Impact Factor : 0.548
- NLM ID: 101723284
- OCoLC: 999826537
- LCCN: 2017202541
B Pinarbasi Simsek1*, D Onel2, M Gulluoglu3, F Akyuz1, H Issever4, S Badur2, K Demir1, F Besisik1, S Ozdil1, G Boztas1, Z Mungan1 and S Kaymakoglu1
Received: December 20, 2017; Published: January 08, 2018
Corresponding author: Binnur Pinarbasi Simsek, MD, Millet Cad.34390 Capa, Istanbul, Turkey
DOI: 10.26717/BJSTR.2018.02.000637
Purpose: We have analyzed the relationships between quantitative HDVRNA, HBsAg and HBV DNA concentrations and biochemical activity, histological activity index and fibrosis stage in 46 patients with chronic hepatitis D (CHD) and 87 with chronic hepatitis B (CHB), including 33 positive for HBeAg.
Methods: All patients were naïve to treatment. HBsAg concentrations were measured using Abbott Architect kits and HDVRNA by real time PCR.
Results and Conclusions: HBsAg and HDVRNA concentrations were significantly and positively correlated (p=0.043). HDV viremia varied greatly, with HDVRNA titer not correlated with biochemical or histological activity or fibrosis stage. Serum transaminase concentrations and histological findings did not differ between HBVDNA-negative and -positive patients with CHD (p>0.05), and HBsAg concentrations did not differ significantly in HBVDNA-positive and -negative patients (p>0.05). HBsAg levels were significantly higher in HBeAg-positive patients with CHD than in those with HBeAg-negative CHB but were similar in patients with CHD and HBeAg-negative CHB. In CHB patients, there was a positive correlation between serum HBVDNA and HBsAg concentrations (p=0.002). As a result, histological activity and fibrosis stage were significantly higher in CHD than CHB patients, but were not influenced by the levels of HDV and HBV viremia in CHD. HBsAg titers were highest in patients with HBeAg-positive CHB, followed by patients with CHD, and lowest in HBeAg-negative CHB patients.
Keywords: Quantitative hepatitis B surface antigen; quantitative HDV RNA; viral interactions; hepatitis B virus; hepatitis delta virus
Chronic hepatitis delta (CHD) is a serious cause of viral hepatitis throughout the world, with high morbidity and mortality rates [1-3]. Estimates suggest that 15-20 million people worldwide have been infected by hepatitis delta virus (HDV). In Turkey, HDV remains a serious health problem, with epidemiologic studies in East and Southwest Anatolia regions showing delta super infection in 9-58% of HBV infected individuals [4]. HDV was first isolated in 1977 from the hepatocytes of hepatitis B virus (HBV) surface antigen carriers (HBsAg) as a nuclear antigen that differed from HBsAg, core (HBcAg) and e (HBeAg) antigens [5,6]. HDV is a defective virus, requiring the HBV envelope proteins, but can replicate within cells in the absence of HBV [7]. HBsAg, however, is required not only for HDV infectivity [8].but also for its packing and extracellular release [9]. Treatment of CHD is complicated by features of HDV. For example, many other viruses encode the enzymes related to their replication and development. Thus, viral replication can be inhibited by inhibiting the synthesis or activity of these enzymes. HDV, however, encodes only HDV RNA-dependent ribosome, with other enzymes involved in viral replication being cellular enzymes. Treatment of CHD should be related to the structure and life cycle of HDV and its relationships with HBV.
Serum HBsAg concentrations are indirect indicators of intrahepatic covalently closed circular (ccc)-DNA concentrations. Since HDV requires HBsAg for virion formation, CHD may be treated by eliminating HBsAg. Administration of the nucleoside analog (NA) clevudine to woodchucks chronically infected with HDV reduced HDV RNA synthesis, with viremia controlled through decreased cccDNA formation [10], thus confirming the importance of HBsAg concentration to HDV viremia. We therefore investigated the relationships between quantitative serum HBsAg and HDV RNA levels and the histopathology and biochemical characteristics of CHD patients, as well as comparing their HBV DNA and HBsAg concentrations.
