ABSTRACT
Introduction: Celiac Disease (CD) is a multisystemic auto-mmune disorder triggered by gluten in HLA genetically predisposed individuals. HLA-DQ genotyping is useful to assess the individual susceptibility to CD but still not sufficient for early diagnosis. Here, we propose HLA-DQA1 and HLA-DQB1 gene typing and exosomes characterization as new tool for CD prevention and diagnosis.
Methods: A Chilean population (n=30) was investigated for SNPs mutations in HLA Class II alleles associated with CD predisposition, using the GenoChip Food Technology. Exosomes have been isolated from donors’ serum by ultracentrifugation and characterized by Western Blotting (for CD63 and CD9) and transmission electron microscopy. Exosomes were also studied for their interleukin-1ra content.
Results: Among the studied population, 45.84, 37.46, and 16.70% were carrying alleles encoding for MHC-DQ heterodimers associated with extremely high, high, and extremely low risk to develop CD. The exosome size decreased significantly (p<0.05) when derived from extremely low CD risk donors (44.58 ± 7.88 nm). In parallel, isolated Exosomes from donors with high and extremely high CD risk showed higher IL-1ra content. The values increase within the extremely high-risk group (108.8 ± 15.91 and 148.8 ± 12.37 pg/mL), as the CD persons were not following any treatment. However, these values were lower (52.50 ± 3.54 and 48.75 ± 6.52 pg/mL) in exosomes isolated from CD patients after treatment.
Conclusion: A relationship between exosomes’ size and IL-1ra content, and genetic susceptibility for CD has been observed, suggesting their possible use as biomarkers for CD prevention and diagnosis.
Keywords: Celiac Disease; HLA-DQA1 and HLA-DQB1; Exosomes; Interleukin- 1ra
Introduction
Health is a state of complete physical, mental, and social wellbeing and not merely the absence of disease or infirmity [1]. Over the past few years, and after the human genome sequence completion, emerging of pharmacogenomics/pharmacogenetics was a very bold step toward personalized medicine intended to develop new and practical pharmacological approaches [2]. In this regard, nutrigenetics has been defined as the effect of genetic variation on dietary response [3]. This genetic heterogeneity is expressed in terms of a combination of specific single nucleotide polymorphisms (SNPs) (at both monogenic and polygenic levels) affecting the nutrient metabolism traits within different ethnic groups [4]. One of the most representative examples of this field is Coeliac disease (CD) associated to SNPs in the human leukocyte antigen (HLA) class II genes. HLA DQ2.5 (encoded by HLA‑DQA1*05 and DQB1*02 alleles) and HLA‑DQ8 proteins (encoded by HLA‑DQA1*03 and DQB1*03:02 alleles) are known as genetic predisposing factors for CD [5].
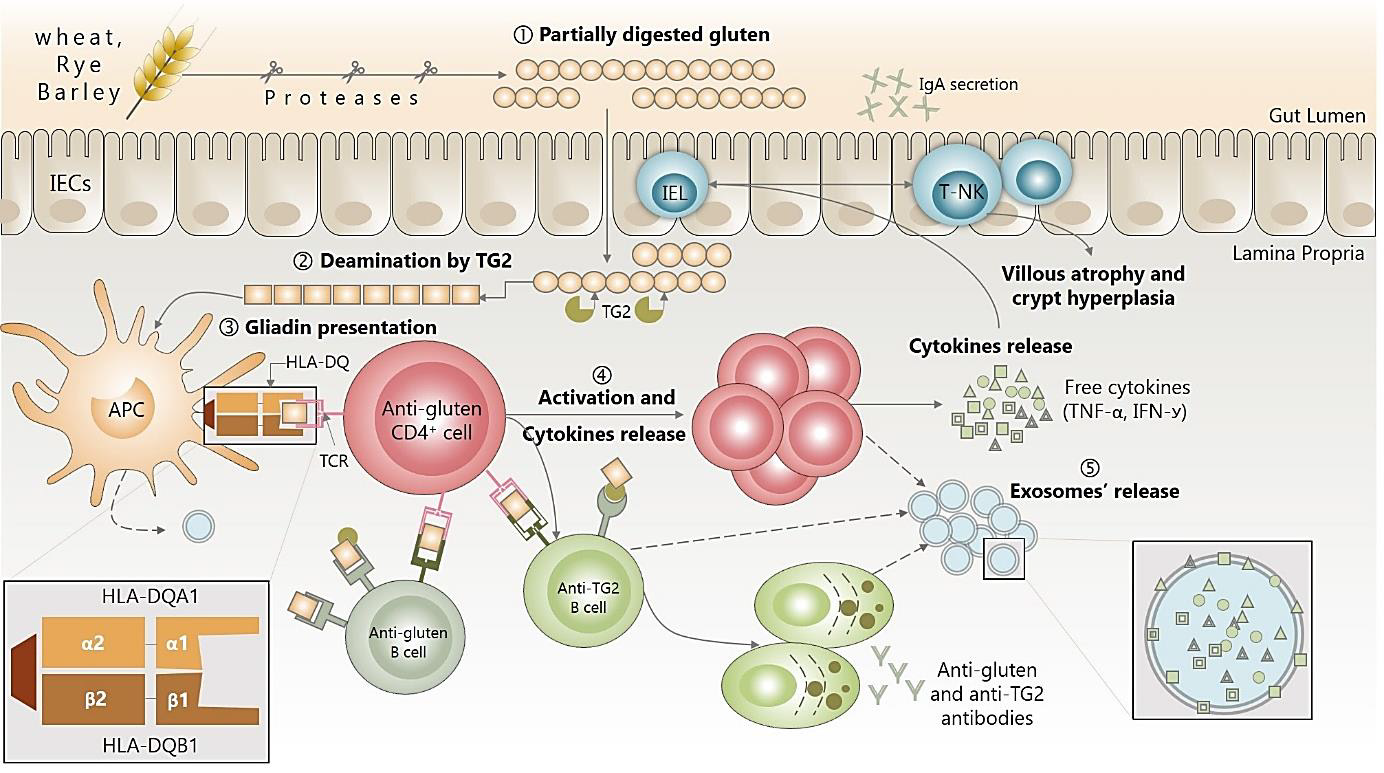
a) Upon digestion, partially digested gluten peptides (mainly gliadin fragments) interact with the epithelium and increase intestinal permeability triggering gliadin translocation into the lamina propria.
