ABSTRACT
Domestic guinea pig is a model animal for human medical research and UDP glucuronosyltransferase 1 family, polypeptide A3 (UGT1A3) is an important human medicine related gene. In this study, the complete coding sequence of domestic guinea pig UGT1A3 gene was amplified by RT-PCR. The open reading frame of domestic guinea pig UGT1A3 gene was 1,605bp encoding a protein of 534 amino acids. Sequence analysis revealed that the UGT1A3 protein of domestic guinea pig shares high homology with the UGT1A3 proteins of human (75%), chimpanzee (75%), sumatran orangutan (75%), northern white-cheeked gibbon (75%), ma’s night monkey (75%) and drill (74%). This gene is structured in five exons and four introns as revealed by computer-assisted analysis. Results also showed that domestic guinea pig UGT1A3 gene has a close genetic relationship with the UGT1A3 gene of ma’s night monkey. The prediction of transmembrane helices revealed that domestic guinea pig UGT1A3 might be a transmembrane protein. The expression profile indicated that domestic guinea pig UGT1A3 gene was differentially expressed in detected domestic guinea pig tissues including small intestine, spleen, lung, muscle, fat, liver, heart and kidney. Our experiment established primary foundation of using domestic guinea pig as model animal to study the UGT1A3-related human diseases.
Keywords: Domestic Guinea Pig; UGT1A3; Molecular Characterization; Tissue Expression
Introduction
UDP glucuronosyltransferase 1 family, polypeptide A3 (UGT1A3) encodes a UDP-glucuronosyltransferase, an enzyme of the glucuronidation pathway that transforms small lipophilic molecules, such as steroids, bilirubin, hormones, and drugs, into water-soluble, excretable metabolites(http://www.ncbi.nlm. nih.gov/gene/54659). UGT1A3 is also an important human medicine related gene. The human UGT1A3 protein catalyzes the glucuronidation of estrogens, bile acids and xenobiotics including non-steroidal anti-inflammatory drugs and lipid lowering drugs (Lankisch, et al. [1,2]) indicated that increased UGT1A3 expression is associated with pancreatic cancer (Trottier J [3]). reported that UGT1A3 enzyme conjugates norursodeoxycholic acid into a C23-ester glucuronide in the liver (Iwai M [4]). revealed that several UGT1A3 polymorphisms exist in the Japanese population and lead to different levels of activity. These polymorphisms are capable of affecting the steady state levels of estrogens and may increase sensitivity to adverse drug effects (Erichsen TJ [5]). also found that UGT1A3 is significantly induced by xenobiotics, and this physiologically links xenobiotic and bile acid metabolism to cholestasis. As mentioned above, UGT1A3 is an important human medicine related gene. Domestic guinea pig is a useful model animal which has been successfully used for human disease research (Schuller S, et al. [6,7]). Until today, the domestic guinea pig UGT1A3 gene has not been reported yet. In present experiment, we will isolate the complete coding sequence of domestic guinea pig UGT1A3 gene, subsequently perform sequence and tissue expression analysis. These will establish the primary foundation of using domestic guinea pig as model animal to study the UGT1A3- related human diseases.
Materials and Methods
Samples Collection, RNA Extraction and First-Strand cDNA Synthesis
Six adult domestic guinea pigs (3 females and 3 males) were killed. Small intestine, spleen, lung, muscle, fat, liver, heart and kidney samples were collected and frozen at – 80 °C (Yang Deming [8]). The total RNA was extracted using the Total RNA Extraction Kit (Gibco, USA). These RNA samples were used to perform reverse transcription to produce the first-strand cDNA (Lankisch [1]). These first-strand cDNAs would be applied to perform RT-PCR and qRT-PCR.
Isolation of the Coding Sequence
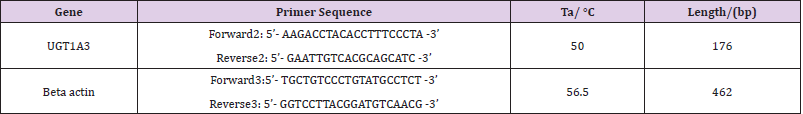
The complete coding sequence of domestic guinea pig UGT1A3 gene was amplified by RT-PCR using the above cDNA as template. The 25 μl reaction system contains 2.0 μl cDNA, 2.5 μl 2 mM mixed dNTPs, 2.5 μl 10×Taq DNA polymerase buffer including Mg2+), 1.0 μl 10 mM forward primer 1(5’- ATG GCA ACA GCA CTG CAG -3’), 1.0 μl 10 mM reverse primer 1(5’- TCA ATG TGT CTT GGA TTT CTG -3’), 2.0 units of Taq DNA polymerase (1 U/1μl), and 14 μl sterile water. The PCR program initially started with a 94°C denaturation for 4 min, followed by 35 cycles of 94°C/1 min, 58°C /1 min, 72°C /1 min, then 72°C extension for 10 min, finally 4°C to terminate the reaction. The PCR product was then sequenced bidirectionally with the commercial fluorometric method.