We assessed 46 patients with CHD and 87 with chronic hepatitis B (CHB), including 33 who were HBeAg positive and 54 who were HBeAg negative; all patients were treated and followed up at our outpatient clinic between 2005 and 2009. Patients were included in the study if they were > 18 years of age, were positive for HBsAg and HBV-DNA (<2000 IU/mL) for ≥ 6 months or were positive for anti-HDV antibody and HDV-RNA for ≥ 6 months, and had elevated (≥2-fold) serum alanine aminotransferase (ALT) concentrations for at least 6 months. In addition, all patients had liver biopsies, obtained on the same day or within the same week that biochemical tests were performed, showing results consistent with chronic hepatitis. Patients with chronic delta hepatitis were excluded if they had any history of antiviral drug use; had any concomitant autoimmune, metabolic or viral chronic liver disease or co-infections, as shown by positivity for autoantibody, anti-HAV IgM, anti-HCV, or anti-HIV. Patients were also excluded if they had decompensate liver cirrhosis (Child B and C stages) or could not undergo liver biopsy; if they had substance addiction and/or alcohol use (> 20 g/day for females,> 30 g/day for males); if they had a concomitant malignancy, including heap to cellular carcinoma; if they had a history of organ transplantation; or if they did not sign the informed consent form. Written informed consent was obtained from each subject following a detailed explanation of the objectives and protocol of the study which was conducted in accordance with the ethical principles stated in the “Declaration of Helsinki” and approved by the institutional ethics committee.
Blood samples were taken from each patient on the day of liver biopsy; at least 0.5 ml of serum was stored in Eppendorf tubes (Eppendorf, Italia srl, Milano, Italy) at -80°C until assayed.
Serum HBsAg concentrations were quantified using an ARCHITECT i2000 SR device and ARCHITECT HBsAg (Abbott Diagnostics, Wiesbaden, Germany) kits. This chemiluminescence microparticle immunoassay (CMIA) of HBsAg in serum and plasma had a confirmation limit of 0.05-250 IU/ml. Samples with HBsAg concentrations higher than 250 IU/ml were diluted 1:500 and, if necessary, 1:1000, as described in the ARCHITECT HBsAg kit protocol [11].
DNA was isolated from the serum samples stored at -80°C using QIAamp DNA mini kits (QIAGEN, Hilden, Germany). HBV DNA was measured using HBV Rotor Gene (HBVRG) PCR kits and Real Time PCR. A 110 bp fragment was amplified using the primers, 5’-GACCACCAAATGCCCCTAT-3’ (forward) and 5’-CCRAGAYYGAGATCTTCTGCGAC-3’ (reverse), which included R and Y markers. Amplification reactions were performed in 30 μl HBV RG master mix (tampon, dNTP, primer, probe and enzyme) and 50 μl reaction mixtures containing 20 μl templates DNA. The amplification protocol consisted of an initial denaturation at 95°C for 10 min, followed by 45 cycles of denaturation at 95°C for 15 seconds, annealing at 55°C for 30 seconds, and extension at 72°C for 15 seconds in a Rotor Gene 2000 device [12]. The copy limit of the test was 26 copies/ml. To prevent contamination, all pupating procedures were performed in laminar flow cabinets and separated chambers and using filtered pipette tips [13].