b) Once there, a series of events lead to transglutaminase (TG2) release and the deamination of gliadin peptides.
c) Antigen presenting cells (ACPs) recognize then the deaminated gliadin and present it via HLA-DQ receptors (mainly DQ2.5/ DQ8 phenotype) to CD4+T lymphocytes, which constitute the key step in the CD pathogenesis.
d) This triggers the activation and proliferation of both CD4 T and B cells, and proinflammatory cytokines secretion (IFN-ꭹ, TNF-α, IL-6, IL-10), which ultimately lead to metalloproteinases secretion and villous atrophy and crypt hyperplasia (not illustrated, since we are evoking here only the CD onset).
e) Exosomes participate in cell-cell communication during this inflammatory process and their content reflects that of parental cells, which make them a very promising biomarkers of CD inflammation.
CD is a multisystemic dietary, gluten-induced auto-immune disorder of the small intestine mucosa [6]. CD is a worldwide health problem occurring at any age in genetically predisposed individuals [7]. The worldwide incidence of CD is about 1%, which can change due to the interplay of genetic, environmental, and immunological factors [7]. CD is associated with chronic inflammation, diarrhea, mal-absorption, and gastrointestinal malignancies [5,6]. With vast clinical manifestations or even asymptomatic disease, undiagnosed and, therefore, untreated CD is mostly associated with other health complications like gastrointestinal cancers, which constitute a heavy burden for health-care systems and decrease life quality [7,8]. Also, it has been recognized that early treatment with a gluten-free diet could prevent CD-associated complications [7].
The personalized intervention is based on the 4P principles, personalized, predictive, preventative, and participative [2]. The investigation of new preventive and predictive approaches for CD also goes through innovative findings from other scientific fields as cellular and extracellular vehicles (EVs) biology, mainly small EVs called exosomes (EXs). Exosomes (EXs) are virus-sized particles (30-150 nm) delimited by a lipid bilayer and commonly released from endosomal multivesicular bodies [9]. They have emerged as new intercellular signaling pathways and biomolecules delivering agents in different physiological and pathological cases, including inflammatory responses [10-12]. In this regard, we have previously demonstrated that inflammatory molecules (TNF-a, IL-6, IL-1ra, and IL-10) are cell-to-cell transported via EXs (unpublished data). Among the studied cytokines, we found that the anti-inflammatory IL-1ra was the most abundant molecule in EXs. Therefore, we suggest that the exosomes’ size and content in cytokines (more particularly IL-1ra) could be a promising marker of inflammation state associated with auto-immune diseases like CD, which could be a milestone toward the prevention of a heavily under-diagnosed disease worldwide. This ongoing study is primarily structured around
(i) The screening of the frequency of CD-predisposing genetic mutations and
(ii) The Exssome content characterization as new tools for CD prevention and diagnosis to mitigate the associated complications of undiagnosed and/or asymptomatic CD. In this paper, we present our preliminary results regarding exosomes’ size and their IL-1ra content when isolated from individuals carrying different alleles of HLA-DQA1 and HLA-DQB1 class II genes.
Methods
Population and Study Design
The current study is designed as a cross‑sectional, population‑based study to investigate the frequency of CD predisposing HLA‑DQ genotypes and the role of exosomes in CD prevention among Chilean individuals. Healthy males and females aged between 20 and 70 years old were voluntarily recruited during February-June 2019 among students and staff of the Medicine Faculty, University of Santiago (Santiago, Chile). The study population includes subjects with average dietary intake of gluten, subjects who self-reported a gluten sensitivity, subjects who self-instituted and strictly adhered to a gluten-free diet, and CD-diagnosed subjects with and without treatment. Participants were also asked to complete a baseline questionnaire, including information about family history of CD, food intolerances history, and inflammatory disease history. Study details were thoroughly explained to all volunteers, and all participants prior to study enrollment signed a written consent. 8 mL fasting blood samples were collected from each participant’s antecubital vein in EDTA tubes (according to the GenoChip Food kit manufacturer). Blood specimens were freshly used for DNA extraction and then centrifuged at 2000 rpm for 10 min. Plasma specimens and DNA aliquots were collected and stored at -20°C and 4°C, respectively.
