Abstract
Objective: To explore the causal relationship between serum calcium concentration
and the risk of COVID-19 in Asian population, this study apply the analysis method of
two samples of Mendelian randomization to demonstrate.
Methods: Through literature search, the summary statistic data of the whole
gene association study (GWAS) related to serum calcium concentration was obtained,
and this data was subjected to secondary data analysis. From the large-sample GWAS
summary data, the genetic variation closely related to the serum calcium concentration
was selected as an instrumental variable. Five models (Inverse variance weighted, MR
Egger, Weighted median, Simple mode and Weighted mode) were used for analyzing and
evaluating the causal relationship between serum calcium concentration and the risk of
COVID-19 with OR value and 95% confidence interval.
Results: A total of 8 SNPs were included as instrumental variables in this study. The
results of gene pleiotropy test showed that there was no gene pleiotropy (P = 0.54). The
IVW results showed that each increase in serum calcium concentration by one standard
deviation would reduce the risk of COVID-19 (OR=0.01, 95%CI: 0.0001-0.73, P=0.03).
Conclusion: Appropriate supplementation of serum calcium concentration can
help prevent the onset of COVID-19.
Keywords: Serum Calcium Concentration; COVID-19; Causal Inference; SNPs; Mendelian Randomization
Abbreviations: MR: Mendelian Randomization; SNPs: Single Nucleotide Polymorphisms; IVs: Instrumental Variables; GWAS: Genome-Wide Association Study; OR: Odds Ratio
Introduction
On February 11, 2020, the Director-General of the World
Health Organization Tan Desai announced in Geneva, Switzerland,
that the 2019 pneumonia infected by the new coronavirus SARSCoV-
2 was named “COVID-19” [1]. Since December 2019, the
new coronavirus is still spreading widely around the world. As
of 0:08 on October 13, 2021, Beijing time, there have been a total
of 237,655,302 confirmed cases of new coronary pneumonia
worldwide, and a total of 4,846,981 deaths, which has brought huge
disasters to society [2]. At present, there is no effective medical
method to kill this virus. Therefore, it is very important to prevent
the new coronavirus infection. In recent years, with the vigorous
development of nutrition, the role of trace elements has gradually
been widely valued by the scientific community. Trace elements
are active substances involved in the intermediate links of the
body’s metabolism, such as calcium. Calcium is the most abundant
inorganic element in the human body [3].
It has the following biological functions [3]:
1. Osteogenesis and blood coagulation;
2. Regulating cell function;
3. Acting as a messenger;
4. Helping to regulate enzyme activity,
5. Maintaining neuromuscular excitability,
6. Reduce the permeability of capillaries and cell
membranes, prevent exudation, and inhibit inflammation and
edema. At present, many studies have proved that serum calcium
concentration is potentially related to the onset and prognosis
of new coronary pneumonia. Some studies have shown that a
lack of serum calcium concentration increases the risk of new
coronary pneumonia infection; but some opinions are that there
is no correlation between serum calcium and the incidence of new
coronary pneumonia. The conclusions are inconsistent. And there
is currently no research on the masses of Asians. In traditional
observational epidemiology, the association between exposure and
outcome may be affected by confounding factors and reverse causal
associations, which limits its use in causal inference. Mendelian
randomization (MR) uses single nucleotide polymorphisms (SNPs)
as instrumental variables (IVs) to infer the causal association
between exposure and outcome, which can overcome confounding
factors and the influence of reverse causality on causal inference
[4,5]. Therefore, this study uses the 2SMR method to conduct a
secondary analysis of genome-wide association study (GWAS)
data to explore the causal relationship between serum calcium
concentration and the risk of COVID-19.
Methods
Study Subjects
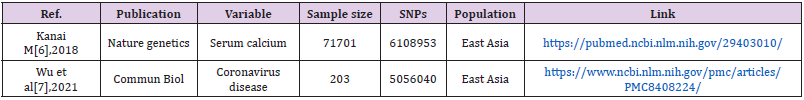
Material Source: Serum calcium concentration is an exposure factor, and whether one person is infected with new coronary pneumonia is the outcome. Download the serum calcium and GWAS summary data of whether or not the new coronary pneumonia is infected through the MR base database and the GWAS Catalog database (Table 1).
