Impact Factor : 0.548
- NLM ID: 101723284
- OCoLC: 999826537
- LCCN: 2017202541
Katerina D Tzimourta1, Athanasios Tsilimbaris2,3, Katerina Tzioukalia3, Alexandros T Tzallas3, Markos G Tsipouras3, Loukas G Astrakas1 and Nikolaos Giannakeas3*
Received: July 31, 2018; Published: August 06, 2018
*Corresponding author: Nikolaos Giannakeas, Department of Computer Engineering, School of Applied Technology, Technological Educational Institute of Epirus, Greece
DOI: 10.26717/BJSTR.2018.07.001535
Sleep disorders have a great impact in the patients’ quality of life. The study of human sleep during the different sleep stages is crucial in the diagnosis of sleep disorders and is mainly performed with polysomnography (PSG). In this work, a methodology for sleep staging using solely Electroencephalographic (EEG) signals from PSG recordings is presented. EEG signals from the ISRUC-Sleep dataset are selected and used, aiming to automatically identify the five sleep stages. Initially, the EEG signal is filtered in order to extract the five EEG rhythms and the energy is calculated in each sub-band and used to train several typical classifiers. Results in terms of classification accuracy reached 75.29% with Random Forests.
Keywords: Sleep Staging, Isruc Database, Automatic Sleep Stage Classification, Assc, Eeg, Energy, Random Foreststreatment
Abbreviations: SAS: Sleep Apnea Syndrome, REM: Rapid Eye Movement, NREM: Non-Rapid Eye Movement, AASM: American Academy of Sleep Medicine, ASSC: Automatic Sleep Staging Classification, PSG: Polysomnography, EEG: Electroencephalogram, EOG: Electrooculogram, EMG: Electromyography, MODWT: Maximum Overlap Discrete Wavelet Transform, SVM: Support Vector Machines, KNN: k-Nearest Neighbor
Sleep is a fundamental restorative process for human mental and physical health [1]. The nightlong study of human sleep and sleep-related behaviors during the different sleep stages is essential in the diagnosis of sleep disorders. Sleep disturbance and disorders, such as the life-threatening Sleep Apnea Syndrome (SAS), can have devastating effects on both the quality of life and essential human activities, including learning and memorization. Monitoring the patient throughout the night sleep and identifying the alterations in sleep patterns, plays a significant role in the accurate diagnosis and in the implementation of the appropriate treatment plan. Sleep is a structured sequenced process comprised of five stages, which progress cyclically. The sleep-wake cycle consists of an awake stage, a non-rapid eye movement stage (NREM), which is further divided into transitional sleep (N1), light sleep (N2) and deep sleep (N3) stages, and then a rapid eye movement sleep stage (REM) [2]. The N3 stage sometimes is considered as two separate stages (N3 and N4) and it is referred as slow wave sleep [3]. Monitoring of these different stages through use of recorded neural, respiratory, and cardiac activity during sleep, can provide an assessment of sleep in patients suffering from sleep disorders.
Sleep stages are characterized by standardized manual scoring. Sleep scoring was previously accomplished using the standard human sleep scoring methods proposed by Rechtschaffen and Kales [4], which were then used as a basis for the new criteria on sleep staging, determined by The American Academy of Sleep Medicine (AASM) [5]. These methods allow for analysis of sleep disorders but suffer from expensive and convoluted procedures, as well as inefficient manual scoring that can be subjective, as scoring results and conclusions differ between experts [2]. Accurate and efficient automatic sleep staging, in order to improve sleep evaluation for the diagnosis of sleep disorders, is therefore of great significance today. One of the methods currently used to monitor and score sleep is polysomnography (PSG). PSG entails simultaneous recording of multiple physiological signals, such as electroencephalogram (EEG), electrocardiography (ECG), electromyography (EMG), electrooculogram (EOG), oxygen saturation (SpO2), and respiration, of a patient asleep over a full night [6]. Subsequently, the recordings are split into short periods of time (30s), called epochs, which are then categorized into the different sleep stages. The procedure usually takes place in a well-equipped hospital, by an experienced sleep specialist who reviews the PSG recordings and detects the sleep-related patterns.
Categorizing these epochs can be done using EEG waves and the characteristic differences in several frequencies (EEG rhythms) observed at each sleep stage. The low frequency band (delta) contains EEG waves of 0.5-4Hz frequency. Theta waves are between the frequencies of 4-8Hz and alpha between the frequencies of 8-13Hz. There are also beta frequency waves (13-35Hz), which are further discriminated into beta 1 waves (13-22Hz) and beta 2 waves (22-35Hz). Special categories of waves, which can be found in the EEG sleep recordings and are characteristic of the sleep stage, are the K-complexes (0.5-1.5 Hz) and the sleep spindles (12-14Hz). The awake stage includes low amplitude mixed frequency waves, whereas the N1 stage has dominant alpha waves. The N2 stage has sleep spindles and k-complexes, whereas the N3 stage has delta waves and the REM stage has sawtooth waves [7].