Genotyping
Genomic DNA was isolated from the blood samples using a QIAamp mini kit DNA extraction protocol, according to the manufacturer’s instructions (QIAGEN GmbH, Hilden, Germany). Samples were then genotyped using the GenoChip Food kit, following the manufacturer guidelines (Pharmgenomics Gmbf, Mainz, Germany). The GenoChip is permanently integrated onto the bottom of a 1.5 mL reaction tube, spanning only 3x3 mm and pre-coated by sequence-specific oligonucleotide probes. This kit is designed to detect SNPs related to some food intolerances, including but not limited to gluten, alcohol, fructose, lactose, and sucrose. The genotyping process consists of:
Multiplex PCR
The target regions in the HLA-DQA1 and HLA-DQB1 genes, including HLA DQA1*05 (145 bp), HLA DQA1*03 (125 bp), HLA DQB1*03:02 (404 bp), and HLA DQB1*02 (147 bp) were amplified by multiplex polymerase chain reaction (PCR) using the Primus 25 advanced® Thermocycler (PEQLAB Biotechnologie GmbH, Erlangen, Germany). Primers and probes were designed and provided by Pharmgenomics Gmbh. Microarray protocol- After a denaturation step for 2 min at 95°C, 5 μL of each amplification product is transferred to the array tube and bind to the corresponding immobilized oligonucleotide probes for 30 min at 55 °C and 550 rpm. A washing step removes unspecific bound fragments. Next, the Conjugation Mix is added, which binds to the probe-PCR fragment complexes for 5 min at 52°C and 550 rpm. Another washing step removing the unbound Conjugation Mix residues is required. The subsequent adding of the substrate leads to a precipitation reaction at those spots where DNA is bound after 15 min at 21°C and 550 rpm. All incubation steps were performed using the Thriller® Thermoshaker Incubator (PEQLAB Biotechnologie GmbH, Erlangen, Germany). The precipitation pattern is detected with the image reader (Alere Technologies Gmbh, Jena, Germany) and interpreted by the GenoChip Food software.
Exosomes Isolation
Exosomes were isolated by ultracentrifugation according to the protocol described by Theiry ,et al. with slight modifications. Briefly, blood samples were subjected to consecutive centrifugation steps (2,000g and 12,000g) to remove cellular debris and large vesicles (J.S 4.3 rotor and the Avanti J-20XP centrifuge, Beckman Coulter, USA). Exosomes were then pelleted two times with ultracentrifugation at 110,000g for 70 and 120 min (SW32 Ti rotor and the Optima LE-80K ultracentrifuge, Beckman Coulter, USA), separated by a filtration step through a 0.22 μm filter (Sarstedt AG & Co. KG, Nümbrecht, Germany). The supernatant was removed, and pellets were resuspended in 200 μL of PBS and stored at -20°C.
Western Blot Analysis
The Exssome preparation has been treated with RIPA lysis buffer and protease inhibitor cocktail. Samples were vortexed for 15s and incubated for 5 min at room temperature to allow complete lysis. Total Exssomal proteins were measured with Bradford assay (Bio-Rad, Germany). About 20 μg of proteins were loaded, resolved, transferred onto the Immuno-blot PVDF membrane, and subsequently blocked with 5% dry milk in TBS-T (Tris Buffered Saline with 0.05% Tween-20) for 1 hour. The membranes were then incubated with rabbit polyclonal CD63 and CD9 (EXSAB-kit-1, System Biosciences, CA, USA) (1:1000) antibodies overnight at 4 °C. The blots were vigorously washed whit TBS-T and then incubated with Goat anti-rabbit antibody (EXSAB-kit-1) (1:20,000) for 1 hour at room temperature. Membranes were detected with Pierce ECL plus Western blotting substrate (Thermo Scientific), and images were taken on Thermo Scientific™ MYECL™ Imager (Thermo Scientific).
ELISA Assay
According to the manufacturer’s instructions, the quantitative sandwich enzyme immunoassay technique was used to quantify exosomes’ IL-1ra content (R&D Systems, USA).
Transmission Electron Microscopy
The typical size and shape of purified exosomes were determined by TEM analysis (Prof. Dr. Osuna’s laboratory, University of Granada, Spain, and their related protocol was used). In brief, exosomes were pelleted down by ultracentrifugation at 100 000 g for an hour using the FiberliteTM F50L-24 x 1.5 FA rotor and Sorvall WX ultra series ultracentrifuge (Thermo Fisher Scientific, USA). Afterward, the supernatants were discarded while the Exssome pellet was fixed with 50 μL of 2% glutaraldehyde and 2% formaldehyde in cacodylate buffer with 0.1 M Saccharose. Fixed samples were then negatively stained using uranyl acetate and analyzed via TEM (Libra® 120, Carl Zeiss AG, Germany).
Statistical Analysis
Data were analyzed using descriptive statistics: arithmetic means, standard deviation, frequency, and percentage (GraphPad Prism version 6.00, GraphPad Prism Inc, San Diego, California). Data were presented as means ± standard deviation, and a oneway analysis of variance (ANOVA) test was used for multigroup (between CD risk groups) comparison followed by Student– Newman–Keuls test. Results with two-sided p-values of <0.05 were considered statistically significant.
