Impact Factor : 0.548
- NLM ID: 101723284
- OCoLC: 999826537
- LCCN: 2017202541
Furtado LL Campos WF, Silva MC, Nicolino RR and Pereira AA*
Received: August 21, 2018; Published: August 29, 2018
*Corresponding author: Pereira AA, Institute of Agrarian Sciences, Federal University of Vale do Jequitinhonha e Mucuri (UFVJM), Brazil
DOI: 10.26717/BJSTR.2018.08.001663
Through the analysis of molecular biology, the relations between organisms and the evolutionary order of the species were inferred, not only through visual characteristics but also through genomic characteristics. Vitamin B1 (Thiamine) is a vital micronutrient required by all organisms. In view of the importance of this coenzyme, this work analyzed the TPK gene of the thiamine biosynthesis pathway in the evolution of terrestrial plants, using phylogenetic analysis tools. Thus, by evaluating the variations in protein level of the TPK gene, it was possible to reflect the evolution of the large groups of plants, and even the proximity between species, families and orders.
Assuming that all organisms and their characteristics come from evolution, it is possible to determine, with comparative analysis, relationships between species. Therefore, studies of comparative and systematic biology classify species to obtain this understanding [1]. The first classifications were based on morphological and environmental characteristics, thus relating ancestors and descendants and determining an order of known plant species [1]. On the other hand, molecular phylogeny is the most accurate way to classify species. There is now a need to define genes that express biological evolution as well as the evolutionary relationship between organisms. Vitamin B1 (Thiamine) is a coenzyme required by all organisms. The active form, thiamin diphosphate (TDP), is an essential cofactor for enzymes of key metabolic pathways, such as glycolysis, acetyl-CoA synthesis, Krebs cycle, alcoholic fermentation, pentose's-phosphate pathway, Calvin cycle, branched chain amino acid pathway and biosynthesis of pigments in photosynthetic organisms [2,3].
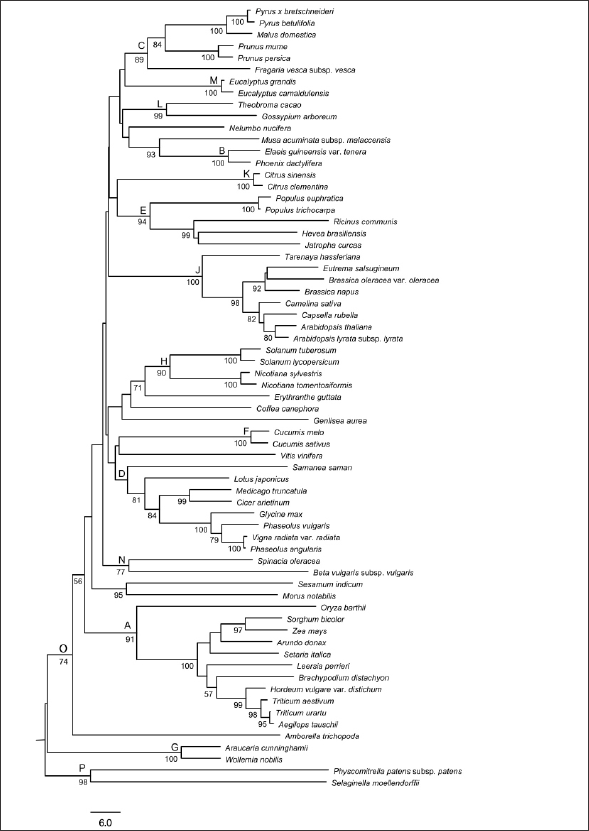
This important coenzyme is biosynthesized in bacteria, fungi and plants but not in animals [3,4] and the genes present in the biosynthesis pathway are well known in plants [5]. Despite this, they have not yet been used as a marker gene in the study of plant evolution. In this sense, the amino acid sequence of the enzyme thiamine-pyrophosphokinase (TPK) of Arabidospis thaliana (AT5G55630), which catalyzes the pyrophosphorylation of thiamine to generate TDP [6], was used to analyze the evolution of plants. The phylogenetic tree indicates the presence of the TPK gene in several species and there was the separation of the clades between gymnosperms and angiosperms (Figure 1), according to the evolution of the groups in the phyla of plants. In addition, within the angiosperms, the species Amborella trichopoda Baill was shown to be basal to all angiosperms (Figure 1), as demonstrated by APG IV (2016). The relations of informal groups within the angiosperms, identified in APG IV (2016), Fabids, Malvids, Commelinids and Lamiids are not defined in the tree presented. However, the formal groups as orders are well delimited (Figure 1). Within Fabids, orders Rosales (branch C) and Fabales (branch D) are monophyletic in all protein sequences (Figure 1).
The thymine biosynthesis pathway TPK gene was used as a marker and showed major similarity with the topology explained by APG IV (2016) and effective separation of large groups (cryptogamas, angiosperms and gymnosperms).
To the National Council for Scientific and Technological Development (CNPQ) for financial support.
Figure 1: Phylogenetic tree based on the TPK protein sequences, using the MEGA 6 software, through the algorithm Neighbor-joining Tree e bootstrap de 1000 replicates. Legend: A= Poales, B=Arecales, C=Rosales, D=Fabales, E= Malpighiales, F=Cucurbitales, G=Pinales (Gminospermas), H=Solanales, J= Brassicales, K=Sapindales, L= Malvales, M=Caryophyllales, N=Caryophyllales, O= Angiosperma e P=Criptogamas.