Electroencephalogram (EEG) is a cost-effective and typically a non-invasive method for monitoring and recording of electrical signals and voltage fluctuations from neurons in the brain, using electrodes attached to the scalp of a patient. These sensors allow for measurement of signals, which can be recorded over a long period of time, such as during sleep. EEG can therefore give great insight into the neural activity of a patient, with the aim of analyzing and evaluating the quality of sleep, in order to provide treatment and resolve health problems associated with sleep disorders. It also allows for potential automation, resulting in automatic sleep scoring to an accurate level with the aim of having a less arduous and objective sleep evaluation. The study of allnight sleep and sleep patterns analysis has gained the research interest over the past years. Methods for automatic sleep staging classification (ASSC) have been evaluated in datasets from various sleep centers. The most well-known databases that have been widely utilized throughout the years are the database provided by Physionet [8] containing PSG recordings from 20 subjects, the Siesta Sleep Database, which comprises PSG data from 300 subjects (100 subjects with sleep disorders and 200 healthy) [9] and the ISRUC-sleep Dataset of 118 subjects, which is used in this study and is subsequently analyzed [3].
Several methods have been proposed based on the PSG recordings by Physionet [6,10,11] or by the Siesta dataset [12-14]. Concerning the ISRUC database, few studies have been proposed [2,3,15-17] that either used the maximum overlap Discrete Wavelet Transform (MODWT) for signal decomposition [2,15,17] or filters [16] and extracted an extensive set of features. Then, the most discriminative features were selected and used to train a Support Vector Machines (SVM) [2,3,15,17] or a Bayesian classifier [16]. In this paper, an EEG-based method for classification of sleep stages is presented. The method evaluates the energy of the EEG rhythms from the ISRUC sleep Dataset in order to train several classifiers and discriminate the 5 stages of sleep. Results for the 5-class problem are presented.
The proposed method consists of two stages: feature extraction and classification in 5 classes. Digital FIR filters are applied to EEG recordings, obtained from 100 patients, aiming to extract the energy in each sub-band. Then, the extracted feature vector is used to train and test several classifiers. The filter design and feature extraction steps are implemented using MathWorks MATLAB platform, whereas the classification step is performed with the Waikato Weka software. A flowchart of the proposed method is presented in Figure 1.
The dataset used for this work is the ISRUC-Sleep dataset [3]. The dataset contains polysomnographic data collected during the night sleep for about eight hours, creating three groups of data;
a) 100 adults with sleep disorders evidence and one recording per subject
b) 8 adults with sleep disorders evidence and two recordings per subject and
c) 10 healthy subjects and one recording per subject. Data collection was performed in the Sleep Medicine Centre of the Hospital of Coimbra University, according to the suggestions of the AASM manual. Each PSG recording consists of signals from 19 channels, containing EEG, EOG and chin EMG signals. Each signal was sampled at 200Hz and was segmented into epochs of 30sec. Two different sleep experts evaluated the recordings and performed the staging according to the AASM guidelines. A comprehensive report of the dataset is presented in [3]. In this methodology, the group of 100 subjects is employed and only the EEG signals of each patient is used in the experiments. Hence, 6 EEG channels (F3-A2, C3-A2, O1-A2, F4-A1, C4-A1 and O2-A1) are extracted and the energy of each sub-band per signal is calculated. Also, the staging according to the evaluation of the one sleep expert is followed. In Table 1 the total number of epochs of each stage (87187 epochs in total) is shown. In Figure 2 the first 30-sec epoch of each EEG channel for Patient 1 is presented.
In order to extract data in the different frequency sub-bands, FIR filters are designed and applied to each EEG signal. Therefore, a low-pass (0-4Hz), a high-pass (30-60Hz) and three band-pass filters (4-8Hz, 8-13Hz and 13-30HZ) are designed and the energy is extracted in each sub-band of interest, corresponding to the EEG rhythms. Since each EEG signal contains information from 6 channels, a feature vector of 30 characteristics (5 sub-bands * 6 channels) is used as input to train the classifiers.
To evaluate the proposed method for automatic sleep staging, five well-known classifiers are used.
Naïve Bayes [18] is a simple classifier that combines a probability model with a decision rule. The classifier operates on the basic assumption that the features comprising the feature vector are statistically independent. Naïve Bayes is based on Bayes decision theory and the purpose is the minimization of classification error probability and the maximization of posterior probability. The classifier adopts the Gaussian assumption, meaning that each marginal is described by two parameters (mean and variance), reducing computational complexity and the need for data. A small number of training data is needed for the classifier to estimate the necessary classification parameters, avoiding overfitting, making the Naïve Bayes suitable for complex classification problems.
