Impact Factor : 0.548
- NLM ID: 101723284
- OCoLC: 999826537
- LCCN: 2017202541
Nazish Mazhar Ali*, Bushra Mazhar and Abdullah Mazhar
Received: May 20, 2018; Published: May 29, 2018
*Corresponding author: Nazish Mazhar Ali, CG University, Lahore, Pakistan
DOI: 10.26717/BJSTR.2018.05.001130
Samples were collected from water, soil, honey and Sewage. Bacteria with ability to produce cysteine were isolated. 50 bacteria were isolated, of which 20 produced amino acids like lysine, cysteine, methionine, glutamic acid, Valine etc. From 20 isolates only 05 produced significant amount of cysteine. The cysteine production by bacteria was improved using different fermentation media and mutation including UV rays and with nitrous oxide. Using different fermentation media, bacterial strains produced different amount of cysteine. Bacterial strain MM5 produced significant cystiene which was 7.8gram/litre.
Keywords: Amino acid production, Cysteine, Microbial production of cysteine
Amino acids are found cheaper to produce by bacterial fermentation rather than by chemical synthesis [1] but some amino acids e.g methionine are still produced chemically more frequently as compared to biologically. Attempts have been made to overproduce biologically active L-cysteine using fermentation [2], but no cysteine fermentation has been commercialized. Lysine [3] another essential amino acid for humans, is also produced from bacteria; it is used as a food additive to enhance its nutritional value. Lysine is produced by Brevibacterium flavum, Corynebacterium glutamicum many other bacterial strains [4,5]. Commercial production of cysteine has not been successfully done yet by microbial fermentation. There is need to isolate those bacterial strains which can not only synthesize cysteine, methionine, lysine and other essential amino acids but also used to produce these amino acids in large quantities, so that they can be commercially fermented and utilized.
Samples of different sources were collected from different localities. The sources include water, soil and honey and these were collected from different areas. The Soil sampling was done from the sugar factories and stables while the honey samples were also from different areas. From these samples, bacterial strains capable of producing specific amino acids like cysteine were isolated. The primary growth media used to isolate these bacterial strains was Nutrient agar and LB agar [6]. MacConkey agar and Nutrient broth were also used for isolation and propagation of bacteria [7,8]. The soil and water sample dilutions were spreaded on LB/Nutrient agar plates. These agar plates were incubated for 24 hours at 37ºC. After 24 hours bacterial CFUs (colony forming units) appeared on surface of agar plate. These colonies were picked and streaked on other agar plates and again for 24 hours. Different fermentation media were prepared to check the ability of these bacterial strains to produce different amino acids especially methionine. The selected bacterial strains were grown in glucose based, urea based and molasses based fermentation media. The fermentation duration was form 24 hours till 96 hours. The broth culture was monitored after every 24 hours. The broth culture was centrifuged at 4000- 5000rpm for 15 minutes. The supernatant containing amino acids was removed from centrifugation tube and filtered using 0.45μm membrane filters. This filtrate containing amino acids was subjected to qualitative and quantitative analysis. The qualitative analysis was done using paper chromatography method. To find out the amount of each amino acid produced, paper chromatography method will be carried out by using Whatman’s paper [9]. The results were confirmed by comparing Rf values with standard amino acids.
For quantification of amino acids the method used was Acidic Ninhydrin method. In this method the ninhydrin reagent is used which reacts with the amino acids and this mixture is analysed via spectrophotometry. The standard amino acid solutions were estimated in same way. The bacterial isolates were also characterized by doing different morphological and biochemical tests [10-12]. These tests included Gram’s staining, Citrate utilization, Indole production and Glucose utilization test [13]. Optimum growth conditions including optimum pH and temperature were also studied. To improve the production of amino acids by bacterial strains, these strains were exposed to mutations which included both Radiations (UV) and Chemical exposures (Nitrous oxide). In radiation method, the selected bacterial strains were exposed to ultra violet (UV) rays and then these bacterial isolates were again be incubated to observe their survival. In chemical mutation, the bacterial strains were treated with different chemicals (mutagens) and then these strains were grown again to observe their viability and production capacity. After obtaining the best producers (bacterial strains) different fermentation media were used to grow them and then the enhanced amount of cysteine produced by these strains was observed.
