Abstract
Nucleic acids that can bind with high affinity and specificity to target molecules are called “aptamers”. Aptamers are single-stranded nucleic acids that directly inhibit a protein’s function by folding into a specific three-dimensional structure that dictates high-affinity binding to the targeted protein. Aptamers exhibit significant advantages relative to protein therapeutics in terms of size, synthetic accessibility, and modification by medicinal chemistry. Despite these properties, aptamers have been slow to reach the marketplace, with only one aptamer-based drug receiving approval so far. Recently, aptamers constitute a new class of oligonucleotides that have gained therapeutic importance and pegaptanib is the first approved aptamer drug that is class of oligonucleotides and is often referred to as ‘chemical antibodies. The development of pegaptanib is looked at for the challenges faced in converting aptamers into therapeutic molecules. Because they inhibit the activity of existing proteins directly, aptamers are more similar to a monoclonal antibody or small molecule drugs than to antisense compounds, and this property greatly increases the number of clinical indications that are potentially treatable by nucleic acid-based compounds.
Keywords: Aptamers; Oligonucleotides; Pegaptanib; SELEX
Introduction
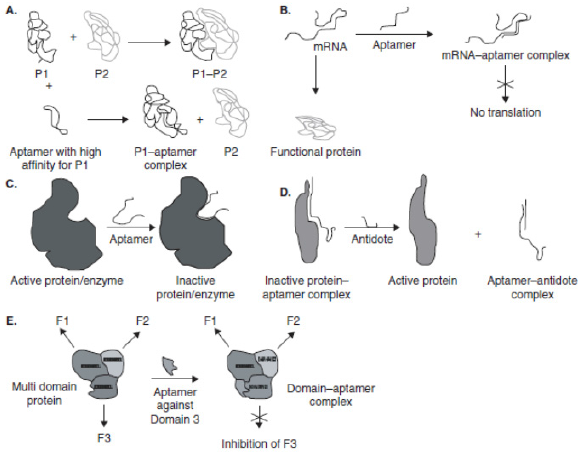
Nucleic acids can not only hybridize to one another based on simple code but can also form complex shapes that may act as scaffolds for molecular interactions and support complex formation with protein and small-molecule targets. Although this is true for biological nucleic acids 1-4, it was only recently that a series of technological advances allowed the development of in vitro evolutionary methods for the discovery of additional, nonbiological oligo nucleotides that can bind to protein targets [1]. Nucleic acid ligands generated using SELEX have been termed as aptamers, an invented Latin term that means ‘to fit’. Aptamers are the only few classes of peptide molecules that, like antibodies, can be crafted to bind to multiple different targets [2,3]. Aptamers are referred to as ‘chemical antibodies. These are short, singlestranded oligo nucleotides (either RNA or DNA) that recognize their targets (organic and nonorganic molecules) based on shape complementarity [1,4,5]. In this, they break away from the oligarchy of oligo nucleotides. Most of the conventional oligo nucleotide therapies target the translational machinery of the cell by destroying the mRNA [6]. Aptamers, on the other hand, target proteins directly and bind with them, modulating their functions. This can be accomplished in several ways (Figure 1): the first one is the function of the protein can be inhibited if the aptamer blocks or distorts the active site of the protein [7]; secondly, the interaction of the protein with another protein/DNA/receptor may be interrupted [8]; or else a specific domain in a multi-subunit protein may be inhibited [9].
Aptamers are in vitro selected DNA or RNA molecules that are capable of binding a wide range of nucleic and non-nucleic acid molecules with high affinity and specificity. They have been conducted through the process known as SELEX (Systematic Evolution of Ligands by Exponential Enrichment) [10]. In a typical SELEX experiment, a random sequence oligo nucleotide library is synthesized that spans 20-100 residues in length, and each nucleotide is flanked by constant sequences required for enzymatic manipulation. The pool will frequently contain between 1 × 1013 to 1 × 1015 members, although it is claimed that even less diverse pools will yield useful aptamers [11,12]. This process serves to reach specificity and considerable affinity to target molecules, including those of viral origin, both proteins, and nucleic acids. Properties of aptamers allow detecting virus-infected cells or viruses themselves and make them competitive to monoclonal antibodies. Specific aptamers can be used to interfere in each stage of the viral replication cycle and also inhibit its penetration into cells. Many current studies have reported the possible application of aptamers as a treatment or diagnostic tool in viral infections, e.g., HIV (Human Immunodeficiency Virus), HBV (Hepatitis B Virus), HCV (Hepatitis C Virus), SARS (Severe Acute Respiratory Syndrome), H5N1 avian influenza and recently spread Ebola and Covid-19 virus [10,13].
