Impact Factor : 0.548
- NLM ID: 101723284
- OCoLC: 999826537
- LCCN: 2017202541
Abdellah Hedjoudje*1, Stéphane Koch1, Lucine Vuitton1 and Allen Zhang2
Received: February 22, 2018; Published: March 08, 2018
*Corresponding author: Abdellah Hedjoudje, Service de gastro-entérologie, Centre hospitalier Régional universitaire de Besancon, CHRU Jean Minjoz, 1 rue Alexandre Fleming, 25030 Besancon, France
DOI: 10.26717/BJSTR.2018.02.000829
Objective: Electronic medical records include detailed information on clinical care. Besides its clinical utility, they afford researchers to evaluate impact of diagnostic and therapeutic decisions on patient outcomes. However, these dataset are not shared mainly for confidential reasons and remain consequently underused. MIMIC-III ('Medical Information Mart for Intensive Care') is a large, single-center database comprising information relating to patients admitted to critical care units at a large tertiary care hospital has been made publicly available. The objective of this study is to investigate and introduce the database content though the descriptive epidemiology of a severe cholangitis patient cohort.
Method: We conducted a retrospective study of patients with severe cholangitis admitted to the intensive care unit of the Beth Israel Deaconess Medical Center using the MIMIC-III v1.4 ('Medical Information Mart for Intensive Care'). We investigated type of data available in the dataset such as patient characteristics, mortality, drug prescription, microbiology data and fluid balance.
Result: MIMIC-III v1.4 stores 26 different tables with an overall number of 46,520 patients accounting for 58,976 hospitalizations. Records are stored in 21 tables and 5 tables are dictionary tables. The largest file contains more than 330 millions monitoring values data. We identified 125 patients with a severe cholangitis associated with a septic shock. Mean age was 70.75 (12.57) years. 52% of patients were male. Of the 125 patients, 27 died at the hospital leading to a in-hospital mortality of 21, 77%. Patients stayed 7.1 (9.35) in intensive care unit on average. Mean fluid balance was 111.34 (73.15), 48.34 (55.94), 34.94 (50.83), 24.25 (48.79), 27.42 (38.92), 27.73 (61.59) and 26.36 (55.66) at 24, 48, 72, 96, 120, 144 and 168 hours. The most frequent microorganism found was E. Coli on 23, 41% followed by K. pneumonia in 6,12% of specimens. Blood culture was the microbiological test most widely prescribed to check for infectious pathogens.
Conclusion: MIMIC-III v1.4 is a large, single-center database that contains a very large number of patients and hospital admissions. As illustrated by the septic shock associated cholangitis patients, a very broad type of data is stored.
Abbreviations: ALT: Alanine Aminotransferase; AST: Aspartate Aminotransferase; BIDM: Beth Israel Deaconess Medical Center; CAM-ICU: Confusion Assessment Method for the Intensive Care Unit; ETOH: Ethanol; ETT: Endo Tracheal Tube; FSPN: Spontaneous Breathing Frequency; GCS: Glasgow Coma Scale; ID: Internal Diameter; INR: International Normalised Ratio; LDH Lactate Deshydrogenase; LLE: Left Lower Extremity; LLL: Left Lower Lobe; LOC: Level Of Consciousness; LUE: Left Upper Extremity; LUL: Left Upper Lobe; MS: Mental State; NBP: Non-invasive Blood Pressure; PAW: Airway Pressure; PEEP Positive End-Expiratory Pressure; PICC: Peripherally Inserted Central Catheter; PTT: Partial Thromboplastin Time; Richmond-RAS Scale - Richmond Agitation Sedation Scale; RLE: Right Lower Extremity; RLL: Right Lower Lobe; RUE: Right Upper Extremity; RUL: Right Upper Lobe; VTI: Velocity Time Integral; WBC: White blood count
It has been estimated in 2009 that as much as 85% of research investment is wasted [1]. This waste concerns all types of research and occurs at all stages of the production of research evidence, from the choice of questions that are not relevant to patients and their physicians to under-reporting of trial methods and results [1-8]. A part of this waste is due to the underuse of data generated from the process of medical care at patient-level. This underuse of precious medical information is due to several reasons including the difficulty of accessing, organizing and using data entered on paper or charts. Even when this data is correctly stored on computers, patient information remains strictly confidential and a requires a careful de-indentification process before being shared. Within that in mind, researchers from Massachusset Institute of Technology and Beth Israel Deaconess Medical Center created the MIMIC-III ('Medical Information Mart for Intensive Care') database. MIMIC III is large, freely-available database comprising deidentified health- related data associated with approimately fifty thousand patients who stayed in critical care units of the Beth Israel Deaconess Medical Center between 2001 and 2012. MIMIC III database was first released on 25 August 2015, as an update of MIMIC II database [9].
