ABSTRACT
Staphylococcus spp. it is one of the genera most frequently implicated in the etiology
of hospital infections, especially in nosocomial wards. Staphylococcus aureus is one of
the most frequently isolated pathogenic microorganisms in hospital infections, capable
of causing septicemia and infections of the skin, respiratory system, and soft tissues.
Furthermore, the spread of infections caused by the methicillin-resistant Staphylococcus
aureus (MRSA) species is constantly increasing and is reaching worrying levels in
various countries of the world, including Italy, in continuous and rapid expansion, even
outside hospitals. In this study, strains of Staphylococcus with methicillin resistance
in hospitalized patients were identified and characterized through a phenotypic and
genotypic approach. In all the methicillin resistant strains analyzed, a high resistance
to other classes of antibiotics tested was found, in accordance with the findings of the
European Center for Disease Prevention and Control (CDC) and numerous studies at
national and world level. On some isolated MRSA strains, a molecular epidemiological
study was conducted to understand the origin and spread of circulating clones. All these
have been identified by molecular approach aimed at genetic research and identification,
by means of Multi Locus Sequence Typing (MLST), typing of the Staphylococcal Cassette
Chromosome mec (SCCmec complex) and spa typing, typing of the repeated variable
region of protein A.
Using the MLST profile, 5 different clones of S. aureus were identified in several
hospital departments, 4 of which already circulating in Italy and worldwide, while one
is not yet reported in Italy. The application and deepening of these techniques have
provided an overview of the spread and development of MRSA strains in the hospital
setting.
Keywords: Staphylococcus; Staphylococcus Aureus; Staphylococcal Protein A; Staphylococcal Cassette Chromosome Mec (SCCmec); Mec Gene Complex; MECA; Multilocus Sequence Typing; Polymerase Chain Reaction; Sequencing; Resistance; Epidemiology
Abbreviations: CA-MRSA: Community-Acquired Methicillin-Resistant S. aureus; CDC: Centers for Disease Control and Prevention; CoNS: Coagulase-negative staphylococc; HAMRSA: Hospital-Acquired Methicillin-Resistant S. aureus; MDR: Multi Drug Resistance; MIC: Minimal Inhibitory Concentration; MLST: Multi-Locus Sequence Typing; MRSA: Methicillin-Resistant S. aureus; PBP: Penicillin Binding Protein; PCR: Polymerase Chain Reaction; PST: Penicillin Streptomycin and Tetracycline – Resistant; PSTE: Penicillin streptomycin tetracycline and erythromycin – Resistant; PVL: Panton-Valentine Leukocidin; SCCmec: Staphylococcal Cassette Chromosome mec; ST: Sequence Type; TSST: Toxic Shock Syndrome Toxin; VRSA: Vancomycin-Resistant Staphylococcus Aureus
Introduction
Staphylococci are gram positive bacteria belonging to the
Staphylococcaceae family. They are catalase positive, spherical
in shape arranged in clusters or tetrads, non-spore-forming, and
immobile. Many staphylococci can grow under various conditions,
in the presence and absence of oxygen, with another market
concentration (10% NaCl) and a temperature between 18 °C and
40 °C. Staphylococci are found mainly on the skin and mucous
membranes of mammals, some species have a preferential host
such as Staphylococcus hominis in humans, while others such as
Staphylococcus aureus, find it in more hosts. S. aureus is present
on the skin and mucous membranes in 20-30% of healthy people.
Adolescents and adults often carry short-term or persistent
S. aureus, approximately 15% of healthy adults are persistent
carriers. The adult is colonized by S. aureus for a 30-50%, 20%
of the population in a persistent way. There are also conditions
such as diabetes, drug addiction, immunodeficiency that support
colonization and proliferation and transmission [1-3]. S. aureus
is one of the most common and important human pathogens,
both in the community and in the hospital. The most common S.
aureus infections, defined as staphylococcal, are of the supportive
type, affect various organs and systems with a high and variable
degree of virulence. Infections affect the skin, cutaneous glands,
and subcutaneous soft tissues. There may be localizations in the
site of abscesses in various organs, therefore infections in surgical
wounds and systemic forms.
Other infections are represented by Ritter’s disease or burned
skin syndrome, due to the epidermolysin staphylococcus produced.
It is a toxin capable of detaching the superficial layers of the skin
and by the toxic shock syndrome, TSST-1, also deriving from action
of a toxin that involves symptoms such as: fever, hypotension,
desquamative erythroderma and organ symptoms [1,4,5]. The
main factors that increase susceptibility to infections are the
prolonged or inefficient antibiotic or corticosteroid therapies, the
use of invasive procedures (vascular and bladder catheterization,
tracheal intubation, etc.), prolonged hospitalization and surgical
interventions [6,7]. S. aureus is also responsible for food poisoning,
due to the multiplication in foods of strains of S. aureus producing
toxins resistant to cooking temperatures and the action of digestive
proteolytic enzymes [8,9]. S. aureus is provided with a polysaccharide
capsule, with phagocytic power, neutralized by specific antibodies.