Results
HLA-DQ Genotypes
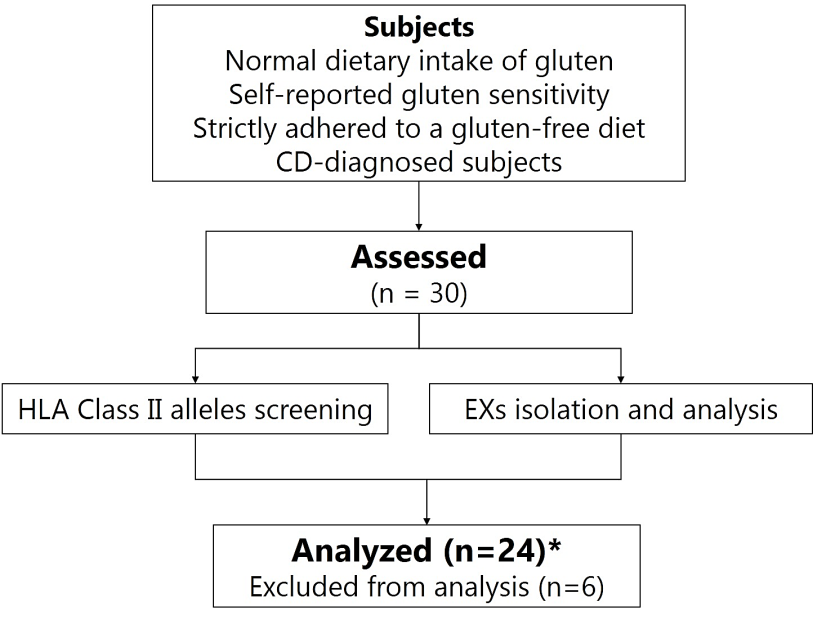
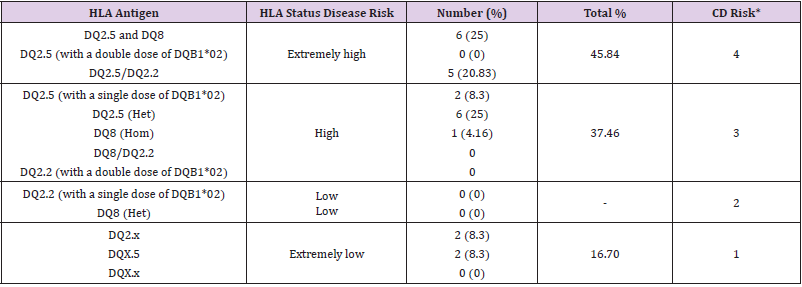
Table 1 shows the frequency of each HLA DQ genotype. As shown in Figure 1, six participants carry alleles that were not detected by the used method. Of the 24 analyzed participants, about 46% carried the CD related HLA DQ molecules, DQ2.5 and DQ8, DQ2.5 (with a double dose of DQB1*02), and DQ2.5/DQ2.2, which are associated with extremely high risk to develop CD. More than 37% carry HLA-DQ2 and HLA-DQ8 haplotypes (DQ2.5 with a single dose of DQB1*02, DQ2.5-heterogenous, DQ8-homozygous, DQ8/DQ2.2, and DQ2.2 with a double dose of DQB1*02), making them highly susceptible to develop CD. 16.6% carried only one of the alleles of the risk HLA DQ2 heterodimers known as “half heterodimer” (HLA DQA1*05 and HLA DQB1*02). Celiac autoimmunity occurs almost exclusively in the presence of DQ molecules. Here, we found that almost all participants were carrying at least one of the DQ heterodimers. However, the calculated general population CD risk was above 3, indicating a high population risk to develop the disease.
Figure 1: Study design. Six participants are carrying alleles that could not be detected by the used method.
Table 1: The HLA-DQA1 and -DQB1 genotype frequencies were observed in the studied population and the associated risk to develop celiac disease.
Note: *CD risk score has been calculated according a 4-point scale (1 = extremely low, 2 = low, 3 = high, 4 = very high)
Exosomes Characterization
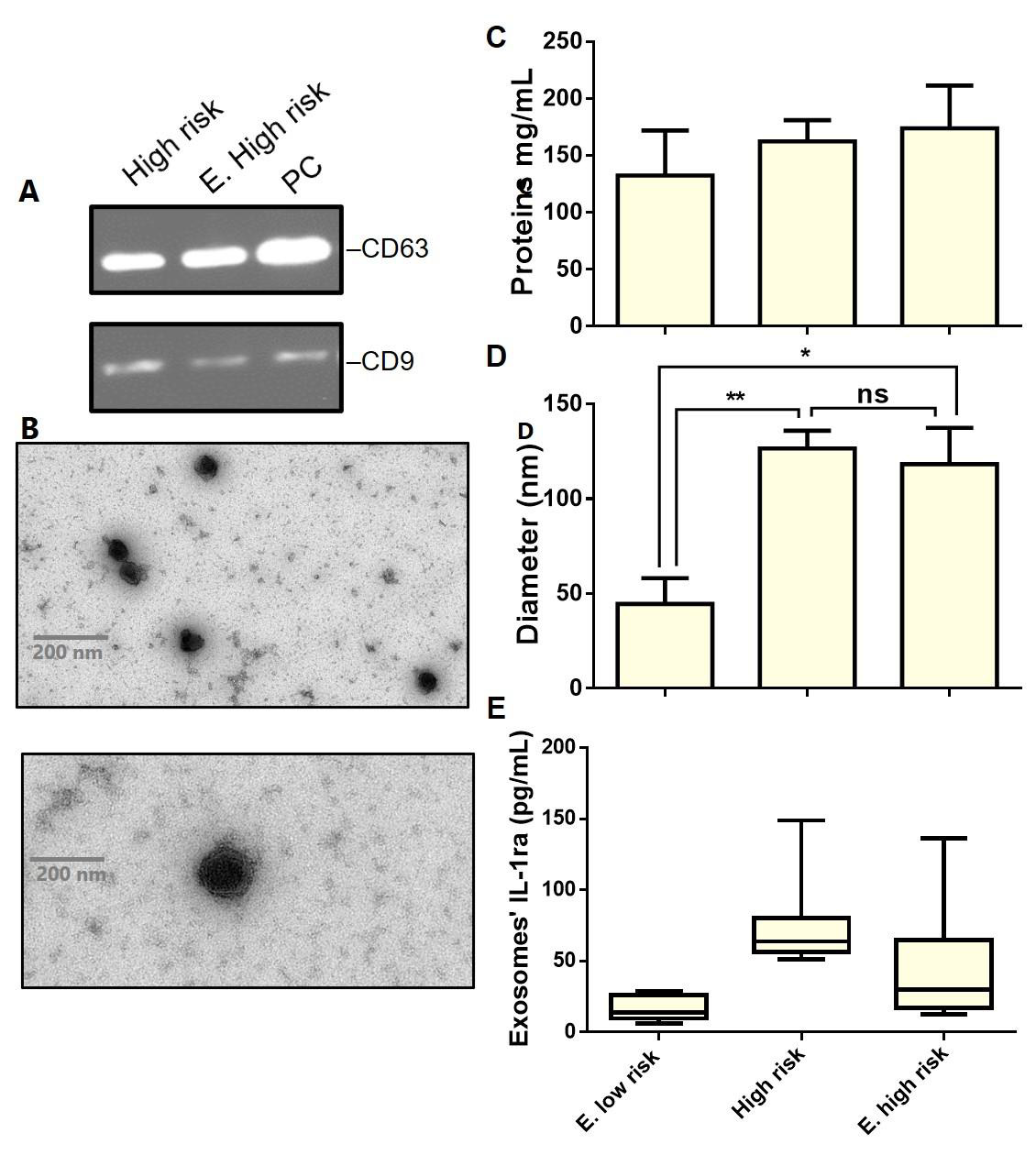
The study of circulating exosomes has emerged as a promising alternative for invasive biopsies, mainly in cancer diagnosis. However, before proceeding with a specific analysis, the characterization of exosomes is a critical step. Nevertheless, not standardized, the identification of isolated particles as exosomes relies on various criteria following a defined working flow. Accordingly, SDS-PAGE and Western blotting firstly characterized the isolated particles; and subsequently, the size, shape, and particles/mL were determined with the help of TEM. SDS-PAGE analysis (data not shown) reveals the presence of intense bands with a molecular weight of about 58 kDa, which could correspond to the Exssome marker CD63, while the slight bands around 25 KDa illustrate the presence of CD9 protein. The presence of CD63 and CD9 as primary Exssome markers was then confirmed on the western blotting membrane (Figure 2a). Furthermore, isolated particles display the expected cup-shaped morphology consistent with pure Exssome preparations (Figure 2b). Despite the genetic background of the donors, almost all isolated particles showed exosomes characteristic size (<150 nm), starting from 23.79 up to 211.22 nm (recorded only once). Taken together, these results demonstrate that the isolated particles certainly belong to the exosome subpopulation.