Screening of Instrumental Variables: The basic requirements
for genetic variation to become IV are summarized below. The IVs
is related to exposure; variation has nothing to do with confusion
issues related to exposure and outcome; and variation has no
effect on the results, unless it is possible to expose through contact.
First, using the whole genome information of Asian populations as
a reference, SNPs with a significant level (P < 5*10-8) associated
with adults up to the whole genome were screened out. In genetics,
it is generally believed that genetic sites that are very close on
chromosomes are “bundled” together and passed on to offspring.
This also results in a large r2 between sites that are very close. In
order to ensure the independence between SNPs, this study set
the linkage disequilibrium parameter (r2) to 0.001 and the genetic
distance to 10000kb. That is, remove all SNPs whose genetic
distance from the top SNP is r2 > 0.001 within 10000kb. Secondly,
in the outcome data set, if the information about the filtered
instrumental variable is not found, the SNiPA website is used to
search for alternative SNPs (r2 > 0.9); if alternative SNPs cannot be
obtained, the instrumental variable is eliminated. After that, align
the allele effect values in the exposure data set and the outcome
data set, and merge the data [6]. Finally,
1. SNPs that are statistically significant (P < 0.05) with the risk
of COVID-19;
2. SNPs whose BMI with confounding factors reaches a genomewide
significance level (P < 5*10-8);
3. F statistics < 10 SNPs.
Statistical Analysis
1. Weak Instrumental Variable Test: When the degree of
correlation between instrumental variables and exposure
factors cannot reach the significance level of the whole
genome, or when instrumental variables can only explain
a small part of phenotypic variation, such instrumental
variables are called “weak instrumental variables”. In a twosample
MR study, the existence of weak instrumental variables
will underestimate the strength of association and reduce
the power of statistical testing [7]. Therefore, this study uses
the formula 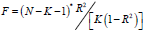 to evaluate whether
there are weak instrumental variables. Among them, N is the
sample size of the exposed data set, K is the number of SNPs,
and R2 is the degree of variation that can be explained by the SNPs in the exposed data set [8]. The calculation formula of R2 is:
to evaluate whether
there are weak instrumental variables. Among them, N is the
sample size of the exposed data set, K is the number of SNPs,
and R2 is the degree of variation that can be explained by the SNPs in the exposed data set [8]. The calculation formula of R2 is: 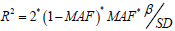 . Among them, MAF is the
minor allele frequency, which is equivalent to the effect allele
frequency EAF when calculating R2, β is the allele effect value,
and SD is the standard deviation.
. Among them, MAF is the
minor allele frequency, which is equivalent to the effect allele
frequency EAF when calculating R2, β is the allele effect value,
and SD is the standard deviation.
2. Gene Pleiotropic Test: When genetic variation can affect
the occurrence of the outcome through other means besides
exposure factors, the instrumental variable is pleiotropic [9].
The existence of gene pleiotropy will cause the “collapse” of
the two-sample Mendelian randomization model. Therefore,
this study used the software package “MR_PLEIOTROPY” in
the R package to test the genetic pleiotropy of instrumental
variables. In addition, the MR-Egger model is used to correct
the bias caused by gene pleiotropy.
3. Linkage Disequilibrium Test: In genetics, it is generally
believed that genetic sites that are very close on chromosomes
are “bundled” together and passed on to offspring, which leads
to r2 between sites that are very close. Will be very large; this
phenomenon is called linkage disequilibrium. Therefore, in
this study, the linkage disequilibrium parameter (r2) was set
to 0.001 and the genetic distance to 10000kb to avoid the
occurrence of linkage disequilibrium. That is, remove all SNPs
whose genetic distance from the top SNP is r2>0.001 within
10000kb.
4. Heterogeneity Test: Cochran’s Q test is used to judge the
heterogeneity of instrumental variables [10].
5. Causal Effect Estimation: Five models of MR Egger, Weighted
median, Inverse variance weighted, Simple mode and Weighted
mode are used to test the causal association between adult
height and the risk of COVID-19. The result is expressed as the
odds ratio (OR) of the risk of COVID-19 for each increase in
height by one standard deviation. In order to strengthen the
reliability of the results, the “Leave-one-out” method is used
for sensitivity analysis, each SNP is eliminated in turn, and the
remaining SNPs are used as IVs for two-sample MR analysis
to determine that a certain SNP will have a strong causal
relationship. Influence.