Besides, changing the name from "Multiparameter Intelligent Monitoring in Intensive Care" to "Medical Information Mart for Intensive Care" [10], the database has also been augmented with collected data between 2008-2012. In addition, many data elements have been regenerated from the raw data in a more robust manner to improve the quality of the underlying data. Several versions of MIMIC III have been developed since its first release with minor improvements. . The latest version (MIMIC III v1.4) has been released on 2 September 2016 and has been used for this report. We therefore anticipate that MIMIC III database will be used at a large scale on multiple field of medicine including hepato- gastroenterology. The open nature of the data allows clinical studies to be reproduced and improved in ways that would not otherwise be possible. In this article, we aims at presenting MIMIC III v1.4 database by describing its tables and content though the descriptive epidemiology of a severe cholangitis cohort.
This a retrospective study describing part of the content of MIMC III database by describing a cohort of critically-ill patient hospitalized from 2001 to 2012 at Beth Israel Deaconess Medical Center (BIDMC) (Boston, MA, USA) due to a septic shock secondary to a severe cholangitis. Information derived from the electronic medical records of 46,476 unique critical care patients admitted to the intensive care unit. Use of the MIMIC-III database has been approved by the Institutional Review Boards of BIDMC and MIT, and waiver of informed consent was granted. After completing a National Institutes of Health (NIH) web-based training course (Protecting Human Research Participants), we obtained approval to download and use MIMIC III database [11].
MIMIC III is available online and patient data were de-identified in a Health Insurance Portability and Accountability Act-compliant manner. We included adult patients older than 18 years admitted directly to the intensive care unit from the emergency department with cholangitis associated with a septic shock. The diagnosis of septic shock and cholangitis were made based on the International Classification of Diseases, 9th Revision (ICD-9), by uing the DIAGNOSES_ICD and D_ICD_DIAGNOSES tables. Severe sepsis and/ or septic shock were defined as presence of infection and acute organ dysfunction. We excluded patients that have been admitted multiple times to intensive care units.
We investigated demographic information such as age, sex, race, type of medical insurance, religion, marital status. Chart notes from CHARTEVENTS table were analysed. Outcome of interest were in-hospital mortality, ICU length of stay and fluid balance. Type of laboratory specimens taken and microorganisms found were also investigated.
Patient with missing data (demographics, clinical or fluid intake/ output), or with alternative diagnoses other than cholangitis on admission to the ICU were excluded. Descriptive data is presented as counts and percentages for categorical variables and mean (SD) or median (IQR) for continuous variables. Statistical analysis was performed on R (R version 4.0.2) on a Linux station (Ubuntu 15.04).
MIMIC-III v1.4 is a relational database consisting of 26 tables (Table 1). Tables are linked by identifiers which usually have the suffix 'ID'. The three most important identifiers are SUBJECT_ID, HADM_ID and ICUSTAY_ID that refer to a unique patient, a unique hospital admission and a unique admission to an intensive care unit respectively. Medical records are stored in 21 tables. Broadly speaking, as shown in Table 1, administrative data and patient characteristics are stored in 10 tables. 5 tables are prefixed with 'D_' and correspond to dictionary tables and provide definitions for identifiers. For example, every row of CHARTEVENTS is associated with a single ITEMID which represents the concept measured, but it does not contain the actual name of the measurement. By joining CHARTEVENTS and D_ITEMS on ITEMID, it is possible to identify the concept represented by a given ITEMID. Similarly, every row of LABEVENTS is associated with a single ITEMID, which concept could be identified by joining to the table D_LABITEMS.
Table 1: An overview of the data tables comprising the MIMIC-III (v1.3) critical care database.