Exosomes as Potential Biomarkers for CD
All blood samples have been collected in EDTA tubes and used for plasma preparation; the exosomes were subsequently isolated by ultracentrifugation. Figure 2d shows the average size of isolated exosomes according to the donors’ genetic predisposition to CD. EXs’ size differs significantly (p< 0.05) when they are isolated from participants with extremely low (44.58 ± 7.88; n=4) and high (126.6 ± 9.334; n=8) or extremely high (118.3 ± 19.09; n=9) risk for CD. Overall, exosomes’ populations were highly homogenous in individuals of each risk group (low risk: 30.02 ± 5.64 to 57.07 ± 7.88; high risk: 120.03 ± 16.73 to 133.20 ± 22.88; E. high risk: 104.80 ± 15.74 to 131.81 ± 22.94) with a shift towards average bigger size when the risk to develop CD increases. When it comes to the exosomes’ content in IL-1ra, a similar pattern has also been observed (Figure 2e). EXs from donors with high to extremely high CD risk showed high IL-1ra content than those isolated from persons with extremely low CD risk. However, this difference has not been statistically borne due to the high heterogeneity of individuals within each group. Besides normal individuals, the study population also includes CD-diagnosed (treated and not treated) and self-reported intolerance to gluten subjects. EXs’ IL- 1ra was very high (108.8 ± 15.91 to 148.8 ± 12.37 in the E. highrisk group), as the CD-diagnosed subjects were not following any treatment. However, values of IL-1ra (in the same risk group) decrease (52.50 ± 3.54 and 48.75 ± 6.52) in exosomes isolated from persons receiving treatment for CD. The difference in exosomes’ IL- 1ra between treated and non-treated CD patients could open new treatment monitoring perspectives towards a fully personalized intervention. Our preliminary findings suggest that both exosomes seize and their content in IL-1ra could serve as biomarkers for CD diagnosis and prognosis in genetically susceptible subjects.
Figure 2: Characterization of serum-derived exosomes enriched from healthy and CD donors.
a) Immunoblots showing expression levels of CD63 and CD9 in the purified exosomes.
b) Representative TEM images of pure exosomes preparations (Scale bar– 200 nm). The graphs represent c) Proteins concentration
d) Size, and interleukin-1Ra content in exosomes enriched from individuals belonging to each CD risk group. For boxplots the center lines mark the median, box limits indicate minimal and maximal values. Data are represented as means ± standard deviation, *p < 0.05, **p < 0.01, ns: no significant difference. S, sample; PC, positive control.