6. The above analysis methods were all realized by R v4.0.5
software, and the two-sided P < 0.05 indicated that the
difference was statistically significant.
Results
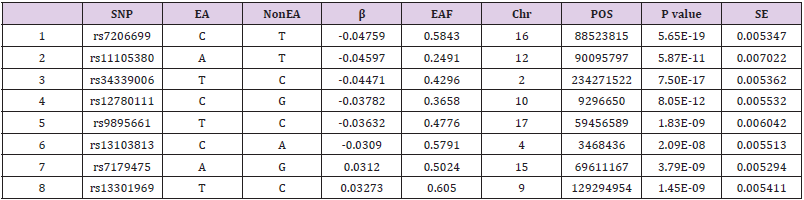
Information About Instrumental Variables
This study finally included a total of 8 SNPs as instrumental variables (Table 2). R2 is 0.25%, the F statistic corresponding to a single SNP has a distribution range of 15.32-38.50, and the F statistic corresponding to 8 SNPs is 22.32, indicating that the causal association is less likely to be affected by weak instrumental variable bias1 [11]. The MR-Egger regression intercept term is 0.33, which shows that there is no genetic pleiotropy between SNPs and the risk of COVID-19. The results of MR_PLEIOTROPY test showed that there was no gene pleiotropic bias (P = 0.54).
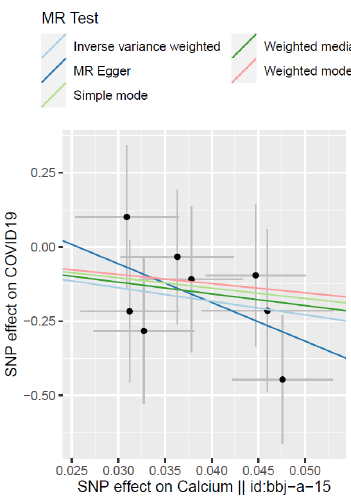
Two-Sample Mr Result
In Asian populations, serum calcium concentration is associated with a reduced risk of COVID-19 (Figure 1 & 2).
Heterogeneity Test Result
The Cochran’s Q test result of IVW is P=0.87, indicating that
there is no heterogeneity in SNPs; the Cochran’s Q test result of MREgger
regression is consistent with it.
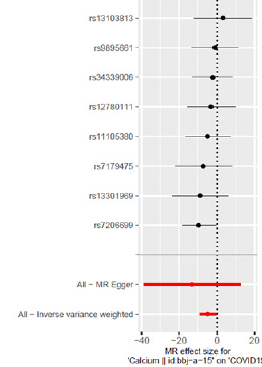
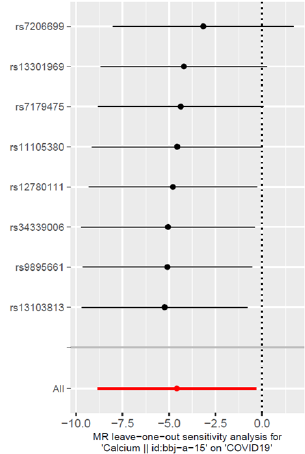
The “Leave-one-out” sensitivity analysis results of this study
are shown in (Figure 3). No matter which SNP is eliminated, the
analysis results of the remaining 7 SNPs are similar to the analysis
results of all the included SNPs, and no influence on the estimated
value of causal association is found. Larger SNPs (Figure 3).
Conclusion
Appropriate supplementation of serum calcium concentration can help prevent the onset of COVID-19.
Discussion
This project uses large-scale GWAS aggregated data and
uses two-sample MR to explore the causal relationship between
serum calcium concentration and the risk of COVID-19 in Asian
populations. The results show that serum calcium can reduce the
risk of COVID-19.
1. Previous studies have shown that in European populations,
serum calcium concentration is negatively correlated with
the risk of COVID-19. This research has reached unanimous
conclusions. This result may be explained by the following
two mechanisms: Calcium ions can regulate cell function and
help regulate enzyme activity, thereby enhancing the body’s
immune function;
2. Calcium ions can reduce the permeability of capillaries and cell
membranes, Prevent oozing, inhibit inflammation and edema.