On the cell surface there are proteins that are able to cooperate
with those of the host, such as fibronectin and fibrinogen, playing
the role of adhesions. Among these, the clumping factor is a protein
which, interacting with fibrinogen, forms aggregates that can be
highlighted on the slide. Another important surface protein of S.
aureus is protein A.
This is involved in complement activation, inhibits the
phagocytosis of the bacterium by polymorphonuclear leukocytes,
invokes hypersensitization and stimulation of lymphocyte
production, contributing significantly to increase the virulence
of S. aureus [3,10]. Furthermore, S. aureus has always been an
absolute protagonist of acquired antibiotic resistance. Of particular
importance and interest was the evolution of the resistance of
S. aureus to β-lactam antibiotics, characterized by two distinct
periods of hospital infections. A first hospital infection, which
developed early (around the early fifties of the last century) and
rapidly spread all over the world, was sustained by penicillinresistant
strains, which became such having acquired the ability
to produce penicillinase [11]. The end after 10 years thanks to the
advent of new antibiotics (such as penicillinase-resistant penicillin
and the first cephalosporin’s), even if the phenotypic and genotypic
characteristic of β-lactamase production remained definitively
acquired by most of both hospital community. A second hospital
infection, still ongoing today, is that sustained by methicillinresistant
strains (internationally known with the acronym MRSA,
methicillin-resistant S. aureus), that is, competent of resisting
methicillin, the progenitor of penicillinase-resistant penicillins [4].
Methicillin is characterized by an acyl group in 6 ‘which sterically
prevents attachment to the β-lactam ring, thus preserving its
activity even in the presence of β-lactamase [12,13].
Furthermore, MRSA are resistant not only to penicillinaseresistant
penicillins but to all β-lactams, and in addition they are
characterized by a demonstrated multi-resistance [9,14]. The onset
of MRSA has occurred over time in at least three different areas
that have seen changes in those involved in infections: hospitalized
people, therefore nosocomial infections, people outside the
hospital community and animals. The presence of MRSA was
reported for the first time as a nosocomial infection (hospital -
acquired MRSA, HA -MRSA), affecting hospitalized patients, so
much so that up to the 1970s strains of MRSA represented the
major cause of hospital infections. The beginning and spread of HAMRSA
has been associated with typical risk factors related to the
hospital environment and isolates from patients who were MRSA
negative at hospital admission or MRSA isolates are still defined as
HA-MRSA. Between 1970 and 1990 several HA-MRSA epidemics
occurred in the USA and Japan; pandemics followed by some cases
in Europe [15-17]. Since the 1990s, invasive MRSA infections of
the skin have occurred in patients who are not hospitalized and
who did not possess characteristics to be attributable to HA-MRSA
strains [18-20]. The S. aureus that affects such infections are called
community-acquired MRSA (CA-MRSA). Described for the first time
in the United States, they are potentially dangerous even for the “healthy” population, and are, unfortunately, responsible for most
of the children’s deaths. It was possible to discriminate between
HA-MRSA and CA-MRSA strains thanks to not only phenotypic but
above all genotypic characteristics.
Most infections caused by CA-MRSA involve skin and soft tissue,
and some also produce the toxin PVL [21-24]. S. aureus owes its
resistance to methicillin to the presence in the SCCmec cassette of
the gene encoding a variant of the penicillin binding protein (PBP)
referred to as PBP2a. Beta-lactam antibiotics work by binding
PBPs to the wall, inhibiting the synthesis of peptidoglycan, the
main component of the bacterial wall, thus causing cell death. The
PBP2 variant is unable to bind β-lactams, so the synthesis activity
can continue, making the action of these ineffective. It is a form of
resistance that develops with the production of a protein like the
drug’s target, but not susceptible to it. The mecA gene is regulated
by the Mecl repressor and the β-lactam sensitive transmembrane
signal transducer, MecRI. In the absence of β-lactam antibiotics,
MecI represses the transcription of all the genes of the mec
complex, therefore not only mecA, but also MecRI and mecI.
MecRI with an autocatalytic cut activates the cytoplasmic metalloprotease
domain, which splits the link between Mecl and the
operator region of the mecA gene, allowing the transcription and
production of PBP2a, in the presence of β-lactam. Therefore, the
staphylococcal chromosomal cassette mec (SCCmec) is the main
genetic determinant able to discriminate between the two groups
of HA and CA-MRSA [11,21,25,26]. SCCmec is a mobile genomic
island that encodes various resistance determinants. Currently 8
different types of SCCmec have been described. Types I, II, III and
VIII are associated with HA-MRSA.