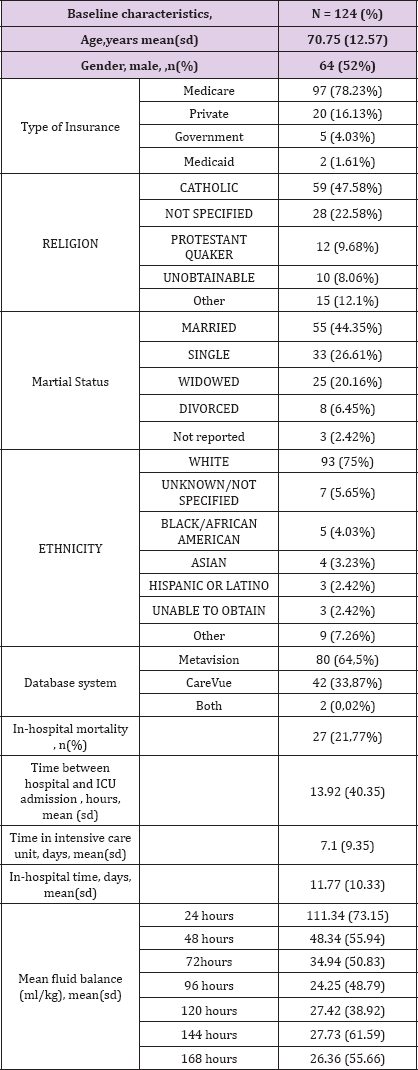
(Table 2) Patient with cholangitis associated with septic shock accounts for 124 hospitalizations between 2001 and 2012. The mean patient age is 70.75 (12.57). 52% of patients are male. MIMIC III database contains very broad type of data including, for instance, the type of patient insurance, religion, marital status or ethnicity Patient insurance were Medicare, Private, Government and Medicaid for 78.23%, 16.13%, 4.03% and 1.61% respectively. We can also find that patient is catholic for 47.58% and protestant for 9.68%. Religion was not specified for 22.58% and unobtainable at admission for 8.06%. We can also find that the largest ethnic group is white Caucasian. (n = 93, 75%). 27/124 (21.77%) patients died during their hospital stay. Mean time between hospital admission and intensive care unit admission was 13.92 (40.35) hours. The mean length of stay at the intensive care unit is 7.1 (9.35) days. The mean length of stay at the hospital is 11.77(10.33) days.
Table 2: Details of MIMIC population

Two different critical care information systems were in place over the data collection period: Philips CareVue Clinical Information System (models M2331A and M1215A; Philips Healthcare, Andover, MA) "carevue" and iMDsoft MetaVision ICU (iMDsoft, Needham, MA) "metavision". In our cohort, 64,5% of patient data was collected by "carevue" system and 33,87% with "metavision" system. Data from these two critical care system were merged in MIMIC III v1.4 except for "luid input which remains divided in 2 tables : inputs for patients monitored with the Carevue system are stored in INPUTEVENTS_CV, whereas inputs for patients monitored with the Meta vision system are stored in INPUTEVENTS_MV. Fluid output is stored OUTPUTEVENTS table. From these tables we can extract mean fluid balance at different times. The mean fluid balance was 111.34 (73.15) , 48.34 (55.94), 34.94 (50.83), 24.25 (48.79), 27.42 (38.92), 27.73 (61.59) and 26.36 (55.66) ml/kg at 24, 48, 72, 96, 120, 144 and 168 hours respectively (Table 2).
Table 3: Bacteria in the dataset.
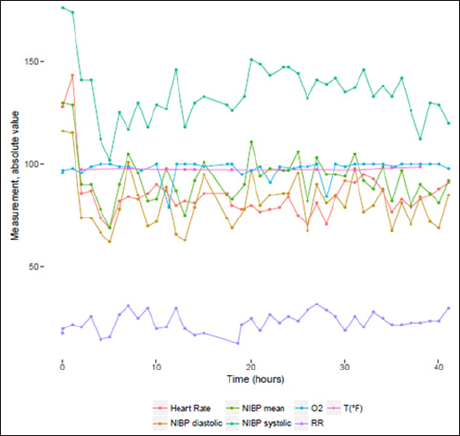
Charted observations are stored in tables CHARTEVENTS. It is by far, the heaviest and largest table in MIMIC III v1.4 database. It contains 330,712,483 rows. It contains all the charted data available for a patient. During their ICU stay, the primary repository of a patient s information is their electronic chart. The electronic chart displays patients' routine vital signs and any additional information relevant to their care: ventilator settings, laboratory values, code status, mental status, and so on. As a result, the bulk of information about a patient's stay is contained in CHARTEVENTS. CHARTEVENTS data could be classified into 21 different categories. Each of these categories contains multiple types of variables as shown in (Table 3). Among the 124 patients, we have a median number of 5402 [2353; 13530] charted observations per patient. For instance, for patient n°77661, we can find 1682 different measurements. A sample of the available measurement is provided in (Figure 1).