RNA was isolated from the serum samples stored at -80°C using QIAMP Viral RNA mini kits (QIAGEN, Hilden, Germany). Viral RNA was transcribed to cDNA using the Primer Design Precision Reverse Transcription kit. Each reaction mixture of 20 μl contained 10 μl RNA and 10 μl of cDNA master mixture (dNTP, MMLV 5X buffer solution, MMLV enzyme, and primer mixture) and was incubated for 1 hour at 42°C. The cDNAs were stored at -20°C until PCR was performed. HDV RNA was measured using Real-Time PCR quantification kits for HDV (Primer Design, Southampton, Hants, UK), which contained primers and probes showing 100% homology with all reference sequences in the phylogenetic tree of HDV.HDV RNA PCR was performed using nested PCR [14], with the first step using the primers 5’-GCCCAGGTCGGACCGCGAGGAGGT-3’ and 5’-ACAAGGAGAGGCAGGATCACCGAC-3’ and the second step using the primers 5’-GAGATGCCATGCCGACCCGAAGAG-3’ and 5’ GAAGGAAGGCCCTCGAGAACAAGA-3’. The PCR products were electrophoresis in 2% agarose gels, with the presence of a 405 bp band defined as positive for HDV RNA. The cut-off value for HDV RNA was 100 copies/mL. Primer and probe mixtures were prepared according to the Taqman principle. During replication, the forward primer for pathogen DNA/cDNA was hybridized with the reverse primers. The Taqman system utilized probes with a reporter fluoro chrome (FAM) at the 5’ end and an inhibiting fluoro chrome (TAMRA) at the 3’ end; these probes bound to the region between the primer binding regions on single stranded target molecules. As hybridization between the probe and targeted molecule continued, the signal from FAM was inhibited by TAMRA.
When primer elongation, initiated after the binding of primers to the targeted nucleic acid, reached the probe binding point, the probe was degraded from the 5’ end by the 5’-3’ nuclease activity of the Taq DNA polymerase. This freed the reporter fluoro chrome and initiated signaling, with the degree of signaling proportional to the amount of amplicon produced during each cycle [12]. The kit included a positive control, in which the concentration of the target sequence was 2x107 copies/ ml. This sample was serially diluted to 2x106 copies/ml, 2x105 copies/ml, 2x104 copies/ml, and 2x103 copies/ml to obtain standard curves for the quantitation of HDV RNA in the serum samples.
All biopsy specimens were evaluated by a single specialist pathologist (M.G.), using the modified Knodell Ishak Scoring System for staging and grading [15]. Stage 3 and/or 4 (presence of bridging fibrosis) was defined as severe fibrosis, and stage 5-6 was defined as cirrhosis. For histological activity indexing, samples with 0-7 points were grouped as having minimal/mild activity and those with 8-18 points were grouped as having moderate-severe activity [16].
All statistical analyses were performed using SPSS 11 version (SPSS Inc, Chicago, IL, USA). Virologic parameters, including HBsAg, HBV DNA and HDV RNA concentrations were converted to log10 using Microsoft Office Excel 2007. Results were expressed as mean ± SD or, when appropriate, as medians. Independent groups were compared using t-tests, continuous variables using the Mann- Whitney U-test, and categorical variables using Pearson’s χ2 test and Pearson correlation coefficient. One-way ANOVA was used for comparisons of more than two groups. ROC curves were used to compare diagnostic results of continuous variables in two groups. P < 0.05 was defined as statistically significant.
We assessed a total of 133 patients, of mean age 37.2 ± 11.8 years (range, 17- 63 years), including 45(34%) females and 88 (66%) males. Of the 133 patients, 46(34.6%), [12(26%) females and 34 (74%) males], had CHD, and 87 (65.4%), [33 (38%) females and 54 (62%) males], had CHB. Of the 46 patients with CHD, 34 (74%) were precirrhotic and 12 (26%) were cirrhotic. Of the 87 patients with CHB, 33 (38%) were positive and 54 (62%) were negative for HBeAg, with 80 (92%) being precirrhotic and 7 (8%) cirrhotic. The demographic and biochemical characteristics of these patients are summarized in Tables 1. We found that the thrombolytic count was significantly lower (p= 0.001), while the GGT (p= 0.001) and γ-globulin (p=0.001) concentrations and INR (p= 0.04) were significantly higher, in patients with CHD than in those with CHB. When we assessed the 46 patients with CHD, we found that mean serum albumin concentration and thrombolytic counts were lower, and that INR and concentrations of AST, GGT, and total and direct bilirubin values were higher, in the 12 patients with than in the 34 without cirrhosis. In analyzing the factors predictive of cirrhosis in CHD patients, we found that low serum albumin concentration, low thrombolytic count, PT elongation, increased INR, increased serum total and direct bilirubin concentrations, and increased AST and GGT were predictive. ROC curves showed that an AST upper limit of 60 IU had a sensitivity and specificity of 70% and that a GGT upper limit of 70 IU had a sensitivity and specificity of 81% in predicting cirrhosis (Figure 1).