While type IV, V, VI and VII are associated with CA-MRSA, virulent
mainly, which mainly affected previously healthy young subjects.
Therefore, according to the single clone theory, the cassette would
have been introduced only once in S. aureus with horizontal transfer
from a species of Staphylococcus, therefore MRSA would have a
single precursor, unlike the multiple clone theory which predicts
that there have been different events and factors involving different
strains of S. aureus [27,28]. Multi-Locus Sequence Typing (MLST)
demonstrated that the 5 pandemic clones of MRSA evolved from
only two genetically distinct ancestral backgrounds: one dating
back to the earliest European MRSA strains and to MSSA strains
circulating in Denmark towards the end of the 1950s, and the other,
a completely different background, attributable to MRSA strains
originally isolated in the USA, Japan and in pediatric patients from
different parts of the world [29,30].
The first European MRSA isolates were characterized by
belonging to the same phage group, resistance to penicillin,
streptomycin, tetracycline (PST) and occasionally to erythromycin
(PSTE), by a low MIC (minimum inhibitory concentration) of
methicillin (6-25 μg/ml), and a heterogeneous expression of
resistance [31,32]. These strains have evolved to the current
clone called Iberic, which has acquired additional resistance
determinants (some resident on mobile elements, such as plasmid
pUB110 and transposon Tn554) and is often resistant to the most
common antibiotics except co-trimoxazole. And glycopeptides.
The Brazilian and Hungarian clones would also have derived
from the first background. The New York / Japan and Pediatric
clones would have derived from the second background. The
Iberic, Hungarian and New York / Japan clones is sensitive only to
co-trimoxazole and glycopeptides. The Brazilian clone is sensitive
only to spectinomycin and glycopeptides. The pediatric clone is
resistant only to oxacillin, penicillin, gentamicin, and occasionally
erythromycin [13,31]. Epidemiologically, the various reports
relating to the isolation of Community MRSA strains outline a
European reality characterized by a polyclonal character. In Italy,
several clones have been described such as ST88, ST30, ST8,
ST72 and ST813. On the contrary in the United States, there is
the diffusion of a clone called USA300, belonging to the ST8 and
USA400 [16,33,34]. The main HA-MRSA clones circulating in the
world belong to the clonal complexes CC5, which includes ST5
SCCmec type II (New York / Japan); ST5-IV pediatric, ST228-I
(southern German); The CC8 with ST250-I (Archaic clone), ST8-IV
(EMRSA-2, -6), ST8-II (Irish), ST239-III (Brazilian / Portuguese),
ST247-I (Iberian); The CC22 with ST22-IV (EMRSA-15); CC30 with
ST36-II (EMRSA-16); The CC45 with ST45-IV (Berlin) [35,36]. The
aim of this work was to characterize the presence of methicillin
resistance in Staphylococcus spp. by phenotypic and genotypic
methods isolated from hospitalized patients.
In addition, an epidemiological-molecular study was performed
on some MRSA isolates from various departments, applying MLST,
to understand the origin and spread of circulating clones.
Materials and Methods
Bacterial Isolates
Eighty-one Staphylococcus spp. strains were isolated and identified. methicillin resistant from patients at the University Hospital of Sassari, Sardinia, Italy. The strains were isolated respectively from 14 blood cultures, 41 samples from the respiratory tract (bronchus aspirate, sputum, nasal, and pharyngeal swabs); 14 from swabs and wound fluids and 12 from other anatomical sites (skin swabs, urine, other). Biochemical identification and antibiogram were performed on all isolates, using the VITEK 2 automated system (Advance Expert System 4.01 software, Biomerieux, Rome, Italy) before being subjected to molecular investigation.
DNA Extraction
Two methods were used for DNA extraction: simple boiling
or boiling prep and the use of the DNeasy Blood & Tissue Kit -
(QIAGEN GmbH, QIAGEN Strasse 1, D-40724 Hilden). Boiling prep.
Some colonies (4 or 5 colonies) were collected and resuspended
in 150μl of sterile double-distilled water and boiled at 100°C for
10 min, to lysate the bacterial wall and obtain the escape of the
DNA. Next it was centrifuged at 10000 rpm for 3 min, allowing
the separation between the pellet (the bacterial lysate) and the
supernatant containing the DNA. One μl of supernatant was used
in the PCR reactions. The DNA thus extracted are stored at - 20 °C.
The instructions of the DNA producers were followed extraction
DNeasy Blood & Tissue Kit (QD). Bacterial strains were grown in
liquid Luria Broth medium under stirring at 37 °C overnight. Pellet
was obtained from 1.5 ml of bacterial culture by centrifugation at
7500 rpm for 10 min. The bacterial pellet was resuspended in 180μl
of enzymatic lysis buffer (20 mM Tris HCl at pH 8.0, 2 mM sodium
EDTA, 1.2% Triton X-100, lysozyme, 20mg/ml) and incubated for
30 min at 37 °C. Then Buffer AL is added with 25μl of Proteinase K
(100mg/ml) and incubated at 56 °C for 30 min for further lysis. The
lysate thus obtained was added with 200μl of ethanol is transferred
to the columns provided by the kit and centrifuged at 8000 rpm for
1 min. This is followed by 2 washes with 500μl of washing Buffer
(AW2).