Exosomes characterization
The study of circulating exosomes has emerged as a promising alternative for invasive biopsies, mainly in cancer diagnosis. However, before proceeding with a specific analysis, the characterization of exosomes is a critical step. Nevertheless, not standardized, the identification of isolated particles as exosomes relies on various criteria following a defined working flow. Accordingly, SDS-PAGE and Western blotting firstly characterized the isolated particles; and subsequently, the size, shape, and particles/mL were determined with the help of TEM. SDS-PAGE analysis (data not shown) reveals the presence of intense bands with a molecular weight of about 58 kDa, which could correspond to the Exssome marker CD63, while the slight bands around 25 KDa illustrate the presence of CD9 protein. The presence of CD63 and CD9 as primary Exssome markers was then confirmed on the western blotting membrane (Figure 2a). Furthermore, isolated particles display the expected cup-shaped morphology consistent with pure Exssome preparations (Figure 2b). Despite the genetic background of the donors, almost all isolated particles showed exosomes characteristic size (<150 nm), starting from 23.79 up to 211.22 nm (recorded only once). Taken together, these results demonstrate that the isolated particles certainly belong to the exosome subpopulation.
Exosomes as Potential Biomarkers for CD
All blood samples have been collected in EDTA tubes and used for plasma preparation; the exosomes were subsequently isolated by ultracentrifugation. (Figure 2d) shows the average size of isolated exosomes according to the donors’ genetic predisposition to CD. EXs’ size differs significantly (p< 0.05) when they are isolated from participants with extremely low (44.58 ± 7.88; n=4) and high (126.6 ± 9.334; n=8) or extremely high (118.3 ± 19.09; n=9) risk for CD. Overall, exosomes’ populations were highly homogenous in individuals of each risk group (low risk: 30.02 ± 5.64 to 57.07 ± 7.88; high risk: 120.03 ± 16.73 to 133.20 ± 22.88; E. high risk: 104.80 ± 15.74 to 131.81 ± 22.94) with a shift towards average bigger size when the risk to develop CD increases. When it comes to the exosomes’ content in IL-1ra, a similar pattern has also been observed (Figure 2e). EXs from donors with high to extremely high CD risk showed high IL-1ra content than those isolated from persons with extremely low CD risk. However, this difference has not been statistically borne due to the high heterogeneity of individuals within each group. Besides normal individuals, the study population also includes CD-diagnosed (treated and not treated) and self-reported intolerance to gluten subjects. EXs’ IL- 1ra was very high (108.8 ± 15.91 to 148.8 ± 12.37 in the E. highrisk group), as the CD-diagnosed subjects were not following any treatment. However, values of IL-1ra (in the same risk group) decrease (52.50 ± 3.54 and 48.75 ± 6.52) in exosomes isolated from persons receiving treatment for CD. The difference in exosomes’ IL- 1ra between treated and non-treated CD patients could open new treatment monitoring perspectives towards a fully personalized intervention. Our preliminary findings suggest that both exosomes seize and their content in IL-1ra could serve as biomarkers for CD diagnosis and prognosis in genetically susceptible subjects.
Discussion
Here, we present the first results regarding exosomes’ size and IL-1ra content when isolated from individuals carrying different alleles of HLA-DQA1 and HLA-DQB1 class II genes. CD is a multifactorial autoimmune disorder triggered by gluten in the small intestine mucosa of genetically susceptible persons. As described in the introduction, the CD is a heavily under-diagnosed disease in which developing complications such as lymphomas and autoimmune disorders is very likely (refractory celiac disease is associated with a 50% risk to develop lymphoma) [7]. However, CD develops rarely in the absence of specific HLA class II haplotypes, making the genetic testing of great importance. So far, only variations in HLA-DQA1 (DQA1*05:01 and DQA1*03) and HLA-DQB1 (DQB1*02, DQB1*03, and DQB1*03:02) alleles have been recognized in the clinical practice of CD [13]. Here, we found that almost all participants are carrying at least one of these alleles leading to genetic predisposition for celiac autoimmunity, which is similar to previous observations on the same population [14-16]. DQ2.5/DQ8 and DQ2.5/DQ2.2 genotypes were the most detected in normal as well as CD participants enrolled in this study. DQ2 homozygosity and DQ2/DQ8 double heterozygosity are associated with the highest CD risk, indicating a very high CD risk in the studied population [16-19]. Despite the CD genetic risk, HLA-DQA1*0501 and HLA-DQB1*02:01 (and their variations: HLADQA1* 05/02 and HLA-DQB1*02) encoding for DQ2 heterodimers (DQ2, DQ2.5, and DQ2.2) were the most dominant CD alleles. In this specific regard, Araya, et al. Have reported that DQ2 was present in 53.9% and 43.9.0% of Chilean CD cases and first-degree relatives [16]. However, Pérez-Bravo, et al. Have earlier found that CD is primarily associated with DQ8 conformation in Chilean children [14]. Therefore, the predominance of DQ2.5/DQ8 and DQ2.5/ DQ2.2 haplotypes were more or less consistent with the previous investigations. Nevertheless, the genotyping method herein used is different, which may explain some discrepancies from previous studies, even with a small sample size [20]. On the other hand, a positive HLA-DQ test means only a genetic predisposition for celiac autoimmunity, whereas the absent HLA-DQ predisposing alleles have an absolute diagnostic value. Along with other markers like anti-transglutaminase (anti-tTGA), HLA-typing would help in the CD screening process to avoid invasive diagnosis methods in the future. Accordingly, more research should also be addressed towards identifying new markers for a complete prediction of CD in genetically susceptible persons.