Figure 1: Parameters predictive of liver cirrhosis in patients with chronic hepatitis delta infection.
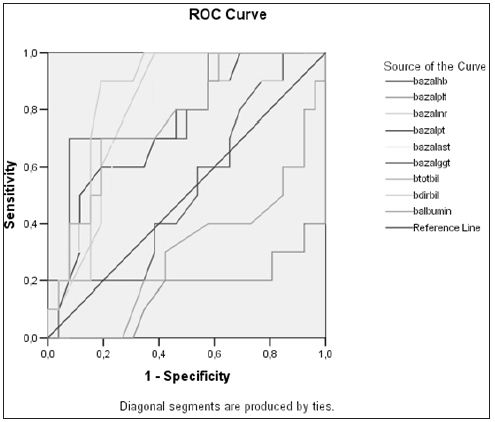
Table 1: Demographic and biochemical characteristics of patients with Chronic Hepatitis B and Chronic Hepatitis D.
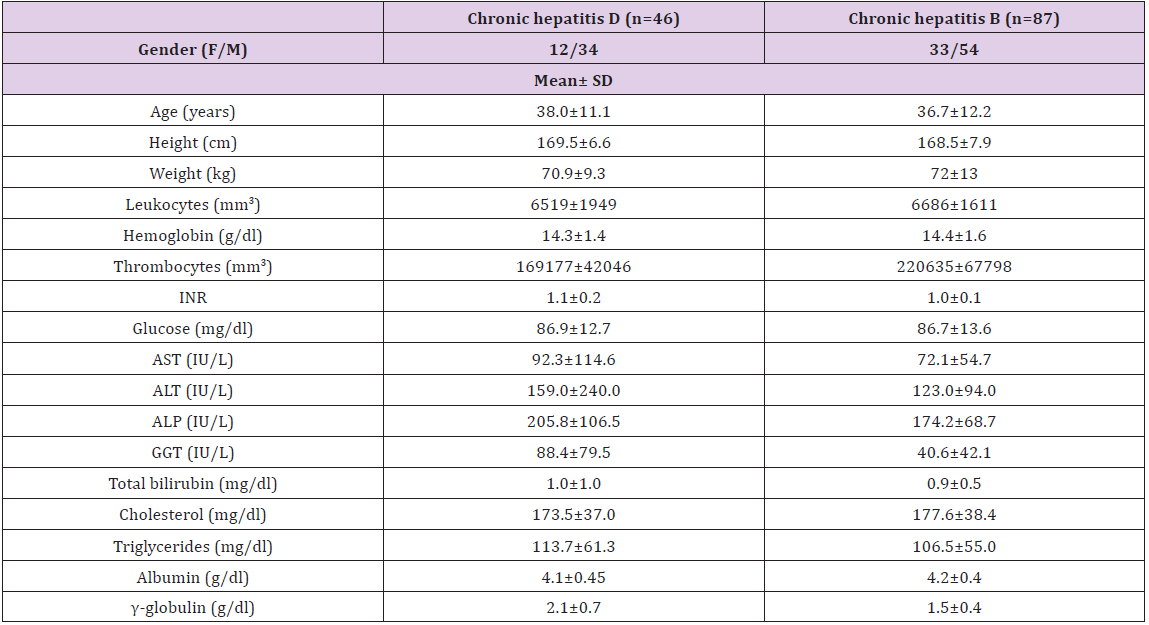
The mean log HDV RNA concentration in patients with CHD was 4.0 ± 0.8 and the mean log HBsAg concentration was 4.7±0.4. We observed a significant positive correlation between HDV RNA and HBsAg concentrations (p = 0.043 and r=0.3). There was no correlation, however, between serum HDV RNA level and serum AST (p=0.39 and r=0.13) or ALT (p=0.30 and r=0.15). Moreover, there was no correlation between HDV RNA concentrations and histological activity (p=0.99, r=0.01), with the mean log HDV RNA concentrations in the groups with minimal (HAI index 0-7 points) and moderate-severe (HAI 8-18 points) activity of 4.0±0.6 and 4.0±0.9, respectively. Likewise, there was no correlation between serum HDV RNA concentration and fibrosis stage (p=0.37, r=-0.13). When we divided patients with CHD into pre-cirrhotic and cirrhotic groups, we found that they had similar virologic parameters (Figure 2). Of the 46 patients with CHD, 17 were negative for serum HBV DNA (< 26 copies/ml), whereas the other 29 were positive (Table 2). The mean serum HDV RNA concentrations were 46563±101239 copies/ml (range 133–359044 copies/ml) and 32036±42556 copies/ml (range 460–154811 copies/ml), respectively, with logarithmic means of 3.8± 0.9 (range 2.1–5.) and 4.1±0.7 (range 2.7– 5.2), respectively (p>0.05). The mean serum HBsAg concentrations were 55416±42608 IU (range 400 – 171030 IU) and 66775±25324 IU (range 18415–116265 