The DNA was then eluted from the column by adding 100μl of
double distilled water and centrifuging at 8000 rpm for 1 min. The
DNA thus extracted is stored at -20 °C until use.
Detention of S. aureus using PCR Amplification
Validation of S. aureus species identification was performed by PCR using the species-specific primers [37]. Primers were as follows: Fw, SAU1 5’AGGGTTTGAAGGCGAATGGG 3’; and RV, SAU2 (reverse) 5’CAATTTGTCGGTCGAGTTTGCTG3’. The reaction was carried out in a final volume of 25μl which included 22μl of Platinum® PCR Supermix (Hot start recombinant Taq DNA polymerase, buffer 22 mM Tris-HCl at pH8.4, 55 mM KCl, 1.65 mM MgCl₂, 220μM dNTPs, Invitrogen), 1μl of DNA sample and 1μl of each primer (final 0.5μM concentration). The amplification program consisted of an initial denaturation step at 95 °C for 10 min, 35 cycles of denaturing at 95 °C for 30 sec, annealing at 61 °C for 30 sec and extension at 72°C for 2 min; and a final extension at 72°C for 10 min. PCR products were analysed by electrophoresis on a 1% agarose gel, previously stained with GelRed® Nucleic Acid Gel Stain, 10,000X (Biotium, Inc. Landing Parkway. Fremont, CA), and run at 5 V/cm for 40 min. The molecular marker used was a 100 bp ladder (Invitrogen, Waltham, Massachusetts, USA). The sizes of the PCR products sequenced after PCR were 296 bp amplicon.
Detection of the mecA, mecC (mecALGA251), spa e pvl genes using Multiplex PCR in S. aureus Sample
Was designed a Multiplex PCR for 13 samples identified
as S. aureus and 14 invasive CoNS strains, isolated from all
blood culture samples, from several departments (intensive
care unit, surgery, hematology, pneumology, medical
pathology, ENT, nephrology, and dialysis departments)
(23,52) to detect the mecA regulatory genes, MecC, spa and
pvl genes. Primers: mecA P4, 5´TCCAGATTACAACTTCACCAGG
3´; mecA P7, 5´CCACTTCATATCTTGTAACG 3´; spa-
1113F, 5´ TAAAGACGATCCTTCGGTGAGC 3´; spa-1514R,
5´ CAGCAGTAGTGCCGTTTGCTT 3´, to amplify mecC,
mecALGA251 MultiFP, 5´ GAAAAAAAGGCTTAGAACGCCTC
3´; mecALGA251 MultiRP, 5´ GAAGATCTTTTCCGTTTTCAGC
3´; pvl-F, 5´ GCTGGACAAAACTTCTTGGAATAT 3´; pvl-R, 5´
GATAGGACACCAATAAATTCTGGATTG 3´. A 50μl PCR reaction
contained final concentration 1 U of Platinum Taq DNA Polymerase
(Invitrogen); 0.25 mmol/L of each dNTP (GeneAmp, Applied
Biosystems, Warrington, UK); 4 mmol/L of MgCl2; 0.4 μmol/L of
each of forward and reverse primers (spa; mecA; mecALGA251; pvl)
and 2 μl of DNA template. The amplification program consisted of an
initial denaturation step at 94 °C for 5 min, 30 cycles of denaturing
at 94 °C for 1 min, annealing at 59°C for 1 min and extension at 72°C
for 1 min: and a final extension at 72°C for 10 min.
The sizes of the expected PCR products were 162 bp for mecA,
138 bp for mecC, 85 bp for the gene encoding Panton Valentine
Leukocidin (pvl) 180-600 bp for spa fragment (the absence of
fragment spa indicates that the isolate is not a S. aureus) [37,38].