According to the current study, two Korean patents describe the use of RNA aptamers for inhibition of SARS viruses. Patent application KR2009128837 identifies RNA aptamers as anti-SARS agents capable of binding to and inhibiting the double-stranded DNA unwinding of the SARS virus helicase. Related patent application KR 2012139512 describes RNA aptamers with a distinct affinity for the nucleo-capsid of SARS-CoV (Covid-19) for potential pharmaceutical use [13]. For therapeutic applications aptamers are frequently in competition with small molecules and antibodies. Initially, aptamers targeted to vascular endothelial growth factor (VEGF) found utility in the treatment of wet macular degeneration-12, in part because they had long half-lives in the ocular compartment [11]. Aptamers that have properties more suitable for systemic administration are now being developed, primarily against targets in the bloodstream, such as thrombin, factor IXa, and von Willebrand factor, or on cell surfaces such as epidermal growth factor receptor (EGFR). It may eventually be possible to use aptamers to access targets inside cells, either by delivering themselves or other drugs across membranes. Also, the types of aptamers that can be discovered are beginning to expand, and now include agonists as well as antagonists. One aptamer has so far been marketed for therapeutic application, and a further eight aptamers are currently being evaluated in the clinic, including those comprising DNA, modified RNA, and Spiegelmers [14].
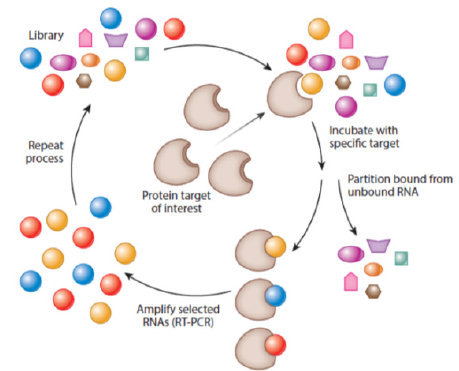
Aptamers are generated in the method referred to as SELEX (Systematic Evolution of Ligands by Exponential Enrichment). This technique was invented in 1990 independently by two teams: Ellington and Szostak as well as Tuerk and Gold and has remained immutable since the discovery of aptamers [15,16]. Cycles of selection and replication followed one by one, are required to obtain target-specific sequences. The combinatorial library, containing all possible sequences with particular length (aptamers have approximately 20-90 nucleotides), is incubated with the target molecule. This process leads to binding with nucleic acids of the highest affinity with the target. At the same time, other oligo nucleotides are removed from the primary sequences pool. Particles of nucleic acids bound with the target molecule, undergo recovering and then replication. The process relies on multiple rounds of selection and replication until high specificity and low dissociation aptamers towards the target molecule are generated [10] (Figure 2). Certain modifications of SELEX (RNA and DNA SELEXs) are possible, including its combination with such techniques as capillary electrophoresis (CE) or surface plasm on resonance (SPR). This is expected to reduce the period of selection time, whereas chemical modifications of aptamers’ increase their bio-stability in vivo [17-19]. Specific aptamers for single atoms, molecules, and also entire bacteria, viruses, cells and tissues can also be engineered by using different selection strategies and modifications of incubation conditions (pH, temperature, etc.). The secondary and tertiary structures of aptamers provide its binding with the target molecule [20,21].
Figure 2: The SELEX (Systematic Evolution of Ligands by Exponential Enrichment) process. Source: Nimjee et al; [11].
