ABSTRACT
Background: This study is cross-sectional and consecutive study carried out to find out the sensitivity and specificity of Quick Vue rapid influenza diagnostic kit test with reference to real time RT-PCR. This study was conducted at National Influenza Center, Reference Laboratory, National Public Health Laboratory, Teku, Kathmandu. This study was undertaken in individuals having Influenza like illness visiting National Public Health Laboratory.
Material & Methods: A total of 1683 throat swab specimens, obtained from patients with Influenza like Illness (ILI) at National Influenza Surveillance Network (NISN) sentinel hospitals, were transported to National Influenza Center, maintaining reverse cold chain, within 48 hours. Viral RNA was extracted using QI Amp viral RNA kit. Polymerase Chain Reaction assay (PCR) was performed following CDC Real-time rRT PCR protocol for detection and characterization of the influenza viruses including pandemic influenza virus A (H1N1) pdm 09. Samples were tested with quick view rapid test.
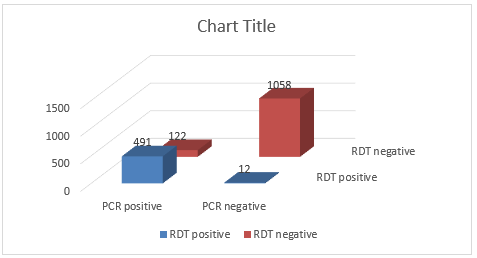
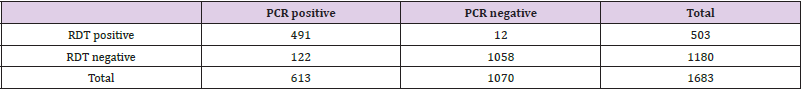
Results: A total 1683 samples were processed for influenza testing by RDT Quick Vue Influenza A + B and PCR simultaneously. Among 1683 samples 491 were positive by rapid testing and PCR as well. Whereas 122 were positive only by PCR. And out of 1520 negative samples, 1508 were negative by rapid testing & PCR as well and 12 were negative only by the PCR.
Conclusion: In our study it shows over all good positive predictive and negative predictive value help to provide about influenza in quickly as point of care testing that help in the clinical management of the patient in the outbreak with less trained manpower and cost-effective kits. However more data and test need to be done to evaluate the clinical performance of RDT for influenza diagnosis on extrapolated to other age group & clinical information for specific symptoms and severity is require.
Keywords: Influenza Virus; Serodiagnosis; Molecular Characterization; Quick View; Rapid Testing
Introduction
Influenza is a highly contagious viral respiratory infection caused by influenza viruses who’s epidemic and pandemic has resulted in significant mortality and morbidity. It has been reported that annual epidemic of influenza results in an estimated 3 – 5 million cases of severe illness and about 290000 – 650000 deaths globally. This study was aimed to investigate the rapid influenza diagnostic tests that detect influenza viral antigens are used to screen patients with suspected influenza and offer the advantage of providing a timely result, compared with real time PCR for influenza, that can influence clinical decision making with in short time [1]. This test can help to decrease unwanted diagnostic testing, to advice antiviral treatment, to decrease inappropriate use of antibiotics and to reduce the duration of treatment in the emergency department or hospitalization [2-8].
Rapid influenza tests have also been used for public health purposes to investigate suspected influenza outbreaks. Identification of influenza virus infection by rapid tests can facilitate prompt implementation of control measures before confirmatory test results are available from shell vial culture, tissue cell viral culture, or RT-PCR. Rapid influenza tests have also been used for public health purposes to investigate suspected influenza outbreaks. Identification of influenza virus infection by rapid tests can facilitate prompt implementation of control measures before confirmatory test results are available from shell vial culture, tissue cell viral culture, or RT-PCR. A wide range of sensitivities of the rapid influenza tests have been reported, whereas specificities have been reported to be high, compared with reference tests [1,8-12]. In the context of enrolling participants in studies to knowledge of transmission of influenza virus, a rapid influenza diagnostic test was used to screen persons with influenza-like illness in the Nepal during the 2016 influenza seasons. Here, we report our findings that compared the accuracy of the rapid test with that of RT-PCR an investigation of these results.
Methods
Study Design
This study is cross-sectional and consecutive study carried out to find out the sensitivity and specificity of Quick Vue rapid influenza diagnostic kit test with reference to real time RT-PCR. This study was conducted at National Influenza Center, Reference Laboratory, National Public Health Laboratory, Teku, Kathmandu. This study was undertaken in individuals having Influenza like illness visiting National Public Health Laboratory. The numbers of the suspected clinical samples included in the study were sixteen hundred eighty-three. This study was performed in 2016.