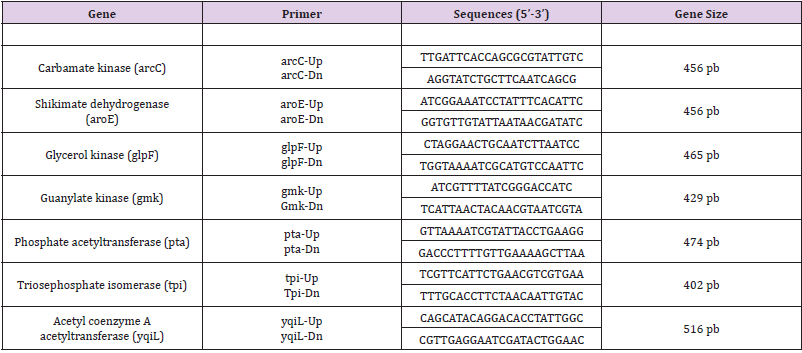
Multilocus Sequence Typing
MLST with standard primers introduced by the MLST database was performed on 7 MRSA isolates based on seven housekeeping genes (arcC, aroE, glpF, gmK, pta, tpiA and yqiL) as described by Enright et al. (2000). The following seven housekeeping genes were used in the final MLST scheme, and the fragments were amplified by using the primers shown in (Table 1). PCRs were carried out with 25 μl reaction volumes containing 1 μL of chromosomal DNA (approximately 0.5 mg), 1.25 μL of each primer, 21,5 μl di Platinum® PCR Supermix (Hot start recombinant Taq DNA polymerase, buffer 22 mmol/L Tris-HCl a pH8.4, 55 mmol/L KCl, 1.65 mmol/L MgCl₂, 220 μM dNTP, Invitrogen). The PCR was performed in a PTC-200 DNA engine (MJ Research, Boston, Mass.) with an initial 3 min denaturation at 94°C, followed by 30 cycles of denaturing at 94 °C for 30 sec, annealing at 55 °C for 30 sec and extension at 72°C for 30 sec; and a final extension at 72°C for 5 min. The amplification products were purified with a MinElute 96 UF PCR purification kit (QIAGEN, Venlo, and The Netherlands) and the samples were sent to the sequencing service, Sequencing Service LMU Munich, Germany (http://www.gi.bio.lmu.de/sequencing). Allele numbers and sequence types (STs) were assigned according to the S. aureus MLST website (http://saureus. mlst.net). Trace files of putative novel alleles and the allelic profiles of novel STs were sent to the database for allele or ST number assignment and admission into the database.
Statistical Analysis
Statistical analysis was performed using Statgraphics Centurion® XV for Windows.
Results
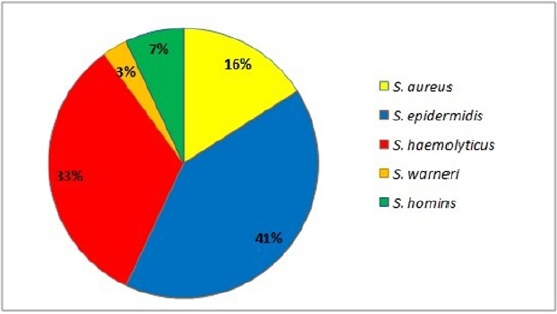
In this study, 81 strains of Staphylococcus spp. were recovered from infected blood samples (17%), respiratory tract samples (51%), wounds (17%) and samples of various kinds (15%). Of the 81 strains, the majority came from inpatients in intensive care (84%). Strains identified included the following Staphylococcus species: 84% Coagulase negative staphylococci (CoNS) of which S. epidermidis, S. haemolyticus, S. hominis, S. warnerii, and S. aureus (16 % n=13) (Figure 1).
Antimicrobial Susceptibility
The following resistance patterns were observed among Staphylococcus spp. isolates: cefoxitin (95%), oxacillin (81%), benzyl penicillin (97%), gentamicin (77%), levofloxacin (85%), erythromycin (86%), clindamycin (48%), and trimethoprim sulfamethoxazole (43%). All isolates were susceptible to vancomycin, teicoplanin, linezolid and tigecycline. On the contrary, all Staphylococcus spp. isolates were sensitive to vancomycin, teicoplanin, linezolid and tigecycline. Of 13 Staphylococcus aureus isolates, 11 (85%) were MRSA and MDR. The predominant resistance profile among MDR isolates included a resistance profile to 7 antibiotics (53.9%) followed by 6 antibiotics (7.7%), 5 antibiotics (15.3%), 3 antibiotic (7.7%) and 2 antibiotics (15.3%) simultaneously.
Distribution of mecA, mecC (mecALGA251), spa and pvl
Multiplex-PCR analysis for detection of different mecA, mecC (mecALGA251), spa and pvl revealed the mecA gene for methicillin resistance in all 14 CoNS (100%) and 11 of 13 of the MRSA (84.6%). The mecC gene was found in 9 MRSA isolates (69.2%). All MRSA samples have showed the presence of spa and the absence of pvl. On the other hand, the previous genes (spa and pvl) were not found in 14 CoNS strains.
MLST
According to the MLST method, isolates were assigned to five different sequence types (STs) (ST5 in 1 strain, ST8 in 1 strain, ST10 in 1 strain, ST22 in 2 strains, and ST228 in 2 strains). Furthermore, the 3 MRSA of care unit were belonged to ST8 (n = 1) and ST228 (n = 2), the strain isolated from the Surgical Clinic showed ST5, from hematology the ST10, while the isolates of Infectious Diseases (n = 1) and of Pneumology (n = 1) were ST22.