A Decision Trees classifier is based on a set of decision rules. The tree root node is at the top and is connected with other nodes through links or branches. The procedure is repeated until no further links exist to connect to other nodes.
According to the architecture of the classifier, only one link can be followed each time and the following node becomes the root node of the next sub-tree until no other decision can be made. Hence, Decision Trees is a straightforward classifier [19].
The k-Nearest Neighbor [20] is a supervised, non-parametric instance-based method and one of the simplest machine learning algorithms used for classification. According to this method, an instance is classified to the most relative class depending on the vote its k nearest neighbors. This approach is a good technique when there is little or no knowledge of the distribution of the data.
Support Vector Machines [21] is a technique for linear and nonlinear classification problems. The input features are represented into a high-dimension (usually much higher than the original) feature space aiming to be linear separated. This projection is performed by the kernel function. The gap that separates the data is called decision hyperplane and the distance from the hyperplane is named margin. The goal in training an SVM is to find the optimal separating hyperplane with the largest margin, leading to better generalization of the classifier. In our experiments, the radial basis function is used as kernel function.
Random Forests [22] is an ensemble learning method for classification that operates by combining decorrelated decision trees. The ultimate goal of the classifier is to decrease the generalization error and improve accuracy. In Random Forests the trees are grown by randomly selecting at each node a group of features to determine the split. The feature set is sub-decomposed into subsets of random values. These subsets are used to train and test each individual decision tree. After a large number of trees is generated, they vote for the most popular class. In our experiments, 100 decision trees is selected.
The 10-fold cross-validation technique is employed to train and test the five classifiers. The performance of each classifier is depicted in Table 2, in terms of True Positive (TP) rate (average for all classes), precision (average for all classes) and accuracy. The best classification accuracy is obtained from the Random Forests (75.29%) followed by Support Vector Machines, k-Nearest Neighbor, Decision Tree and Naïve Bayes. (66.59%, 65.61%, 64.77% and 55.97% respectively). Table 3 presents the confusion matrix for the Random Forests classification, which as can be seen from Table 1 indicated the best classification accuracy. The percentage of classified instances for each class (row) is calculated and depicted in the table, thus the main diagonal includes the TP rate of each respective row class, while all other values in the same row are the misclassification rates of this class to all other classes. The confusion matrix that is presented in Table 3, gives a thorough description of the classification. The percentage of N1 epochs correctly classified by as N1 is less than 50% while the majority of the instances were classified as N2 (37.63%), Awake (19.29%) and REM (11.10%). Also, epochs of REM sleep are misclassified as N2 (22.27%), N1 (6.96%) and Awake (5.91%), leading to a total accuracy for the REM stage equal to 63.50%. The percentages of correctly classified N2 and N3 epochs are above 80% (82.78% and 81.82% respectively) and the majority of errors is due to misclassified epochs as Awake (26.10%) and N2 (17.12%) respectively. The best discrimination, matching with the scoring assigned by the expert, is provided with epochs of Awake stage (90.43%).
In this work, an EEG-based method for 5-class automatic sleep staging is presented. The EEG signals from the ISRUC Sleep Database of 100 subjects with evidence of sleep disorders are extracted from PSG recordings and utilized for the analysis. FIR Filters are applied to the EEG signals, extracting the EEG rhythms. Then, the energy from each sub-band is used to train several classifiers aiming to discriminate the five stages of sleep. Table 1 shows the obtained results for the five classifiers described above. The best classification accuracy (75.29%) is obtained with Random Forests. Concerning the errors in classification of each sleep stage, the misclassification problems are mainly related to epochs N1 and REM, as can be also confirmed by previous studies [2,3,16,17]. This error is mainly observed due to the similarity between the EEG patterns presented in Rapid Eye Movement stage and N1 stage (slow eye movement) [16]. Furthermore, the applied FIR filters decompose the EEG signals into specific frequencies of interest that correspond to the EEG rhythms.