Quantitative Real Time PCR (qRT-PCR) for Tissue Expression Profile Analysis
qRT-PCR for evaluating the level of mRNA for UGT1A3 gene was performed on the ABI Prism 7300 Sequence Detection Systems (Applied Biosystems, Foster City, CA, USA). PCR reactions for each sample were carried out in 25μl reaction volume containing 1μl SYBR Green real-time PCR Master Mix, 100 ng cDNA template and 200 nM each primer. Conditions for real-time PCR were: an initial denaturation at 95 °C for 3 min, 40 cycles of 95 °C for 15 s, Ta(optimal annealing temperature for each specific primer) for 15 s (Table 1), 72°C for 20 s. For each sample, reactions were set up in triplicate to ensure the reproducibility of the results. The gene relative expression levels were quantified relative to the expression of the reference gene, beta actin (GenBank Accession No. AF508792), by employing the 2 -ΔΔCt value model (Yang Deming, et al. [8,9]).
Sequence Analysis
cDNA sequence prediction was conducted using GenScan software (http://genes.mit.edu/GENSCAN.html). The protein analysis was performed using the tool of BLAST at the National Center for Biotechnology Information (NCBI) server (http://www. ncbi.nlm.nih.gov/BLAST) and the Clustalw software (http://www. ebi.ac.uk/clustalw). The theoretical isoelectric point (pI) and molecular weight (Mw) of protein were also computed using the Compute pI/Mw Tool (http://www.expasy.org/tools/pi_tool.html). The prediction of transmembrane helices in protein was performed using the TMHMM Server v. 2.0 (http://www.cbs.dtu.dk/services/ TMHMM/).
Results and Discussion
Isolation Result for Domestic Guinea Pig UGT1A3 Gene
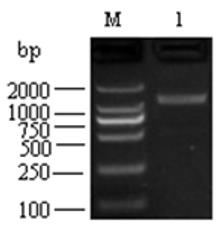
For domestic guinea pig UGT1A3 gene, through RT-PCR with pooled tissue cDNAs, the resulting product was 1,605-bp in length (Figure 1).
Figure 1: PCR result for domestic guinea pig UGT1A3 gene. M DL2000 DNA markers; 1, PCR product for domestic guinea pig UGT1A3 gene.
Sequence Analysis
The cDNA sequence analysis revealed that this PCR product was not homologous to any of the known domestic guinea pig gene and it was then deposited into the Genbank database (Accession number: KT160224). The sequence prediction showed that the 1,605-bp cDNA sequence represents one single gene which encodes 534 amino acids (Figure 2). The pI of domestic guinea pig UGT1A3 is 8.93. The molecular weight of this putative protein is 60484.70Da. Further BLAST analysis of this protein revealed that domestic guinea pig UGT1A3 protein shares high homology with the UGT1A3 proteinsof human (accession number: NP_061966, 75%), chimpanzee (accession number: XP_001151100, 75%), sumatran orangutan (accession number: NP_001125615, 75%), northern white-cheeked gibbon (accesession number: XP_003278593, 75%), ma’s night monkey (accesession number: XP_012289584, 75%) and drill (accesession number: XP_011835648, 74%) (Figure 3). To obtain the genomic DNA of UGT1A3, the publicly available domestic guinea pig genome database at the NCBI Genome Resources (http://www.ncbi.nlm.nih.gov/projects/genome) was screened using the full-length cDNA sequence of UGT1A3 gene as a seed by BLAST analysis. A bacterial artificial chromosome (BAC) clone CH234-244A3 (GenBank accession no. AC190431.2) was found to encompass the entire UGT1A3 gene (38,894bp-64,028bp).
Note: *indicates the stop codon.
Figure 2: The complete coding sequence of domestic guinea pig UGT1A3 gene and its encoding amino acids.