Despite the aptamers have obvious advantages, further clinical studies are necessary before they can be applied in therapy; especially aptamer safety and efficacy should be considered. The first and for the time being the only pharmaceutical aptamer, Macugen (pegaptanib sodium), has been admitted in 2004 by the US Agency for Food and Drug Administration (FDA) in the therapy of Age-Related Macular Degeneration (AMD) [1,22]. Several aptamers are currently being evaluated in phases II and III (Macugen and E2F decoy) of clinical trials, for example in hemophilia (ARC19499) [19] von Willebrand’s disease (ARC1779) [23,24] and lung cancer (AS1411) [19,25]. Apart from cancers, neurodegenerative, autoimmunological, and bacterial diseases, there is currently another significant group of therapeutic targets, i.e., viral infections. However, after decades of research, no treatment of many viral diseases has moved past clinical trials to reach the market or the standard therapy is not satisfactory enough. Thus, aptamers could be potentially used in the detection of specific viral molecules and treatment of viral infections [26,27]. Therefore, this review highlights the current achievements of using aptamers in diagnostic and therapeutic methods of viral infections.
History of Aptamer
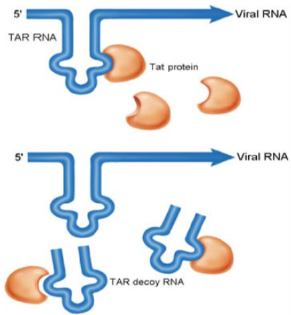
The concept that nucleic acid ligands could modulate the activity of target proteins emerged from basic science studies of viruses. In the 1980s, research on the human immunodeficiency virus HIV and adenovirus discovered that these viruses encode several small, structured RNAs that bind to viral or cellular proteins with high affinity and specificity. Not surprisingly, functional analyses of these viral RNA ligands demonstrated that the viruses had evolved these aptamers either to modulate the activity of proteins essential for their replication [28] or to inhibit the activity of proteins involved in cellular antiviral responses [29,30]. For example, HIV has evolved a short, structured RNA ligand called TAR (trans-activation response) that binds to a viral protein called Tat (transactivation of transcription) as well as the cellular protein cyclin T1 to control viral gene expression and replication (Figure 3) [28]. Adenovirus has evolved a short, structured RNA aptamer, termed Virus Associated RNA (VA-RNA), to inhibit interferoninduced Protein Kinase R (PKR) activity and thus subvert the cell’s antiviral defense system [31]. The observation that viruses utilize RNA ligands for their ends suggested to translational researchers in the late 1980s that RNA ligands might also be useful for therapeutic ends. The first study performed to determine if an RNA aptamer could be used to inhibit the activity of a pathogenic protein was published in 1990. This groundbreaking work demonstrated that the TAR aptamer, evolved by HIV to recruit viral and cellular proteins to viral transcripts, could be turned against the virus to inhibit its replication (Figure 3) [32].
Figure 3: TAR decoy RNA aptamer preventing Tat protein from binding to viral TAR RNA. By inhibiting TAR-Tat interaction the aptamer inhibits HIV RNA transcription and replication. Nimjee et al; [20].
The TAR aptamer sequence was expressed from a transferase ribonucleic acid (tRNA) promoter to act as a decoy for the viral Tat and cellular cyclin T1 proteins in CD4+ T cells. Cells that expressed high levels of the TAR aptamer were shown to be highly resistant to viral replication and cytotoxicity [32,33]. Thus, these studies described a novel method to inhibit HIV replication and established that RNA ligands could serve as therapeutic agents to directly bind and inhibit the activity of clinically relevant proteins [5,34,35]. Two other groundbreaking studies, also published in 1990, demonstrated that large libraries of RNAs could be screened in vitro for RNA ligands that bind T4 DNA polymerase and a variety of organic dye, respectively [16,36]. This selection process was termed SELEX by Tuerk and Gold [16] and RNA ligands were named aptamers by Ellington and Szostak [36]. The 1990 studies on in vitro evolution of RNA aptamers suggested that the concept of therapeutic aptamers [16,36]. It is now becoming clear that SELEX can rapidly generate aptamers to many therapeutically relevant targets, and these aptamers are on the verge of becoming an exciting new class of therapeutic agents. Three aptamers are now being evaluated in clinical trials for inhibition of HIV replication, angiogenesis, and intimal hyperplasia. Two of these aptamers evolved in nature and the third was generated in vitro. Numerous aptamers are in preclinical development, and a few of these are slated to begin clinical evaluation soon [20].