Figure 1: Monitoring values for admission id n°116460.
a. NIBP systolic , Non Invasive Blood Pressure systolic,
b. NIBP diastolic, Non Invasive Blood Pressure diastolic,
c. NIBP mean , Non Invasive Blood Pressure mean,
d. RR , Respiratory Rate,
e. O2 , O2 saturation pulseoxymetry,
f. T (°F), Temperature Fahrenheit.
Pharmacological treatments are stored in PRESCRIPTIONS table. Regarding our cohort, 20, 531 prescriptions have been made for the entire population (107.5 [69;196.8]) prescriptions/ patient). The most widely prescribed drug. Regarding antibiotics prescription, the following drugs have been used. Prescription table contains prescribed medications. It does not guarantee any form of administration. On the other hand, the input events tables contain information on intravenously administered medication, and will tell you the exact amount administered. However, it is not possible to know if oral drugs are truly administered after being prescribed.
Laboratory measurements are contained in LABEVENTS table which contains 27,854,055 rows. This table contains all laboratory measurements for a given patient, including outpatient data. Each row associated with one ITEMID, that could be identified from D_LABITEMS table. A median number of 699.5 [322;1308] laboratory measurement are available for each patient; 3 types of laboratory measurements are available ; biochemsitry, hematology and blood gas. On the other side, microbiological data are stored in MICROBIOLOGYEVENTS table. This table stores 631,726 rows. The median number of rows is 21 [8.75;39] for our biliary septic shock cohort. The most frequent germ found was E. Coli in 654 measure accounting for 23.41% of specimens, then K. pneumoniae and Staph aureus coag + in 171 (6.12%) and 127 (4.55%) respectively. Specimens were sterile in 46.53% of cases. (Table 4). Specimens mostly collected were blood culture, urine then sputum in 1363 (48.78%), 283 (10.13%) and 250 (8.95%) patients respectively (Table 5).
Table 4: Origin of specimens.
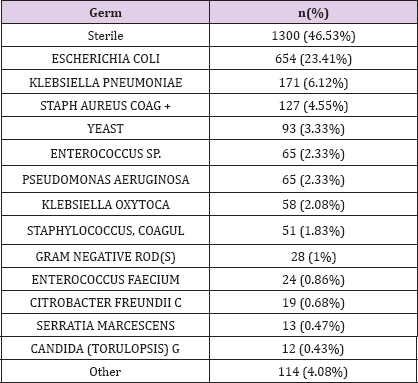
Table 5: Elements from CHARTEVENTS file.
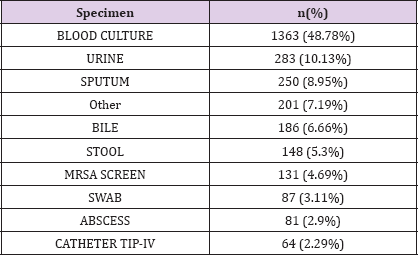
Table 6: Ten most frequent ICD9-9 code procedures.
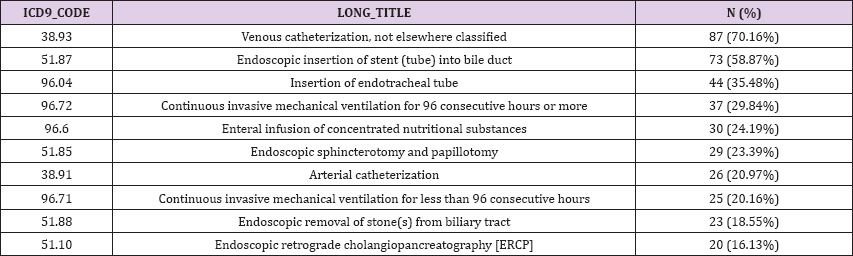
The PROCEDURES_ICD table contains ICD procedures for patients, most notably ICD-9 procedures. The ICD codes are generated for billing purposes at the end of the hospital stay. Regarding cholangitis patient dataset, the moste frequent reported ICD9 procedures were « venous catherization, not elsewhere classified » (n = 87, 70,16%), « endoscopic insertion of stent (tube) into bile duct » (n=73/124 , 58.87%) and « insertion of endotracheal tube » (n=44 , 35.48%) (Table 6).