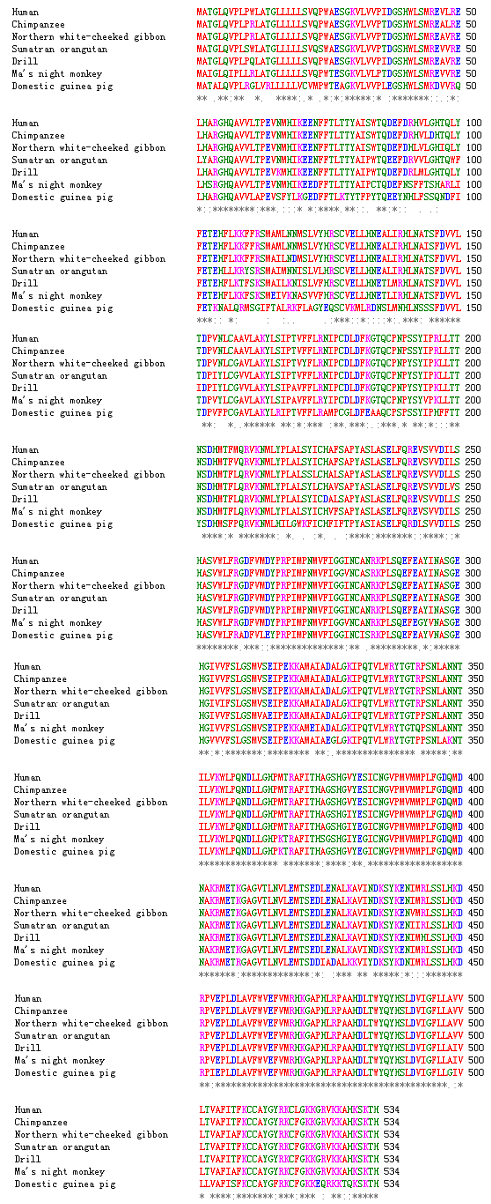
Figure 3: The alignment of the proteins encoded by domestic guinea pig UGT1A3 gene and six other kinds of UGT1A3 proteins.
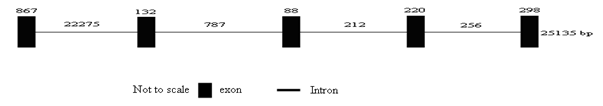
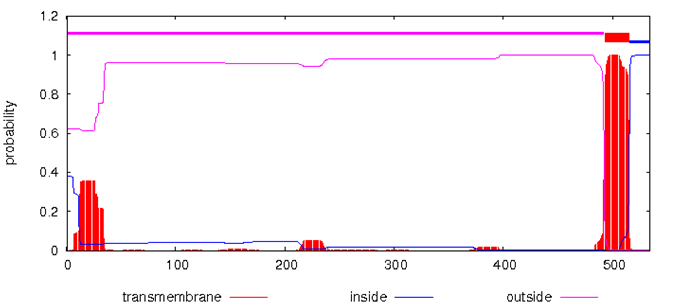
The domestic guinea pig UGT1A3 gene is 25,135 bp in length and consists of five exons and four introns. All exon–intron splice junction sequences conform to the GT–AG rule (Figure 4). The prediction of transmembrane helices in protein showed that domestic guinea pig UGT1A3 protein might be a transmembrane protein (Figure 5). Based on the results of the alignment of different species of UGT1A3proteins, a phylogenetic tree was constructed as shown in (Figure 6). The phylogenetic tree analysis revealed that the domestic guinea pig UGT1A3 gene has a close genetic relationship with that of ma’s night monkey (Figure 7).
Figure 4: The genomic sequence organization representing the ORF of the domestic guinea pig UGT1A3 gene.
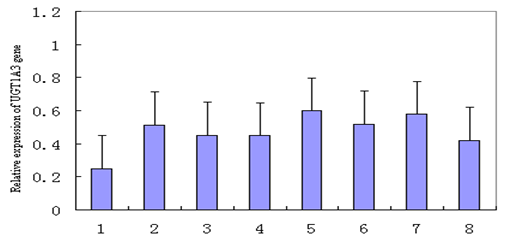
Figure 6: Expression analysis of UGT1A3 gene mRNA in various tissues.