General Properties of Aptamer Compounds
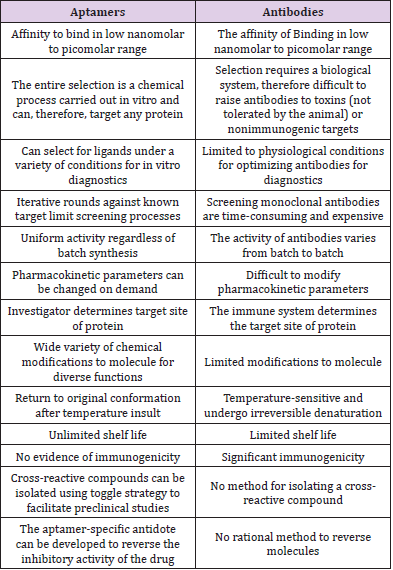
Unique properties of aptamers make them competitive to monoclonal antibodies, currently used in conventional laboratory practice. First of all, the time of aptamers selection is relatively short (a few weeks) in comparison to monoclonal antibodies production (a few months). Unquestionable aptamers’ advantage is an opportunity to acquire ligands directed against toxic molecules. It is almost impossible in the case of monoclonal antibodies because of obligatory animals’ immunization. Also, aptamers are relatively small in size. Thus, it makes them attractive for in vivo applications, in comparison to the large molecule of monoclonal antibodies. Moreover, aptamers could be easily modified with drugs and immune fluorescence dyes without losing their primary properties [19,21]. Affinities of aptamers for the targeted proteins tend to be very high, with typical dissociation constants ranging from low picomolar (1 × 10–12 M) to low nano molar (1× 10–9 M). As in vitro selection techniques have improved, the generation of aptamers with sub-nano molar affinities for the target has become increasingly common. These affinities are similar to those measured for interactions between monoclonal antibodies and antigens. However, since the dissociation constants measured for aptamer-target proteins are true affinities, reflecting a bimolecular interaction in solution, they are more accurately compared to the affinities of Fab fragments for their target antigens. On average, the affinities of aptamers for a targeted protein are stronger than is typical for interactions between Fab fragments and their target antigens [14,37].
High-affinity nucleic acid-protein interactions require specific complementary contacts between functional groups on both the nucleic acid and the protein. Because the specific three-dimensional arrangement of complementary contact sites that mediate the protein aptamer interaction is unlikely to be recapitulated in other proteins, aptamers are generally specific for their targets. For example, aptamers have been generated that are capable of discriminating between isoforms of protein kinase C that share a high degree of identity [2,38]. Because of the specific affinity that aptamers display for their respective target molecules, they constitute very good affinity ligands for the capture of the target molecule from complex mixtures. A classic example is the purification of recombinant L-selection receptor globulin from complex Chinese hamster ovary (CHO) cell medium, using a DNA aptamer directed against human L-selectin [3,39]. Single-step purification, using mild elution conditions, resulted in 83% recovery, with 1500-fold enrichment of the target protein. Immunoaffinity chromatography, on the other hand, would normally have required denaturing conditions for the recovery of the target molecule from the affinity column [14,15]. Modified-RNA aptamers to coagulation factor VIIa (FVIIa) exhibit a greater than 500-fold specificity for F VIIa relative to the coagulation factor Xa and greater than 1000-fold relative to coagulation factor IXa, although these proteins share a common set of structural domains [19,40]. Employing subtractive selection strategies can yield aptamers with an even greater ability to discriminate between the target and related nontargeted proteins [41,42]. Furthermore, because the selection process is performed in vitro, schemes to improve the specificity of a given aptamer for the target, or to direct the binding of the aptamer to a particular site on the target, are only limited by the knowledge of the target and the investigator’s imagination [5] (Table 1).