In this respect, EXs have been widely studied as cancer biomarkers focusing on their protein and nucleic acid content, while little is known about their role(s) in pathologies like CD [21,22]. Starting from our previous observations on EXS’ behavior during inflammation, we hypothesize that EXS’ cytokines alongside their size could serve as a promising marker of CD inflammation. As stated in Figure 3, deaminated gluten peptides presentation (to CD4+T cells) via HLA-DQ (mainly DQ2.5/DQ8) is the crucial step determining the magnitude of the inflammatory response in CD. The cytokines network (IL-6/1ra/10, TNF-α, IFN-ꭹ) that is released directly or packed in EXs, leads to the metalloproteinase secretion and triggers villous atrophy and crypt hyperplasia. It has been recently demonstrated that serum cytokines rise upon gluten ingestion, and this could differentiate CD patients from selfreported gluten sensitivity [22,23]. Indeed, both studies have only focused on pro-inflammatory cytokines release (mainly IL-2), and no information about IL-1Ra or even IL-1 has been evoked. We, for the first time, report that EXS carries different amounts of IL-1Ra depending on genetic predisposition to CD and pathological status (treated or not), suggesting their role(s) in cell-cell communication during CD inflammation. IL-1Ra is a natural anti-inflammatory cytokine that tightly regulates the IL-1 activity creating a balance between innate immunity amplification and uncontrolled inflammation [24,25]. However, IL-1Ra has a short half-life, and a high concentration is required to inhibit IL-1 activity [25]. However, preliminary, these findings may provide clues about using EXS (by cells) as an alternative but stable way for cell-to-cell carrying of vulnerable IL-1Ra, reinforcing the idea to use them as stable biomarkers in the clinic.
Figure 3: Immune dysregulation in CD onset and potential use of exosomes as new trends for diagnosis and screening.
Despite the growing interest in EXS, many secrets related to the releasing mechanism(s), like the variety of EXS size, remain unrevealed. Many microscopic methods are now widely used to study the physical features of EXS as an essential step in the characterization process of pure EXS preparations. To our knowledge, the EXS size itself was not yet considered as a marker since different EXS populations enriched from body fluids or cell cultures originate from cells with different morphological characteristics as it has been accepted. By contrast to that, Sokolova, et al. Have reported that EXS size was comparable when even isolated from cell lines that are morphologically highly heterogeneous [26]. This may indicate that cells release different subpopulations (regarding size) of EXS depending on their physiological status. Here, EXS populations were found to be highly homogenous in individuals belonging to the same CD risk group, while the size was significantly more prominent in favor of high CD risk groups. We should also underline that size distribution depend significantly on the isolation method [27,28]. However, the classical EXS isolation technique of ultracentrifugation herein utilized enriches particles up to a diameter of 250 nm without affecting their shape and integrity [26-28]. Accordingly, CD genetic predisposition seems to affect not only the IL-1Ra in EXS but also their size, suggesting the potential use of these two parameters (alongside HLA-DQ typing) as biomarkers for CD diagnostic and prognostic.
Conclusion
Celiac disease can be diagnosed at any age, and apparently, its worldwide prevalence is increasing due to environmental changes and the absence of non-invasive diagnostic and screening methods. In this paper, we have provided results (yet preliminary) supporting our starting hypothesis and demonstrating that HLA-DQA1/DQB1 gene typing, and exosomes characterization could be useful tools for CD assessment. EXS can be found in almost all body fluids, and their content seems to be more stable than circulating biomarkers due to protection by a lipid bilayer. Hence, developing a system for rapid characterization of active EXS (with specific cargo like the IL-1Ra) could open up possibilities of performing “liquid biopsy” in CD patients to avoid serious complications like lymphoma and colorectal cancer.
Acknowledgment
Not applicable.
Conflict of Interest
The authors declare no conflict of interest.
Funding
This work was supported by grants from Deutscher Akademischer Austauschdienst (DAAD) to Weisshaar M.P.
References
- (2005) Organization, W H Basic documents, World Health Organization, including amendments adopted up to 31 December 2004.
- Bernsen EC, Melanie M Hagleitner, Theodorus W Kouwenberg, Lidwien M Hanff (2020) Pharmacogenomics as a Tool to Limit Acute and Long-Term Adverse Effects of Chemotherapeutics: An Update in Pediatric Oncology. Front Pharmacol 11: 1184.
- Lorenz M, Marco Klemm, Moritz Eidens, Merle Fleischer, Alexander Weise, et al. (2013) Actionable Nutrigenetics for Genetically Based Diseases? A New Critical Path to P4 Medicine 11(2): 167-180.
- Vesnina A (2020) Genes and Eating Preferences, Their Roles in Personalized Nutrition. Genes (Basel) 11(4): 357.
- Caio G, Umberto Volta, Anna Sapone, Daniel A Leffler, Roberto De Giorgio, et al. (2019) Celiac disease: a comprehensive current review. BMC Med 17: 142.
- Kaukinen K (2014) Advances in the treatment of coeliac disease: an immunopathogenic perspective. Nat Rev Gastroenterol Hepatol 11(1): 36-44.