1. Fat;
2. Heart;
3. Lung;
4. Spleen;
5. Muscle;
6. Small intestine;
7. Liver;
8. Kidney.
Tissue Expression Profile
Tissue expression analysis indicated that porcine UGT1A3 genes are moderately expressed in muscle, liver, heart, spleen, lung, kidney and small intestine, and weekly expressed in fat (Figure 6). Comparative genomics research has revealed that the extensive conservation in protein-coding regions implied that this conservation of protein-coding sequences may be expected in domestic guinea pig and other mammalian animals (Hardison RC, et al. [10,8]) From the sequence analysis of UGT1A3 genes, it can be seen that the coding sequences of UGT1A3 genes were highly conserved in some mammalians including human and domestic guinea pig. This implied that these mammalian UGT1A3 genes should have similar functions. We can use the domestic guinea pig as model animal to study the UGT1A3-related diseases of other mammalians including human. We also noticed that domestic guinea pig UGT1A3 protein show 75% identity to human UGT1A3 protein. For UGT1A3 is a important human diseases related gene (Yilmaz et al., 2015), so that, when we use the domestic guinea pig as model animal to study the human UGT1A3-related diseases we should pay more attentions to this difference.
From the tissue distribution analysis in our experiment it can be seen that UGT1A1gene was obviously differentially expressed in some tissues. For UGT1A3 functions as an enzyme of the glucuronidation pathway that transforms small lipophilic molecules, such as steroids, bilirubin, hormones, and drugs, into water-soluble, excretable metabolites http://www.ncbi.nlm. nih.gov/gene/54659 [1], the suitable explanation for this under present condition is that the biological functions of UGT1A1protein might be presented diversely in different tissues. In conclusion, we first isolated the domestic guinea pig UGT1A3 gene and performed necessary sequence analysis and tissue expression profile analysis. Our experiment established primary foundation of using domestic guinea pig as model animal to study the UGT1A3-related human diseases.
Acknowledgment
This work was supported by grants from the Open Project of Key Laboratory for Fertility Regulation and Eugenics of Minority Research of Yunnan Province and National Nature Science Foundation of China (No. 31460645).
Disclosure Statement
No potential conflict of interest was reported by the authors.
References
- Lankisch TO, Gillman TC, Erichsen TJ, Ehmer U, Kalthoff S, et al. (2008) Aryl hydrocarbon receptor-mediated regulation of the human estrogen and bile acid UDP-glucuronosyltransferase 1A3 gene. Arch Toxicol 82(9): 573-582.
- Yilmaz L, Borazan E, Aytekin T, Baskonus I, Aytekin A, et al. (2015) Increased UGT1A3 and UGT1A7 expression is associated with pancreatic cancer. Asian Pac J Cancer Prev 16(4): 1651-1655.
- Trottier J, El Husseini D, Perreault M, Pâquet S, Caron P, et al. (2010) The human UGT1A3 enzyme conjugates norursodeoxycholic acid into a C23-ester glucuronide in the liver. J Biol Chem 285(2): 1113-1121.
- Iwai M, Maruo Y, Ito M, Yamamoto K, Sato H, et al. (2004) Novel UDP-glucuronosyltransferase (UGT1A3) polymorphisms with varying activity. J Hum Genet 49(3): 123-128.
- Erichsen TJ, Aehlen A, Ehmer U, Kalthoff S, Manns MP, et al. (2010) Regulation of the human bile acid UDP-glucuronosyltransferase 1A3 by the farnesoid X receptor and bile acids. J Hepatol 52(4): 570-578.
- Schuller S, Sergeant K, Renaut J, Callanan JJ, Scaife C, et al. (2015) Comparative proteomic analysis of lung tissue from guinea pigswith leptospiral pulmonary haemorrhage syndrome (LPHS) reveals a decrease in abundance of host proteins involved in cytoskeletal and cellular organization. J proteomic 3(122): 55-72.
- Clark S, Hall Y, Williams A (2014) Animal models of tuberculosis: Guinea pigs.Cold Spring. Harb Perspect Med 18(5): 5.
- Yang Deming, An Qingcong, Liu Yonggang (2016) Molecular cloning, sequence identification and expression profile of domesticguinea pig (Cavia porcellus) UGT1A1 gene. BIOTECHNOLOGY & BIOTECHNOLOGICAL EQUIPMENT 30(1): 106-111.
- Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods (4): 402-408.
- Hardison RC (2003) Comparative genomics. PLoS Biol 1(2): E58.

 Research Article
Research Article