Indications and Factors Affecting Therapeutic Potency of Aptamers In Vivo
Given the current state of aptamer technology, the ideal indication for a therapeutic aptamer is an acute condition for which an important, spatially confined, an extracellular target has been identified and for which no good alternative therapies exist. In such a situation, local delivery of even a small dose of aptamer may be effective, both in terms of clinical response and cost. Aptamers targeting conditions that meet some or all of these criteria have made the most progress towards clinical utility. We will discuss three indications for which aptamers have progressed to animal and/or human models. By no coincidence, all three falls under the umbrella of vascular biology. For these common clinical conditions, several plasmas- or endothelium-based targets have been characterized. Drug delivery can thus be accomplished either intra vascularly or locally in the setting of surgery or percutaneous vascular intervention [5]. Although the selection of aptamers for therapeutic applications is relatively straightforward, the adaptation of aptamers for use in vivo has required extensive research since many factors are affecting its therapeutic potency. Relevant features of much protein therapeutics, such as those used in replacement therapies, can be extrapolated based on characteristics of known, circulating proteins. For example, therapeutic antibodies tend to have long circulating half-lives because of their large molecular mass and Neonatal Fc receptor (fcRn) r-mediated recycling. However, there are few circulating nucleic acids from which similar comparisons can be drawn [11,15]. Conversely, even though aptamers are chemicals, as outlined above, they are large by the standards of traditional non-protein drugs and do not readily cross biological barriers such as cell membranes. Most targets for therapeutic aptamers are either in solution in the blood plasma or displayed on the surface of cells that are accessible from the blood plasma, such as on the surface of the vasculature. Aptamers in this medium are subject to nuclease-mediated degradation by serum nucleases, renal filtration, and uptake by the liver and other tissues such as the spleen [14,15,19].
Application of Aptamers in Diagnostic and Therapeutic of Viral Infections
Appropriate diagnosis is the key factor for the treatment of viral diseases. Nevertheless, viral infections are difficult to distinguish, especially at the onset. If the acute infection appears, the patient presents the set of nonspecific signs and symptoms. Time is the most important factor in rapidly developing and epidemiologically dangerous diseases, such as influenza, Ebola, and SARS (Severe Acute Respiratory Syndrome) [43]. On the other hand, chronic viral diseases are asymptomatic or oligo symptomatic. The therapeutic success focused on organ protection from chronic destruction and failure, e.g., in HIV-1 (Human Immunodeficiency Virus 1) or HCV (Hepatitis C Virus), depends on early detection of an infective agent. It has been more than twenty years since aptamers were constructed [34,36]. Apart from the above cases where aptamers have been used for the discovery and delivery of drugs, the oligonucleotides themselves have also found use as therapeutic drugs. Many aptamers have completed various stages of preclinical development, some are in Phase I of clinical trial and one drug is already in the market. The therapeutic potential of single-stranded aptamers lies in the fact that they directly inhibit the function of the protein by folding into a specific three-dimensional structure that dictates high-affinity binding to the targeted protein. This property is being exploited to develop drugs against different validated protein targets. Some of these cases are discussed in detail below [14].
The Therapeutic Relevance of SELEX /Aptamers as Therapeutics
Aptamers can be used for therapeutic purposes in much the same way as monoclonal antibodies. However, unlike traditional methods for producing monoclonal antibodies, no organisms are required for the in vitro selection of oligonucleotides. This freedom from cellular biochemistry offers a huge advantage in manipulating the process of directed evolution. As per the investigation, chemistry, selection conditions, and targets can be manipulated in vitro in ways that would be difficult or impossible if organisms were involved [11,42]. Correspondingly, aptamers have a unique niche relative to other oligonucleotide therapeutics. For antisense oligonucleotides or siRNAs, the therapeutic target is intracellular, whereas aptamer therapeutics can be developed for intracellular, extracellular, or cell-surface targets. Targeting proteins in these latter two classes alleviate the necessity for the therapeutic to cross the cell membrane much like monoclonal antibodies [30]. Aptamers can theoretically be used therapeutically in any disease for which extracellular blockade of protein-protein interactions is required. The focus on extracellular targets has so far not been a limiting factor for aptamer development, as aptamers are currently undergoing clinical evaluation for ocular diseases, hematological diseases, and cancer [42,44]. Many aptamers that are selected to bind to a specific protein also inhibit its function. This is possible because protein active sites offer more exposed hetero atoms for hydrogen bonding and other interactions. Another explanation is that aptamers have a limited number of interactions that they can make with a protein target, and therefore aptamers that ‘fit’ into a crevice on a protein, such as an active site, are more likely to be selected known as homing principle [1,11].