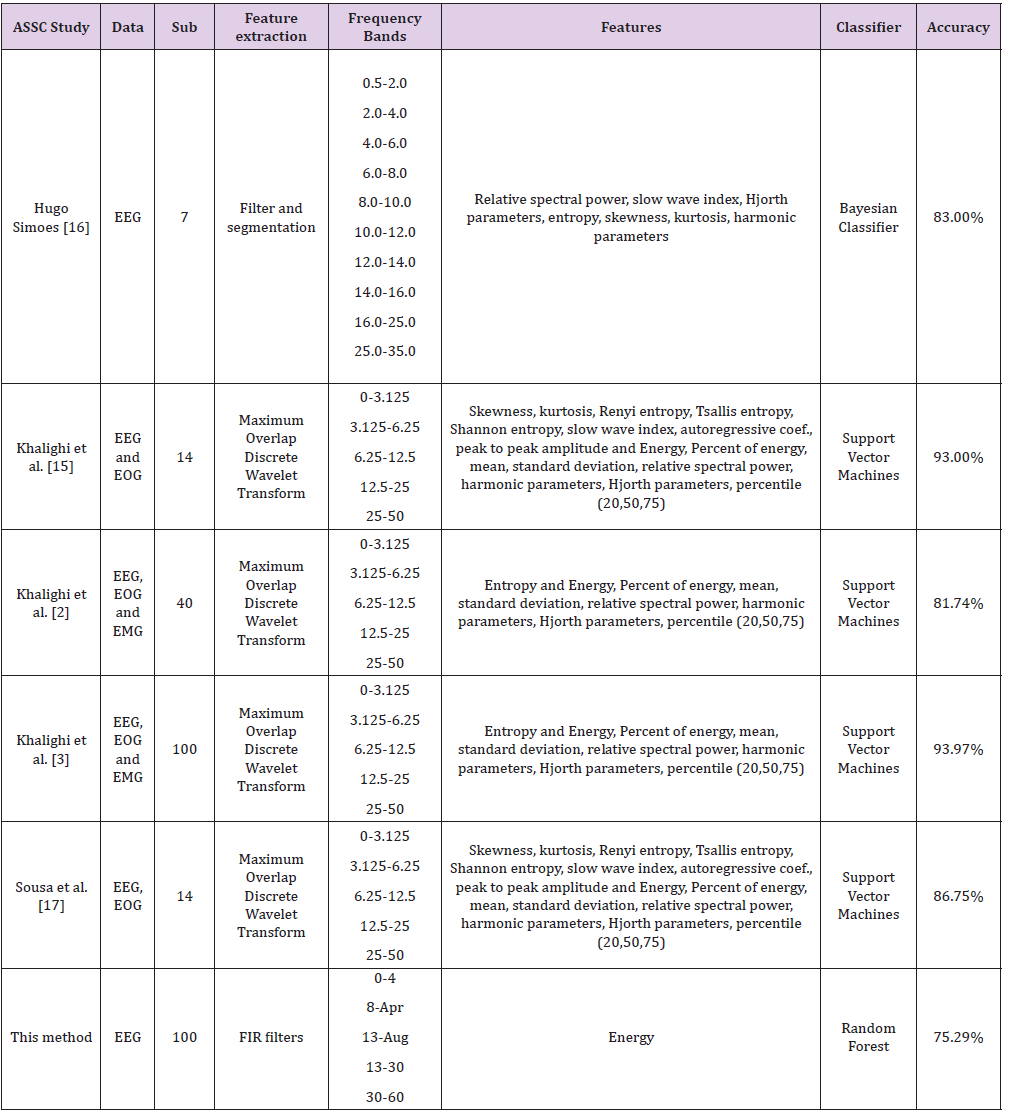
However, the maximum overlap Discrete Wavelet Transform, which is proposed in previous studies [2,15,17] decompose the signal depending on the sampling frequency into frequencies that correspond approximately to EEG rhythms. (e.g. the sub-band related to alpha rhythm is 6.25-12.5Hz with MODWT instead of 8-13Hz). Since the sleep stages are characterized by specific patterns found to certain EEG rhythms, a filter-based approach seems more appropriate. To the best of the authors’ knowledge, only one previous study [3] has utilized the total of 100 subjects with evidence of sleep disorders. Previous studies have been validated only in a small number of the entire dataset (14 subjects [15] and 40 subjects [2]). Also, in the proposed approach only the EEG channels of the recordings are used. However, most of the previous studies [2,3,15,17] utilized EOG and chin EMG signals in combination with EEG signals to provide the best classification results. Hence, greater accuracy values in previous studies can be partially explained due to the smaller number of subjects involved in the experiments and the combination of EEG signals with EOG and EMG (Table 4).
Table 4: A Comparison of performances of the various methods proposed in the literature that have utilized the ISRUC Sleep Dataset for automated sleep staging.
Sleep studies are usually performed for the diagnosis of sleeprelated pathologies. Sleep scoring is performed by sleep experts who review the polysomnographic recordings and obtain the different sleep stages. Scientific research has been focused on automatic sleep staging classification, aiming to assist the sleep experts in sleep scoring and hence, in identification of sleep disorders. In this work, an EEG-based method for automatic sleep stage classification based on the energy extracted from the EEG rhythms is presented FIR filters were applied to EEG signals to extract the frequencies of interest and results showed 75.29% accuracy with Random Forests. In the future, more features will be examined and the ability of several feature selection methods will be evaluated.