Collection of Sample and Transportation
After obtaining Ethical approval, informed consent, patient information, demographic characteristics, clinical history, two throat swabs were collected. According to the WHO (2011), throat swab is a suitable specimen for upper respiratory tract infection, which can easily be collected. Out of two throat swabs collected, one was used for the rapid detection of influenza at the site of sample collection while another was transported to laboratory for viral RNA extraction by using WHO manual for the collection and transportation of clinical sample. Clinical samples which could not be processed on the collection day were refrigerated at 4oC no longer than 4 days and for longer storage than that storage at -70oC were done. The clinical specimens collected were processed according to the standard protocols for detection of influenza virus.
Rapid Antigen Diagnosis of Influenza Virus
The Quick Vue Influenza A+B test involves the extraction of influenza A and B viral antigens. The patient specimen is placed in the extraction reagent tube, during which time the virus particles in the specimen are disrupted, exposing internal viral nucleoproteins. After extraction, the test strip is placed in the extraction reagent tube where nucleoproteins in the specimen will react with the reagents in the test strip. If the extracted specimen contains influenza antigens, a pink-to-red test line along with a blue procedural control line will appear on the test strip indicating a positive result. The test line for Influenza A or B will develop at separate specified locations on the same test strip. If influenza type A or type B antigens are not present, only a blue procedural control line will appear. In case of presence of very low levels of influenza type A or type B antigens, the test kit cannot detect viral antigen and gives result as in negative, a blue control line.
RNA Extraction
The RNA was extracted manually using the QAIamp® Viral RNA mini kit (QIAGEN GmbH, Hilden, Germany) as per the manufacturer’s instructions.
Nucleic Acid Detection of the Influenza Virus
The CDC Real-time RT-PCR protocol for detection and characterization of Influenza includes a panel of oligonucleotide primers and dual-labeled hydrolysis (taqman®) probes to be used in real time RT-PCR assays for the in-vitro qualitative detection and characterization of human Influenza Virus in respiratory specimens and viral cultures. The Influenza A and B primer and probe set was designated for universal detection of type A and type B influenza viruses. Influenza A sub typing primer and probe sets were designed to specially detect contemporary human A/H1, human A/H3, and A/H1N1 pdm09 influenza viruses.
Data Collection and Analysis
Data were collected after the valid results were obtained for each specimen. The data were entered in Microsoft Excel 2013 as well as IBM SPSS Statistics Version 20.
Laboratory Safety and Infection Control
Influenza is air-borne disease. It is very contagious disease and transmitted to the person through the air borne droplet up to 6 feet away. If proper safety method is not used, then it can cause serious health hazard to the laboratory workers. It was mandatory to wear PPE (personal protective equipment) before handling the clinical samples. No laboratory staff handling clinical samples could touch pen, copy/files, computers, door locks, hair, eyes etc. Triple layers packing of the contagious samples of influenza virus was mandatory to prevent the possible spread of virus. Every clinical sample was processed in the safety cabinet. There was a different room for allocation, extraction, master mix and PCR. All the waste materials were discarded only after autoclaving. All the diagnosis was carried out in BSL II laboratory.
Results
A total 1683 samples were processed for influenza testing by RDT Quick Vue Influenza A + B and PCR simultaneously. Among 1683 samples 491 were positive by rapid testing and PCR as well. Whereas 122 were positive only by PCR (Tables 1 & 2). And out of 1520 negative samples, 1508 were negative by rapid testing & PCR as well and 12 were negative only by the PCR (Figure 1).
Discussion
Influenza can be a major public health burden. Both influenza A and influenza B have caused estimated 3 – 5 million cases of severe illness and about 290000 – 650000 deaths globally. The diagnosis of influenza infections includes viral isolation in cell culture, immunofluorescence assays, nucleic acid amplification tests, immunochromatography‐based rapid diagnostic tests, etc. The rapid diagnosis which is relatively simple and easy to perform test than other test and take shorten period to diagnose that help to manage the patient treatment and reduce the overall cost of the treatment. Here we have evaluated rapid influenza test through Quick Vue Influenza A+B test kit in comparison to RT-PCR. The Performance of the rapid test was moderately good, with a sensitivity (80%) and specificity (98%) in comparison with a real time RT-PCR assay. In Janice K. Louie et al study on the Quick Vue test for influenza have found comparable specificities (98%) but lower sensitivities (26%) R. However other previous studies show related to this show comparable sensitivity and specificity R. The discrepancies in the result might be due to the study design and sample size, across the range of Ct value.
Conclusion
In our study it shows overall good positive predictive and negative predictive value help to provide about influenza in quickly as point of care testing that help in the clinical management of the patient in the outbreak with less trained manpower and costeffective kits. However more data and test need to be done to evaluate the clinical performance of RDT for influenza diagnosis on extrapolated to other age group & clinical information for specific symptoms and severity is require [13-15].