Discussion
S. aureus is one of the species most frequently implicated in
the etiology of hospital infections in different parts of the world,
especially in the intensive care, pneumology, hematology, and
surgery departments [39,40]. Although with lower percentages,
CoNS are also emerging as important opportunistic pathogens,
and are often involved in hospital epidemics [41,42]. This study,
in agreement with these studies, highlighted beyond the isolation
of S. aureus, a high percentage of CoNS from clinical samples from
acutely patients, confirming the growing involvement of these
problems in nosocomial infections. The MRSA spread infections
is increasing and is achieving worrying levels in several countries,
including Italy. Since Staphylococcus spp., in particular MRSA is
transmitted through infected people, or vehicles, the first strategy
to contain this spread may therefore concern the implementation
of prevention, as suggested by the guidelines [43,44]. In this work,
all methicillin resistant strains were found to have high resistance
to other classes of tested, in accordance with what was reported by
the European Center for Disease Prevention and Control (CDC) [45].
The mecA gene was considered the “golden standard” for detecting
methicillin resistance in MRSA, however, recently methicillinresistant
mecA negative strains have been found, in which the
presence is associated with the mecC analogue (mecALGA251).
In this work 97% of methicillin-resistant staphylococci had
showed the presence of the mecA gene. Instead, in two isolates,
despite being resistant to methicillin from the analysis with Vitek2,
they did not possess the mecA and cC genes, highlighting, as reported
by other authors, the limits of the phenotypic systems [46,47]. The
data confirmed that HA-MRSA showed the virulence gene of Protein
A (spa) but not the Leukocidin Panton - Valentine (pvl) gene, usually
associated with CA-MRSA a community circulation [48]. Through
the MLST profile have been identified 5 different clones of S. aureus,
4 of which ST5, ST8, ST22 and ST228 already circulating in Italy and
worldwide, while the ST10 was not yet reported in Italy, was present
only at community and veterinary level, confirming the trend of
diffusion and exchange between CA-MRSA and HA-MRSA [49]. The
ST5 profile strain from surgical clinic, linked to the type of sequence
of a HA-MRSA widespread throughout the world and responsible
for nosocomial, tract, mucosal and wound complications. Strains of
ST8 and ST228 were identified in the intensive care unit isolates,
detecting the circulation of at least two different clones in this
unit. The presence of strains with characteristics such as to be
included in ST8 and ST228, found to be circulating in both hospital
and community settings, has been reported throughout the world
[3,31,43].
Furthermore, MRSA with ST22 type sequence had been isolated
from different types of samples from infectious disease and
pneumology department, clone was found mainly in hospital and
outpatient clinics, but also in communities and in animals in close
contact with humans (dogs and cats) [3,46]. Finally, in this work, a
type of ST10 sequence never reported in Italy was found coming
from a nasal swab of the hematology department.
Conclusion
In conclusion, this study demonstrated the importance of constant supervision of the clones circulating in the several hospital departments, colonization, and the probable, but already possible, diffusion and exchange of strains found in the hospital and then in the community. This study was conducted on clinical samples that were chosen to represent the reality nosocomial situation. Although conducted on a restricted number of samples, it provides a database for the design of targeted screening and preventive molecular diagnostics.
Conflict of Interest
Declare that I have no economic interest or conflict of interest.
References
- Antonelli G, Clementi M, Pozzi G, Rossolini GM (2012) Principi di microbiologia medica. Italy, CEA.
- Bistoni F, Blasi E, Castelli F, Ceccherini Nelli L, Cermelli C, et al. (2014) La Placa-Principi di Microbiologia Medica.
- Odonkor ST, Addo KK (2011) Evaluation of three methods for detection of methicillin resistant Staphylococcus aureus (MRSA). Int J Biol Med Res 2(4): 1031-1034.
- Farahani A, Mohajeri P, Gholamine B, Rezaei M, Abbasi H (2013) Comparison of different phenotypic and genotypic methods for the detection of methicillin-resistant Staphylococcus aureus. North American journal of medical sciences 5(11): 637-640.
- Food Safety Authority (2009) Analysis of the baseline survey on the prevalence of methicillin‐resistant Staphylococcus aureus (MRSA) in holdings with breeding pigs, in the EU, 2008‐Part A: MRSA prevalence estimates. EFSA Journal 7(11): 1376.
- Chambers HF (1988) Methicillin-resistant staphylococci. Clinical microbiology reviews 1(2): 173-186.
- Enright MC, Robinson DA, Randle G, Feil EJ, Grundmann H, et al. (2002) The evolutionary history of methicillin-resistant Staphylococcus aureus (MRSA). Proceedings of the National Academy of Sciences 99(11): 7687-7692.
- Gleeson TD (2008) Prevention and control of methicillin-resistant Staphylococcus aureus. Disease-a-month 54(12): 801-806.