IU), respectively, with logarithmic means of 4.6±0.6 (range 2.6-5.3) and 4.8±0.2 (range 4.1–5.0), respectively (p>0.05).The mean HBV DNA concentration in the 29 CHD patients positive for HBV DNA was 72064638 ± 382515197 copies/ml (range 59-2060821400 copies/ml), with a logarithmic mean of 3.9±1.8 (range 1.8-9.3). There was no correlation between serum HBV DNA and HBsAg titers in these CHD patients positive for HBV DNA (p= 0.61, r=-0.98) or between HBV DNA concentrations and HAI (p=0.53, r=-0.8). The fibrosis scores of CHD patients positive and negative for HBV DNA were similar (3.5±1.7 vs. 3.3±1.6, p= 0.6). Similarly, HDV RNA concentration was not correlated with necroinflammatory activity or fibrosis score. Patients with CHD whose HBV-DNA’s positive were subdivided into two (as (<10000 copy/ml and ≥ 10000 copy/ml) and investigated in terms of basal ALT, necroinflammatory activity and fibrosis. Although each of three parameters was higher in the group with HBVDNA level of ≥ 10000 copy/ml, this difference did not suggest any significance when statistically evaluated (p > 0.05). When we assessed HDVRNA and HBV DNA concentrations in patient’s positive for both, we found that low HBV viremia (< 4 log) was usually accompanied by high HDV viremia (Figure 3).
Figure 2: Comparison of viral markers in patients with chronic hepatitis delta.
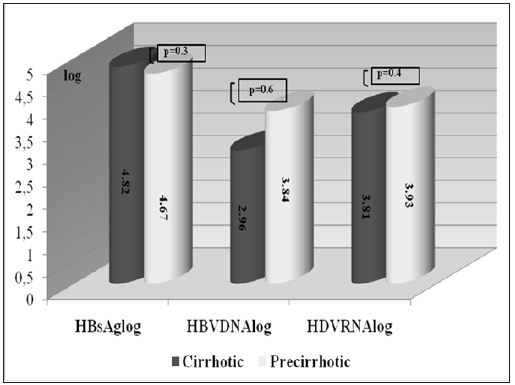
Figure 3: Distribution of serum HDV RNA and HBV DNA concentrations in patients with chronic hepatitis delta (n= 29, when HBV DNA negative cases were excluded).
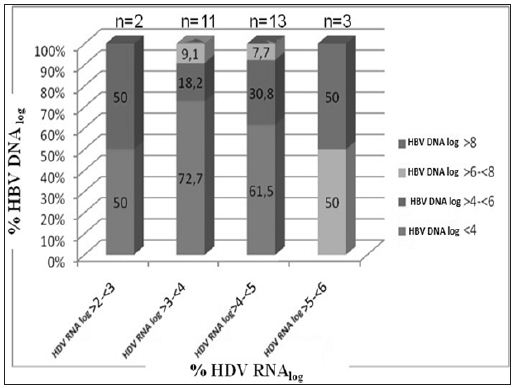
Table 2: Virologic and histopathologic characteristics of patients with chronic hepatitis D and positive or negative for HBV DNA.