In other words, the biological pathway involved in calcium ions
may partly coincide with the pathogenesis of COVID-19.
This study has the following advantages: First, the use of public
GWAS data can save research cost and time; Second, the two GWAS
databases are both from Asian populations and are independent of
each other, avoiding population bias; Third, it is not the same as
using a single SNP as the IV In comparison, using 8 SNPs together
as IVs can increase the proportion of calcium ion variation that
can be explained by genetic variation; fourth, compared with
traditional experimental studies, MR simulates a more realistic
random allocation process, and the research design Relatively
simple, research and implementation will not violate ethics. At the
same time, this study also has some limitations. 1. [12] First of all,
MR cannot further explore and explain the biological mechanism of
the influence of genetic variation on the pathogenesis of COVID-19.
Second, MR assumes that there is a linear relationship between
exposure and outcome. If there is a non-linear relationship between
calcium ion concentration and the risk of COVID-19, MR will not
be suitable for causal inferences between the two. Third, this study
uses two GWAS aggregated databases, lacking individual level data,
unable to conduct subgroup analysis of age or gender, and unable
to compare the differences in causal effects between subgroups. At
the same time, the small sample size of the COVID-19 database will
cause problems.
The statistical power of causal inference is reduced. Finally, this
study did not adjust the immune-related indicators, and the MR
method can only make preliminary inferences about its causality,
and further research is needed to explore it. In summary, this
study used two samples of MR to investigate the serum calcium
concentration and the risk of COVID-19. The results showed that
there is a negative causal relationship between the two, that is, the
increase in serum calcium concentration can reduce the incidence
of COVID-19. Risk, which can provide a certain reference value for
the clinical treatment process.
References
- Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, et al. (2015) Global cancer statistics, 2012. CA Cancer J Clin 65(2): 87‐108.
- Fitzmaurice C, Dicker D, Pain A, Michael F MacIntyre, Christine Allen, et al. (2015) The Global Burden of Cancer 2013. JAMA Oncol 1(4): 505‐527.
- Hecht SS (2003) Tobacco carcinogens, their biomarkers and tobacco-induced cancer. Nat Rev Cancer 3(10): 733‐744.
- Ebrahim S, Smith GD (2008) Mendelian randomization: can genetic epidemiology help redress the failures of observational epidemiology? Hum Genet 123(1) :15-33.
- Thomas DC, Conti DV (2004) Commentary: the concept of ‘Mendelian randomization’. Int J Epidemiol 33(1): 21-25.
- Hartwig FP, Davies NM, Hemani G, George Davey Smith (2016) Two-sample Mendelian randomization: avoiding the downsides of a powerful,widely applicable but potentially fallible technique. Int J Epidemiol 45(6): 1717-1726.
- Wu P, Ding L, Li X, Siyang Liu, Fanjun Cheng, et al. (2021) Trans-ethnic genome-wide association study of severe COVID-19. Commun Biol 4(1): 1034.
- Hartwig FP, Davies NM, Hemani G, George Davey Smith (2016) Two-sample Mendelian randomization: avoiding the downsides of a powerful,widely applicable but potentially fallible technique. Int J Epidemiol 45(6): 1717-1726.
- Davies NM, Holmes MV, Davey Smith G (2018) Reading Mendelian randomisation studies: a guide, glossary, and checklist for clinicians. BMJ 362: k601.
- Pierce BL,Ahsan H, Vanderweele TJ (2011) Power and instrument strength requirements for Mendelian randomization studies using multiple genetic variants. Int J Epidemiol 40(3): 740-752.
- Xue Gao, Hui Wang, Tong Wang (2019) Introduction to the correction method of pleiotropic bias in Mendelian randomization. Chinese Journal of Epidemiology 03: 360-365.
- Bowden J, Spiller W, Del Greco MF, Nuala Sheehan, John Thompson, et al. (2018) Improving the visualization, interpretation and analysis of two-sample summary data Mendelian randomization via the Radial plot and Radial regression. Int J Epidemiol 47(4): 1264-1278.

 Research Article
Research Article