Different bacterial strains MM1, MM2, MM3, MM4 and MM5 were isolated from different sources. Bacterial strain MM1 was isolated from soil sample. It is Gram -ve rod and is catalase positive. Optimum temperature is 37ºC and optimum pH is 7.0. it showed production of cysteineMM2 was isolated from honey sample. It is Gram +ve bacteria which can utilize glucose, indole and citrate in medium. The optimum temperature for its growth is 30 ºC and optimum pH is 7.0. MM3 sample was also isolated from honey sample and it is a Gram –ve rod and is able to utilize catalase and can ferment glucose. MM4 is a Gram +ve and branched rod bacterial strain which was isolated from honey sample. MM5 strain showed better cysteine production as compared to MM4. Its production is quite significant as compared to other three strains. It is a Gram +ve rod and isolated from soil sample. Its optimum temperature is 37 ºC and optimum pH is 7.0 (Tables 1-3) (Figure 1).
Table 1: Amount of cysteine produced by different bacterial strains during different time intervals.
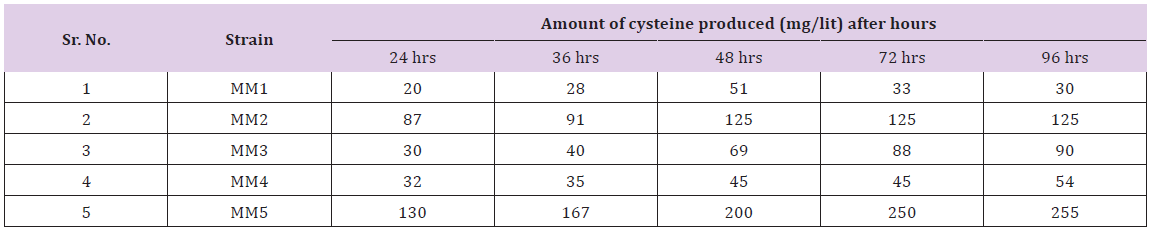
Table 2: Amount of cysteine produced by different bacterial strains (after mutation with nitrous oxide) during different time intervals.
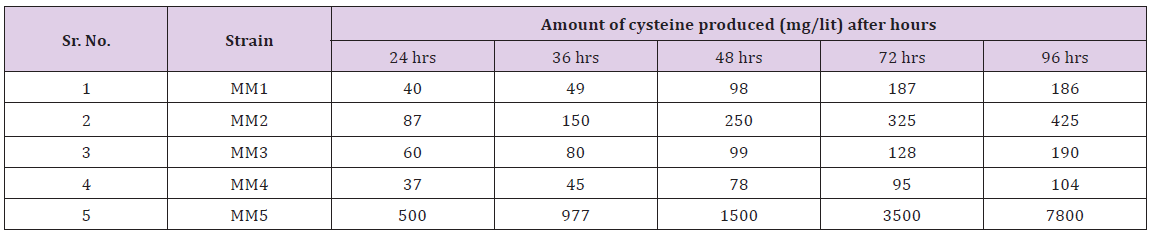
Table 3: Amount of cysteine produced by different bacterial strains (after mutation with UV rays) during different time intervals.
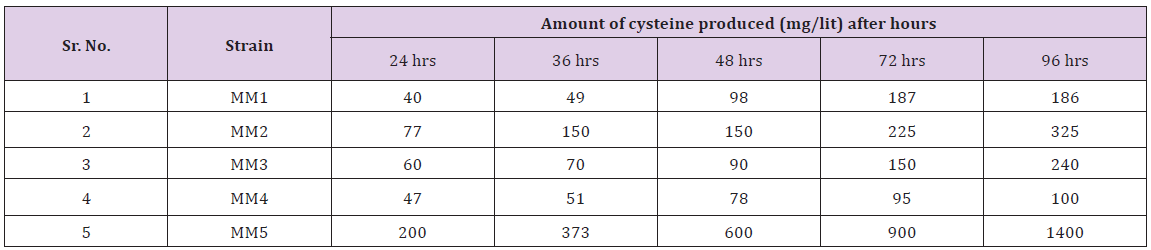
The micro-organisms are now commercially used to produce amino acids via microbial fermentation [13,14]. The amino acid lysine is commercially produced now days using different bacterial strains. The most important of them is Corynebacterium glutamicum which is involved in large scale production of methionine. Other bacteria include E. coli and some species of Bacillus. Corynebacterium glutamicum, B. subtilis, Streptomyces, E. coli have the ability to produce the amino acid cysteine. These were identified on the basis of different biochemical tests. But for further confirmation their ribotyping will be done. Ribotyping is the latest technique by which the chromosomal DNA of bacteria is isolated and sequenced. The bacterial strains under study were grown on different fermentation media which also include the sugar factory waste and corn factory waste i.e, molasses and corn steep liquor respectively. These are considered the cheap source of carbon and other organic compounds. During this study, the strains were grown on these media and were incubated for 96 hours. During these hours, the sample was taken and processed and readings were observed. MM5 gave significant amount of cysteine.