We found that mean HAI was higher in CHD patients negative than positive for HBV DNA, but the difference was not statistically significant (9.1±2.8 vs. 8.2±3.2, p=0.17). Of the 17 CHD patients negative for HBV DNA, 5(29%) had HAIs of 0-7 points and 12(71%) had HAIs of 8-18 points. In comparison, of the 29 CHD patients positive for HBV DNA, 12(41%) had HAIs of 0-7 points and 17(59%) had HAIs of 8-18 points (Table 2) and (Figure 3). Serum ALT, AST, ALP and GGT concentrations were similar in CHD patients positive and negative for serum HBV DNA. We found that the mean HBsAg concentration was significantly higher in CHB patients positive than negative for HBeAg (p=0.001), but was similar in CHB patients positive for HBeAg and CHD patients (p = 0.42). Similarly, HBsAg concentrations were significantly higher in patients with CHD than in CHB patients negative for HBeAg (p = 0.006) (Figure 4). In the patients with CHB, we observed a positive and moderate correlation between serum HBV DNA and HBsAg concentrations (p=0.002, r=0.33). When we assessed histological characteristics, we found that mean HAI (8.5±3.0 vs. 6.1±2.8, p=0.001) and mean fibrosis stage (3.4±1.6 vs. 2.3±1.3, p=0.001) were significantly higher in CHD than in CHB patients. Among the 46 patients with CHD, 23(50%) had severe fibrosis (stage 3-4) and 11(24%) were cirrhotic (stage 5-6), with only 12(26%) having stage 1-2. In contrast, of the 87 patients with CHB, 58 had early stage fibrosis (stage 1-2), 22 (25%) had stage 3-4 and only 7(8%) were cirrhotic (stage 5-6).
Figure 4: Virologicmarkers in patients with chronic hepatitis B and chronic hepatitis delta.
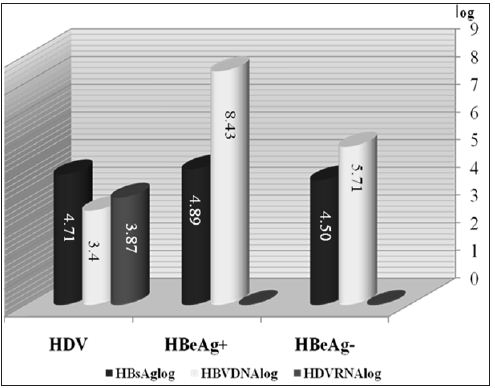
We also compared the demographic, biochemical, virologic and histopathology characteristics of CHB patients positive and negative for HBeAg. We found that the mean ages of these groups were 31.4±11.8 years (range 18-58 years) and 40.1±11.4years (range: 17-63 years), respectively, showing that patients positive for HBeAg were significantly younger (p= 0.001). The mean weight of these patients also differed significantly (67.5±13.6 kg vs. 74.7±12.8 kg, p= 0.01). Median thrombolytic counts were higher in the HBeAg positive than in the HBeAg negative group (237500 vs. 207000, p= 0.02), but there were no other differences in biochemical characteristics. Mean log serum HBV DNA (8.40±1.51 vs. 5.66±1.55, p= 0.001) and HBsAg (4.8±0.4 IU vs. 4.5±0.5 IU, p=0.001) titers were significantly higher in HBeAg positive than in HBeAg negative patients. In contrast, mean HAI (6.1±2.7 vs. 6.08±2.9) and mean fibrosis stage (2.4±1.4 vs. 2.2±1.3) were similar in the two groups.