Declarations
Author Contribution Statement
a) Bimalesh kumar jha: Conceived and designed the experiments, performed the experiments, wrote paper.
b) Rachana Mehta: Performed the experiment, analyzed the data.
c) Krishna Das Manandhar: Conceived and designed the experiments; Contributed reagents, materials, analysis tools or data; Wrote the paper.
Finding Statement: This research did not receive any specific grant from funding agencies in the public, commercial or non-forprofit organization.
Competing Interest: The authors declare that they have no competing interests.
Availability of Data: All necessary data are included in paper. Remaining data will be provided by corresponding authors on reasonable request.
Acknowledgement
We express our deep gratitude to all laboratory staffs of National Influenza Centre, National public health laboratory, Teku and Department of biotechnology Tribhuvan university, Nepal for their kind support to completing this work. We are also thankful to head of Department who provided us to chance to carry the study.
References
- Korsun N, Angelova S, Trifonova I, Georgieva I, Tzotcheva I, et al. (2019) Predominance of influenza B/Yamagata lineage viruses in Bulgaria during the 2017/2018 season. Epidemiology & Infection 147.
- Scorza FB (2017) Advancing new vaccines against pandemic influenza in low-resource countries. Vaccine. 35(40): 5397-5402.
- Odun-Ayo F, Odaibo G, Olaleye D (2018) Influenza virus A (H1 and H3) and B co-circulation among patient presenting with acute respiratory tract infection in Ibadan, Nigeria. African Health Sciences 18(4): 1134-1143.
- Kawakami C, Yamayoshi S, Akimoto M, Nakamura K, Miura H, et al. (2019) Genetic and antigenic characterisation of influenza A (H3N2) viruses isolated in Yokohama during the 2016/17 and 2017/18 influenza seasons. Eurosurveillance 24(6).
- Wu C, Wang MH, Lu X, Chong KC, He J, et al. (2016) Concurrent epidemics of influenza A/H3N2 and A/H1N1pdm in Southern China: A serial cross-sectional study. Journal of Infection 72(3): 369-376.
- Bragstad K, Nielsen LP, Fomsgaard A (2008) The evolution of human influenza A viruses from 1999 to 2006: A complete genome study. Virology Journal 5(1): 40.
- Daum LT, Shaw MW, Klimov AI, Canas LC, Macias EA, et al. (2005) Influenza A (H3N2) outbreak, Nepal. Emerging infectious diseases 11(8): 1186-1191.
- Pan M, Yang H, Jian J, Kuang Y, Xu J, et al. (2019) Association of meteorological factors with seasonal activity of influenza A subtypes and B lineages in subtropical western China. Epidemiology & Infection, pp. 147.
- Guiomar R, da Silva SP, Conde P, Cristóvão P, Maia AC, et al. (2017) Cross-protection to new drifted influenza A (H3) viruses and prevalence of protective antibodies to seasonal influenza, during 2014 in Portugal. Vaccine 35(16): 2092-2099.
- Segaloff H, Melidou A, Adlhoch C, Pereyaslov D, Robesyn E, et al. (2019) Co-circulation of influenza A (H1N1) pdm09 and influenza A (H3N2) viruses, World Health Organization (WHO) European Region, October 2018 to February 2019. Eurosurveillance 24(9).
- Upadhyay BP, Ghimire P, Tashiro M, Banjara MR (2017) Molecular epidemiology and antigenic characterization of seasonal influenza viruses circulating in Nepal. Journal of Nepal Health Research Council 15(1): 44-50.
- Derrar F, Voirin N, Khanafer N, Izri K, Gradi EA, et al. (2019) Influenza surveillance during the 2009‐2010, 2010‐2011, 2011‐2012, and 2012‐2013 seasons in Algeria. Journal of medical virology 91(8): 1394-1399.
- Sun S, Fu C, Cong J, Li Y, Xie S, et al. (2019) Epidemiological features and trends of influenza incidence in mainland China: a population-based surveillance study from 2005 to 2015. International Journal of Infectious Diseases 89: 12-20.
- Suzuki Y, Taira K, Saito R, Nidaira M, Okano S, et al. (2009) Epidemiologic study of influenza infection in Okinawa, Japan, from 2001 to 2007: changing patterns of seasonality and prevalence of amantadine-resistant influenza A virus. Journal of clinical microbiology 47(3): 623-629.
- Yang L, Chan KH, Suen LK, Chan KP, Wang X, et al. (2015) Impact of the 2009 H1N1 pandemic on age-specific epidemic curves of other respiratory viruses: a comparison of pre-pandemic, pandemic and post-pandemic periods in a subtropical city. PLoS One 10(4): e0125447.

 Review Article
Review Article