (Figure 5B). The mechanism of this reversed effect may be due to the biphasic function of HMGA1a RNA decoy expressed above, but maybe simply due to reduced effect of the HMGA1a RNA decoy because it is easy to imagine that stably expressed would be much lower than transiently expressed. However, this would only decrease the ratio and would not reverse ERα46 and ERα66 or induce HMGA1a expression.
Figure 5: Reversed effect of HMGA1a RNA decoy.
A. HMGA1a RNA decoy stably expressed MCF-7 tumors exhibit a reversed effect of ERα46 and ERα66 protein ratio (upper panel), due to re-expression of HMGA1a (lower panel).
B. The reversed effect of ERα46 and ERα66 mRNA ratio by HMGA1a RNA decoy stably expressed in MCF-7 tumors.
We have reported the first splicing factor that regulates splicing of 5’ non-coding exons of the ERα gene. An HMGA1a RNA decoy inhibits HMGA1a RNA-binding function, reduce ERα46 expression, and sensitize MCF cells to tamoxifen by the restored estrogendependent growth [7]. Since HMGA1a is also known as a DNAbinding factor, in vitro splicing assays were performed to exclude the possible regulation through promoter selection [8]. Here, we show that the seed sequence of miR-16, an HMGA1a-targeting micro-RNA, is complementary to the HMGA1a RNA-binding site. Since it is known that increase of HMGA1a expression in cancer cells increase its malignant potential [24] the increased HMGA1a may decrease its own targeting miR-16 by binding to the antisense strand of primary miR-16.
This possible miR-16-mediated RNA-binding feedback loop of HMGA1a (Figure 6) also explains the results we showed here. Prior to transplantation, HMGA1a RNA decoy prevents the HMGA1ainduced exon-skipping of ERα exon 1 in MCF-7 cells [7] and by in vitro splicing assays [8]. However, the long incubation period of cloning stable MCF-7 transfectants of HMGA1a RNA decoy induces exon-skipping rather than exon-inclusion by inducing HMGA1a expression (Figure 5) a result of attenuated miR-16 expression (Figure 4C). A feedback loop mediated by micro-RNA has been reported for SF2/ASF (SRSF1) [25]. The micro-RNA feedback loop of HMGA1a is the second of such to be reported and differs in that it negatively regulates its own targeting micro-RNA. The limitations of this study is that we have not shown direct in vitro analyses of HMGA1a binding to the primary miR-16. Future studies will focus on the simple induction of miR-16 in tamoxifen-resistant MCF-7 (TAMR1) cells that may lower HMGA1a expression and its RNAbinding properties.
KO conducted all the experiments and wrote the manuscript Y Ha, YHo, YB, HH, HT, MM, YM, TT, MT, IA, KA, KK, TN, TU, TY, ME with assist of KO and Ta Y, KO. Wrote the paper. IA, MT, TN, KK, ME, TY supervised and arranged the experiments.