In this study, we provided with a description of MIMIC III v1.4 database through the descriptive epidemiology of a biliary septic shock cohort. We identified from this database 124 hospitalizations with a mean patient age of 70, 75 (12,57). We presented for that cohort a glimpse of available data by describing some baseline characteristics, fluid balance, length of hospital stay and mortality, laboratory and microbiology, and vital sign measurements. MIMIC database has already been analyzed for clinical investigation in the field of hepato-gastroenterology. For example, Aboelsoud [12] has shown that hospital mortality in patient with acute pancreatitis is lower with the use of lactated ringers compared to isotonic saline; the same author Aboelsoud [13] that among 565 patients diagnosed with hypoxis hepatitis, all-cause hospital mortality was 44.1%. On multivariate analysis, older age, higher SAPS-II, higher INR, higher bilirubin, higher LDH, acute kidney injury and the need for vasopressors were independently associated with mortality.
In another study conducted by Dan-Qin Sun et al. [14], authors used MIMIC III database to develop a specific prognostic score for critically ill cirrhotic patients with acute kidney injury that they called the acute kidney injury - Chronic Liver Failure - Sequential Organ Failure- Assessment score (AKI-CLIF-SOFA) with a valuable discriminative ability compared to other well established scores in critically ill cirrhotic patients with acute kidney injury. In another study, Hu et al. [15] investigated an earlier version of MIMIC III and found that red blood cell width can be used to predict hospital mortality for acute pancreatitis patients admitted to intensive care unit. In other medical specialities, Zhang [16] investigated the relation between do-no-ressucite order and risk of death. Authors included a total of 17,168 subjects and found that patients with do-not-ressucitate order were more likely to die during hospital stay than patients without (42.0 vs. 11.0%; p < 0.001) with mild to moderate severity of illness .They also found that the impact of do-not-ressucitate order was not so prominent in patients with severe illness. In another study, Li et al. [17] analyzed 9000 patients and more than 500,000 records of central venous from MIMIC III database in order to evaluate the association between mean central venous pressure and 28-day mortality after intensive care unit admission.
Authors found that the highest quartile of mean central venous pressure [17.4 (4.1)mmHg] was associated with a 33.6% (95% CI 1.12-1.60) higher adjusted risk of death compared to the lowest quartile [7.4 (1.9) mmHg]. In another study, Yanfei Shen et al. [18] investigated whether negative fluid balance and restricted fluid intake were associated with an improved outcome in critically ill patients. Authors analyzed 2068 patients and found that negative fluid balance were associated with decreased hospital mortality Input and output data are useful information when studying patient in intensive care unit and have also been investigated in multiple other studies. For instance, Acheampong et al. [19] have shown that a persistence of a positive daily fluid balance over time was associated with a higher mortality rate in septic patients (HR = 1.01[1.007-1.02] per ml/kg increase). MIMIC III database is a unique database for multiple reasons; To our knowledge, this is the unique freely accessible critical care database of its kind. Secondly, it spans more than a decade. Data are prospectively collected from 2001 to 2012. MIMIC III database is intended to be accessible to a large number or clinical researchers from a diverse specialty background In order to use it, researchers are required to formally request access via a process documented online on the MIMIC website.
In order to facilitate collaboration and prevent duplicate work, a public code repository has been created to encourage researchers to develop and share code collectively: https://github.com/ MIT-LCP/mimic-code. Limitations of MIMIC are that it requires computational skills. Data of patients need to be extracted using structure query language (SQL), an open source administration and development platform for Postgre SQL which could be challenging for researchers with little or no background of database management. Also, even though data are collected prospectively, studies using MIMIC III are retrospective studies and might before be prone to its inherent bias. Finally, a last limitation is that data come from a unique tertiary care center in the United States that might not be representative of intensive care units in other hospitals or other countries. Nevertheless, MIMIC III database represents a new opportunity for clinical research in hepatogastreonterology and other medical specialties. In this new era of data sharing, publicly available large hospital databases such as MIMIC III will be more and more available and will constitute promising alternative for primary clinical research and validation studies.