We have compared the demographic, biochemical, virologic and histopathology characteristics of CHB and CHD patients naïve to treatment; to our knowledge, this is the first such study to compare virologic parameters quantitatively in these patients and to assess the relationships between virologic and histopathology characteristics. Although we observed a correlation between serum HDV RNA and HBsAg concentrations, we observed no relationship between HAI and fibrosis stage. Serum HBsAg concentrations were higher in HBeAg positive than in HBeAg negative CHB patients, but were similar in HBeAg positive CHB and CHD patients. HBV viremia was not detected in 37% of the patients with CHD, whereas 59% of CHD patients positive for HBV had low HBV titers (< 4 log HBV DNA copies/ml). However, there was no inverse correlation between HBV DNA and HDV RNA concentrations. In patients with CHD, biochemical parameters for cirrhosis included increased AST, GGT, γ-globulin, and total bilirubin concentrations, and decreased thrombolytic counts and albumin concentrations. Measurements of HDVRNA by qualitative PCR methods do not indicate the exact magnitude of viremia. However, HDV RNA quantification is difficult. HDV RNA has the appearance of a thick rod, due to 70% base pairing in the inner regions of the HDV RNA. This can decrease PCR efficacy, to as low as 83%, by inhibiting DNA synthesis [17]. Since the sequences of HDV types differ by more than 37% [18], the primers and probes chosen for preserved regions (i.e. ribosome’s) may not be suitable for every type. Furthermore, since there is no international standard or control, calibration of quantification is impossible. New-onset real-time PCR systems are very sensitive [19]. And have been employed in a few studies [16,20,21]. In agreement with our findings, one previous study found no correlation between HDV RNA concentrations and histopathological findings [16]. That study, however, evaluated only patients with CHD, not those with CHB.
Importantly, we found that concomitant HBV viremia did not affect biochemical and histopathological parameters in patients with CHD. In contrast, the progression of chronic liver disease was found to be more rapid and severe in patients with concomitant HDV and HBV infection [22]. Our findings are in agreement with studies showing that biochemical and histological findings in patients with CHD were not related to HBV DNA concentrations [16,23]. Moreover, no correlation was observed between HBV DNA and ALT concentrations in patients with CHD, suggesting that liver damage in these patients was mainly caused by HDV, not HBV; infection [2]. Liver damage in patients with CHD was not affected by HBV confection. In contrast, HDV was found not to have direct cytopathic effects, with hepatocellular damage in CHD being immune-mediated [24]. We observed no correlation between serum transaminase and HDV RNA concentrations. Although ALT concentrations were found to be higher in patient’s positive than negative for HDV RNA [25], a later study, in which HDV RNA was measured semi quantitatively, showed no correlation between ALT and HDV RNA [23]. The correlation we detected between serum HBsAg and HDV RNA titers in patients with CHD suggests that HBsAg may be a key to the treatment of CHD. Antiviral agents that decrease HBsAg titer may therefore suppress HDV RNA.A study on 53 patients with CHD showed that HBsAg and HDV RNA titers decreased in parallel in patients responsive to IFNα-2b, but neither was affected in interferon-unresponsive patients [26]. Our results indicate that serum HBsAg may be a marker of HDV viremia in patients with CHD.
HBV and HDV are thought to occur together, with HDV suppressing HBV viremia. Our findings indicated, however, that decreased HBV DNA did not correlate with increased HDV viremia. Many of our CHD patients were negative for HBV DNA or had low titers (< 4 log copies/ml). The low numbers of CHD patients positive for HBV DNA may have caused a discrepancy between low HDV viremia (≥2 and <3 log) accompanied by somewhat high HBV viremia (≥4 and <6 log) and high HDV (≥5 and <6 log) accompanied by high HBV (≥6 and ≥8 log) viremia. A cross-sectional and longitudinal study of HDVRNA and HBVDNA in 37 CHD patients naive to treatment showed that both viruses had dynamic and complex replication profiles: The cross-sectional evaluation showed that both viruses were replicative in 15 patients (40.5%), HDV alone was replicative in 12 (32.4%), HBV alone was replicative in 6 (16.2%), but both were inactive in 4 (10.8%). Longitudinal evaluation, however, showed that HBV replication increased in the presence of HDV replication [24].
In conclusion, we found that HDV viral load correlated with HBsAg titer, but not with histological, activity, fibrosis stage, or HBV viremia. Our findings indicate that CHD is a more serious disease than CHB.