Most therapeutically useful aptamers tend to inhibit proteinprotein interactions, such as receptor-ligand interactions, and thereby function as antagonists. However, at least some aptamers have been shown to have agonist-like activities. For example, aptamers isolated against the extracellular domain of the protein human epidermal growth factor receptor 3 (HER3; also known as ERBB3) can promote oligomerization (although this does not result in inhibition of downstream phosphorylation) [45]. By contrast, a DNA aptamer isolated against an isoleucyl tRNA synthetase enhanced editing activity [19,46]. For antagonists, therapeutic effects are only observed as long as the aptamer can physically dock with the target. If the binding affinity of the aptamer is high, the therapeutically relevant effect will likely be more prolonged. Therefore, as with many other drugs, the key features of aptamers that must be optimized for drug development include high affinity and specificity, and a long half-life in the relevant biological compartment. SELEX protocols have now been developed to increase target affinity (by decreasing off-rates) and to increase specificity. For example, specificity can be selected by using a specific sub domain of a protein as a target, or by immobilizing the target using a specific method or affinity tag 9. Iteratively toggling or switching between multiple targets during selection (for example, between the full-length protein and a specific sub domain) can drive binding to a particular epitope [47,48]. Negative selection against closely related targets together with positive selection against the desired target can reduce cross-binding that might elicit toxicity. Alternatively, sequential selection against different targets can be used to ensure that aptamers cross-bind to species homologs, yielding aptamers that can be used in both animal models as well as in humans [49,50].
Aptamers in the Treatment of Viral Infections
The life-threatening common viral diseases include HCV, HIV- 1, SARS, MERS (Middle East Respiratory Syndrome), and mentioned above avian flu variants, e.g., H5N1 [51,52]. The reason for inefficient medications and vaccines includes high virus mutation variability, low specificity, and avoiding the host immune response [51,53]. It also must be remembered, that many of the existing antiviral drugs, cause side effects and may lead to the development of other diseases than initially treated. They also interact with a variety of medicines, which may weaken or enhance their primary activity [54,55]. Many methods used in the treatment of viral infections have been only partially effective. For example, the standard treatment in HCV (with ribavirin and interferon-alpha) is effective in 50% of cases [42,56], whereas about 0.5 million patients die every year [57]. These problems should be the basis for searching new therapeutic tools, more effective and simultaneously less dangerous for patients’. One of the potentially promising solutions might be aptamers directed against any protein of the infected cells and any viral component [13,58].
Aptamers might be used not only to treat the infection but also to prevent it viral infection can be inhibited in almost any step of the disease. Many studies confirmed that the most effective therapeutic strategy is to block the penetration of viruses into the cells and/or inhibition of enzymes involved in their replication [59,60]. It is also believed that aptamers can selectively stimulate the immune system [61]. High variable viral genome regions are the common cause of virus resistance to currently used therapies. Thus, there is a requirement to generate aptamers specific to highly conserved nucleic acid regions, where mutations appear relatively rare. The most attention has been paid to HCV and HIV-1 infections, due to their prevalence, severe complications, and well-known therapeutic problems. Other viral diseases considered as aptamer targets include influenza, HSV (Herpes Simplex Virus) and HBV infections, i.e., the diseases, with commonly occurring immune antiviral response [10].