- Pan A, Cappelli V, Parenti M, Pantosti A, Pompa MG, et al. (2011) Raccomandazioni sul controllo della diffusione nosocomiale dello Staphylococcus aureus resistente alla meticillina (MRSA). Bologna, maggio.
- Loncaric I, Kübber Heiss A, Posautz A, Stalder GL, Hoffmann D, et al. (2014) mec C‐and mec A‐positive meticillin‐resistant S taphylococcus aureus (MRSA) isolated from livestock sharing habitat with wildlife previously tested positive for mec C‐positive MRSA. Veterinary dermatology 25(2): 147-148.
- Deurenberg RH, Vink C, Kalenic S, Friedrich AW, Bruggeman CA, et al. (2007) The molecular evolution of methicillin-resistant Staphylococcus aureus. Clinical Microbiology and Infection 13(3): 222-235.
- Appelbaum PC (2007) Reduced glycopeptide susceptibility in methicillin-resistant Staphylococcus aureus (MRSA). International journal of antimicrobial agents, 30(5): 398-408.
- Deurenberg RH, Stobberingh EE (2008) The evolution of Staphylococcus aureus. Infection, genetics and evolution 8(6): 747-763.
- Barber M, Rozwadowska Dowzenko M (1948) Infection by Penicillin-Resistant Staphylococci. Lancet 2(6530): 641-644.
- Boucher HW, Corey GR (2008) Epidemiology of methicillin-resistant Staphylococcus aureus. Clinical infectious diseases 46(Supplement_5): S344-S349.
- Campanile F, Bongiorno D, Borbone S, Mongelli G, Jeddari S, et al. (2008) Looking back, current status, and future trends of MRSA clones in Italy: P1426. Clinical Microbiology & Infection 14.
- Campanile F, Bongiorno D, Borbone S, Stefani S (2009) Hospital-associated methicillin-resistant Staphylococcus aureus (HA-MRSA) in Italy. Annals of clinical microbiology and antimicrobials 8(1): 22.
- Gattuso G, Palvarini L, Tomasoni D, Ferri F, Scalzini A (2009) A case of community-acquired MRSA (CA-MRSA) sepsis complicated by meningoencephalitis and cerebral abscess, successfully treated with linezolid. Infez Med 17(4): 244-248.
- Pan A, Battisti A, Zoncada A, Bernieri F, Boldini M, et al. (2009) Community-acquired methicillin-resistant Staphylococcus aureus ST398 infection, Italy. Emerging infectious diseases 15(5): 845-847.
- Qu T, Feng Y, Jiang Y, Zhu P, Wei Z, et al. (2014) Whole genome analysis of a community-associated methicillin-resistant Staphylococcus aureus ST59 isolate from a case of human sepsis and severe pneumonia in China. PLoS One 9(2): e89235.
- De Lencastre H, Tomasz A (2011) The CEM-NET initiative: molecular biology and epidemiology in alliance–tracking antibiotic-resistant staphylococci and pneumococci in hospitals and in the community. International Journal of Medical Microbiology 301(8): 623-629.
- Gonzalez BE, Rueda AM, Shelburne SA, Musher DM, Hamill RJ, et al. (2006) Community-associated strains of methicillin-resistant Staphylococcus aureus as the cause of healthcare-associated infection. Infection Control & Hospital Epidemiology 27(10): 1051-1056.
- Stefani S (2009) Evolution in the antibiotic susceptibility and resistance. Le infezioni in medicina 17: 5-12.
- Stefani S, Chung DR, Lindsay JA, Friedrich AW, Kearns AM, et al. (2012) Meticillin-resistant Staphylococcus aureus (MRSA): global epidemiology and harmonisation of typing methods. International journal of antimicrobial agents 39(4): 273-282.
- Archer GL, Bosilevac JM (2001) Signaling antibiotic resistance in staphylococci. Science 291(5510): 1915-1916.
- D’Orio V, Festino AR, Costanzo C, Di Ciccio P, Colavita G, et al. (2008) Biomolecular identification of methicillin-resistant strains of Staphylococcus aureus (mrsa) isolated from meat and meat processing environments. Italian Journal of Food Safety p. 35-38.
- Zhang HZ, Hackbarth CJ, Chansky KM, Chambers HF (2001) A proteolytic transmembrane signaling pathway and resistance to β-lactams in staphylococci. Science 291(5510): 1962-1965.
- Zhang K, McClure JA, Elsayed S, Louie T, Conly JM (2005) Novel multiplex PCR assay for characterization and concomitant subtyping of staphylococcal cassette chromosome mec types I to V in methicillin-resistant Staphylococcus aureus. Journal of clinical microbiology, 43(10): 5026-5033.