- Tack G J (2010) The spectrum of celiac disease: epidemiology, clinical aspects and treatment. Nat Rev Gastroenterol Hepatol 7(4): 204-213.
- Charlesworth R P (2020) Diagnosing coeliac disease: Out with the old and in with the new? World J Gastroenterol 26(1): 1-10.
- Théry C, Kenneth W Witwer, Elena Aikawa, Maria Jose Alcaraz, Johnathon D Anderson, et al. (2018) Minimal information for studies of extracellular vesicles 2018 (MISEV2018): a position statement of the International Society for Extracellular Vesicles and update of the MISEV2014 guidelines. J Extracell Vesicles 7(1): 1535750.
- Chan BD, Wing-Yan Wong, Magnolia Muk-Lan Lee, William Chi-Shing Cho, Benjamin K Yee, et al. (2019) Exosomes in Inflammation and Inflammatory Disease. Proteomics 19(8): e1800149.
- Console L, Mariafrancesca Scalise, Cesare Indiveri (2019) Exosomes in inflammation and role as biomarkers. Clin Chim Acta 488: 165-171.
- Gao W, Haibo Liu, Jie Yuan, Chaoneng Wu, Dong Huang, et al. (2016) Exosomes derived from mature dendritic cells increase endothelial inflammation and atherosclerosis via membrane TNF-α mediated NF-κB pathway. J Cell Mol Med 20(12): 2318-2327.
- Ludvigsson JF, Green PH (2011) Clinical management of coeliac disease. J Intern Med 269(6): 560-571.
- Pérez-Bravo F, M Araya, A Mondragón, G Ríos, T Alarcón, et al. (1999) Genetic differences in HLA-DQA1* and DQB1* allelic distributions between celiac and control children in Santiago. Chile Hum Immunol 60(3): 262-267.
- Araya M, A Mondragón, F Pérez-Bravo, J L Roessler, T Alarcón, et al. (2000) Celiac disease in a Chilean population carrying Amerindian traits. J Pediatr Gastroenterol. Nutr 31(4): 381-386.
- Araya M, Amaya Oyarzun, Yalda Lucero, Nelly Espinosa, Francisco Pérez-Bravo, et al. (2015) DQ2, DQ7 and DQ8 Distribution and Clinical Manifestations in Celiac Cases and Their First-Degree Relatives. Nutrients 7(6): 4955-4965.
- Megiorni F, Pizzuti A (2012) HLA-DQA1 and HLA-DQB1 in Celiac disease predisposition: practical implications of the HLA molecular typing. J Biomed Sci 19(1): 88.
- Almeida LM, Lenora Gandolfi, Riccardo Pratesi, Rosa Harumi Uenishi, Fernanda Coutinho de Almeida, et al. (2016) Presence of DQ2.2 Associated with DQ2.5 Increases the Risk for Celiac Disease. Autoimmune Dis 5409653.
- Margaritte-Jeannin P, M C Babron, M Bourgey, A S Louka, F Clot, et al. (2004) HLA-DQ relative risks for coeliac disease in European populations: a study of the European Genetics Cluster on Coeliac Disease. Tissue Antigens 63(6): 562-567.
- De Miguel Pérez D, Alba Rodriguez Martínez, Alba Ortigosa Palomo, Mayte Delgado Ureña, Jose Luis Garcia Puche, et al. (2020) Extracellular vesicle-miRNAs as liquid biopsy biomarkers for disease identification and prognosis in metastatic colorectal cancer patients. Sci Rep 10: 3974.
- Zheng X, Kailun Xu, Biting Zhou, Ting Chen, Yanqin Huang, et al. (2020) A circulating extracellular vesicles-based novel screening tool for colorectal cancer revealed by shotgun and data-independent acquisition mass spectrometry. J Extracell Vesicles 9(1): 1750202.
- Tye-Din J A, Gry I Skodje, Vikas K Sarna, John L Dzuris, Amy K Russell, et al. (2020) Cytokine release after gluten ingestion differentiates coeliac disease from self-reported gluten sensitivity. United Eur Gastroenterol J 8(1): 108-118.
- Goel G (2019) Serum cytokines elevated during gluten-mediated cytokine release in coeliac disease. Clin Exp Immunol 199(1): 68-78.
- Garlanda C (2013) The Interleukin-1 Family: Back to the Future. Immunity 39(6): 1003-1018.
- Aksentijevich I, Seth L Masters, Polly J Ferguson, Paul Dancey, Joost Frenkel, et al. (2009) An Autoinflammatory Disease with Deficiency of the Interleukin-1–Receptor Antagonist. N Engl J Med 360: 2426-2437.
- Sokolova V, Anna-Kristin Ludwig, Sandra Hornung, Olga Rotan, Peter A Horn, et al. (2011) Characterisation of exosomes derived from human cells by nanoparticle tracking analysis and scanning electron microscopy. Colloids Surfaces B Biointerfaces 87(1): 146-150.
- Patel G K, Mohammad Aslam Khan, Haseeb Zubair, Sanjeev Kumar Srivastava, Moh’d Khushman, et al. (2019) Comparative analysis of exosome isolation methods using culture supernatant for optimum yield, purity and downstream applications. Sci Rep 9: 5335.
- Lobb R J, Melanie Becker, Shu Wen Wen, Christina S F Wong, Adrian P Wiegmans, et al. (2015) Optimized exosome isolation protocol for cell culture supernatant and human plasma. J Extracell Vesicles 4: 27031.

 Research Article
Research Article