Anti-HIV1 Aptamers
Historically speaking, viral DNA polymerases have been the first target for aptamers [16,34] probably because they have an inherent capacity to bind to nucleic acids. It turns out that out of the large library that was used for selection, the best binders were the sequence of the wild-type version found in bacteriophage mRNA that interacts with the polymerase and a sequence bearing a high degree of homology to it [16]. Thus, evolution had already zeroed onto the best binder. This group was also the first to report the selection of an RNA aptamer against the HIV1 reverse transcriptase, which affects its replication capacity [35,62]. Small molecules have constituted the major weapon to fight HIV1. The development of resistant/mutant species and the inability of small molecules to inhibit the replication of HIV1 completely have forced scientists to look for other inhibitors. One approach has been to use oligo nucleotide-based strategies including antisense, RNAi, and aptamers [63]. The vector expresses a 937-base antisense gene that targets multiple sites on the HIV1 envelope, which circumvents the main problem of most of the anti-HIV strategies, namely that of mutation of the viral genome. Four of the five patients considered in this study, all of who had exhibited resistance to at least two antiviral therapies, showed an enhanced cellular response to HIV1, following the autologous transfection of transformed CD4 cells. In total, 60% of patients (three out of five) demonstrated improved memory retention in T cells. One patient actually exhibited improved antiviral effects. Studies with multiple doses of the drug are in progress. Therapeutics has conducted Phase I clinical trials with an HGTV43 gene vector, which helps to transfect three antisense genes directed against the genes responsible for HIV1 replication into hematopoietic stem cells of patients infected with HIV1 [1,11]. These engineered stem cells (after ex vivo gene therapy) were able to survive in vivo and to produce CD4+ cells that expressed anti-HIV1 antisense genes for more than 24 months after infusion. This is in contrast to ribozymes directed against specific sequences of the HIV1 genome discussed earlier, which exhibit transient expression [48,64]. Phase II trials will focus on increasing the fraction of CD4+ cells carrying the anti-HIV1 antisense genes. Because of the important role played by the enzyme reverse transcriptase (RT) in HIV1 replication, it has been a major target of all the above approaches. The reverse transcription reaction catalyzed by RT consists of more than one step and hence offers more than one site of inhibitory action by aptamers. A comparison of RNAi and aptamers for inhibition of RT activity has shown that at high multiplicity of infection, aptamers exhibit higher efficiency in blocking HIV1 replication [35].
Different studies have shown that this may be because of the ability of aptamers to be encapsulated in virion particles which allows them to inhibit two successive rounds of reverse transcription. The anti-RT aptamers are mostly classified as ‘primer/ template analogue RT inhibitors’ (TRTIs) as structural studies have revealed that the pseudoknot fold of the RNA aptamers competes with and binds at a cleft that overlaps with the primer/template binding site [65]. The aptamers were found to inhibit polymerase as well as RNAse H activities. As replication of the viral genome requires a precise synergy among all these steps, disruption of anyone stage causes a failure in the replication mechanism of HIV1. It has been shown that RNA pseudo knot aptamers raised against RT of HIV-1 B subtype can recognize and bind to recombinant RTs from phylogenetically diverse lenti viruses. The degree of inhibition however depended on the subfamily of the aptamer structure [14,34]. A second target is the surface glyco proteins of HIV1, which promote the fusion of the viral and host cell membranes, facilitating infection by HIV1. The selection of an aptamer has been carried out against the surface glycoprotein (gp120) as the target, by following the binding via surface Plasmon resonance [66]. This aptamer was able to protect human lymphocytes when challenged with HIV1. Surprisingly, even though the carbohydrate content of gp120 is implicated in viral infectivity, the binding of the aptamer to the glycoprotein was not dependent on the sugar residues, as it bound to the deglycosylated gp120 with a similar affinity as the glycosylated version [67,68].