- Enright MC, Day NP, Davies CE, Peacock SJ, Spratt BG (2000) Multilocus sequence typing for characterization of methicillin-resistant and methicillin-susceptible clones of Staphylococcus aureus. Journal of clinical microbiology 38(3): 1008-1015.
- Livermore DM (2000) Antibiotic resistance in staphylococci. International journal of antimicrobial agents 16: 3-10.
- Humphreys H, Grundmann H, Skov R, Lucet JC, Cauda R (2009) Prevention and control of methicillin-resistant Staphylococcus aureus. Clinical microbiology and infection 15(2): 120-124.
- Manso E, Varaldo PE (2002) La resistenza agli antibiotici negli stafilococchi.
- Padmanabhan RA, Fraser TG (2005) The emergence of methicillin-resistant Staphylococcus aureus in the community. Cleveland Clinic journal of medicine 72(3): 235-241.
- Pantosti A, Venditti M (2009) What is MRSA? European Respiratory Journal 34(5): 1190-1196.
- Pantosti A (2006) Ca-MRSA in Italia e nel mondo. Microbiologia Medica 21(3).
- Vignaroli C, Mancini A, Varaldo PE (2014) Composite SCCmec element in single-locus variant (ST217) of epidemic MRSA-15 clone. Emerging infectious diseases 20(5): 905-907.
- Strommenger B, Kettlitz C, Werner G, Witte W (2003) Multiplex PCR assay for simultaneous detection of nine clinically relevant antibiotic resistance genes in Staphylococcus aureus. Journal of clinical microbiology 41(9): 4089-4094.
- García Álvarez L, Holden MT, Lindsay H, Webb CR, Brown DF, et al. (2011) Meticillin-resistant Staphylococcus aureus with a novel mecA homologue in human and bovine populations in the UK and Denmark: A descriptive study. The Lancet infectious diseases 11(8): 595-603.
- Geraci DM, Giuffre M, Bonura C, Matranga D, Aleo A, et al. (2014) Methicillin-resistant Staphylococcus aureus colonization: A three-year prospective study in a neonatal intensive care unit in Italy. PLoS One 9(2): e87760.
- Dufour P, Gillet Y, Bes M, Lina G, Vandenesch F, et al. (2002) Community-acquired methicillin-resistant Staphylococcus aureus infections in France: Emergence of a single clone that produces Panton-Valentine leukocidin. Clinical Infectious Diseases 35(7): 819-824.
- Almeida GC M, Dos Santos MM, Lima NGM, Cidral TA, Melo MCN, et al. (2014) Prevalence and factors associated with wound colonization by Staphylococcus and Staphylococcus aureus in hospitalized patients in inland northeastern Brazil: A cross-sectional study. BMC infectious diseases 14(1): 328.
- Hagiya H, Matsumoto M, Yamasawa T, Haruki Y, Otsuka F (2014) A Case of Vascular Graft Infection Caused by Staphylococcus lugdunensis after Femoropopliteal Bypass Operation. Acta Medica Okayama 68(3): 171-175.
- Stefani S, Varaldo PE (2003) Epidemiology of methicillin-resistant staphylococci in Europe. Clinical Microbiology and Infection 9(12): 1179-1186.
- Stegger M, Lindsay JA, Moodley A, Skov R, Broens E M, et al. (2011) Rapid PCR detection of Staphylococcus aureus clonal complex 398 by targeting the restriction-modification system carrying sau1-hsdS1. Journal of Clinical Microbiology 49(2): 732-734.
- European Centre for Disease Prevention and Control, Suetens C, Hopkins S, Kolman J, Högberg LD (2013) Point Prevalence Survey of Healthcare-associated Infections and Antimicrobial Use in European Acute Care Hospitals: 2011-2012. Publications Office of the European Union.
- Campanile F, Bongiorno D, Falcone M, Vailati F, Pasticci MB, et al. (2012) Changing Italian nosocomial-community trends and heteroresistance in Staphylococcus aureus from bacteremia and endocarditis. European journal of clinical microbiology & infectious diseases 31(5): 739-745.
- Cartwright E J, Paterson GK, Raven KE, Harrison EM, Gouliouris T, et al. (2013) Use of Vitek 2 antimicrobial susceptibility profile to identify mecC in methicillin-resistant Staphylococcus aureus. Journal of clinical microbiology 51(8): 2732-2734.
- Vandenesch F, Naimi T, Enright MC, Lina G, Nimmo GR, et al. (2003) Community-acquired methicillin-resistant Staphylococcus aureus carrying Panton-Valentine leukocidin genes: Worldwide emergence. Emerging infectious diseases 9(8): 978-984.
- Ho PL, Chiu SS, Chan MY, Gan Y, Chow K H, et al. (2012) Molecular epidemiology and nasal carriage of Staphylococcus aureus and methicillin-resistant S. aureus among young children attending day care centers and kindergartens in Hong Kong. Journal of Infection 64(5): 500-506.

 Research Article
Research Article