Targeting Therapeutics with Aptamers
Aptamers that bind to the cell surface can specifically cause therapeutics (such as drugs, toxins, or siRNA) to persist in the vicinity of a specific cell or tissue type. This can also potentially cause an increase in the rate of internalization into cells by receptormediated endocytosis; for example, by acting as escort molecules to deliver intracellular therapeutics [14,69]. One popular epitope that has been targeted for therapy has been prostate-specific membrane antigen (PSmA), a protein that is widely expressed but rarely found on the cell surface. PSmA is observed on the surface of some prostate cancer cells [70,71]. The Coffey group at Johns Hopkins University School of medicine, Maryland, USA, selected two aptamers against this receptor, termed A9 and A10, that demonstrate low nanomolar Integrated Circuit (IC) 50 values for this target [72]. As PSmA is constitutively internalized, the PSmA-specific aptamer was also an excellent candidate as an escort aptamer that could mediate delivery via endocytosis [3]. Conventional small-molecule therapeutics have been delivered by aptamers. Doxorubicin, an anthracycline drug, is widely used for anticancer treatment and is well known to interact with the double helix of DNA. Doxorubicin has been directly bound to the PSmA specific aptamer A10 and delivered to cells [73]. Other aptamers, including the PTK7-specific sgc8c, have also been conjugated to doxorubicin and have proved their utility as vectors for drug delivery [74]. A more innovative application involves the generation of phototoxic aptamers for the targeted therapy of specific cancer cells [75].
DNA aptamers selected against short O-glycan peptides specifically expressed only on the surface of cancer cells were modified at their 5′-ends with chlorin e6, a photodynamic agent, and internalized into epithelial tumor cells. Light-activated cytotoxicity relative to the drug alone was found to be enhanced 500-fold and resulted in the tissue-specific killing of cancer cells. The delivery of biopolymer therapeutics has also been examined. Gelonin is a toxin that has been conjugated to antibodies or other proteins for delivery to tumor cells, and as it lacks a translocation domain in the absence of conjugation it has little inherent cytotoxicity [2,76]. Gelonin conjugated to the PSmA-specific aptamer A9 can target and specifically destroy PSmA-overexpressing prostate cancer cells. The conjugates have an increased potency of at least 600- fold relative to cells that do not express PSmA [77]. Interestingly, DNA aptamers selected against short O-glycan peptides specifically expressed only on the surface of cancer cells were modified at their 5′-ends with chlorin e6, a photodynamic agent, and internalized into epithelial tumor cells. Light-activated cytotoxicity relative to the drug alone was found to be enhanced 500-fold and resulted in the tissue-specific killing of cancer cells. The delivery of biopolymer therapeutics has also been examined. Gelonin is a toxin that has been conjugated to antibodies or other proteins for delivery to tumor cells, and as it lacks a translocation domain in the absence of conjugation it has little inherent cytotoxicity [2,76]. Gelonin conjugated to the PSmA-specific aptamer A9 can target and specifically destroy PSmA-overexpressing prostate cancer cells. The conjugates have an increased potency of at least 600- fold relative to cells that do not express PSmA [77]. Interestingly, Several standard options for improving siRNA processing were introduced into the original construct, and the identification of a cell line amenable to uptake enabled the demonstration of efficacy after systemic administration [81]. Aptamer-siRNA chimeras have also been generated by fusing the HIV gp120-specific aptamer and a tat/revspecific siRNA. This construct targets cells infected with HIV-1, and can inhibit HIV replication through both the aptamer and the siRNA components [27]. Finally, aptamers can be used to direct the delivery of supra molecular structures using the PSmA-specific aptamer A10. Farokhzad, Langer and their co-workers directed the delivery of nano particles to tumor cells [82]. Farokhzad’s group further delivered an encapsulated cancer drug, docetaxel, to LNCaP xenografts in nude mice. All seven of the mice treated by intra-tumoral injection survived and showed reductions in tumor volume, compared with the survival of only four out of seven mice in a control group that was treated with drug encapsulated nanoparticles lacking the PSmA-specific aptamer. Some efficacy was also demonstrated with the chemotherapeutic drug cisplatin delivered to tumor cells via aptamer functionalized PLGA-PEG nanoparticle conjugates [18,83]. Similarly, a nucleolin-specific aptamer was conjugated to a liposome that encased the potent chemotherapeutic cisplatin and led to the delivery of the drug to tumors. Furthermore, the controlled release of the conjugate to tumors was achieved by using an antisense oligonucleotide. These targeted encapsulation strategies may decrease systemic toxicity normally associated with chemotherapies [84].
Anticancer Aptamer
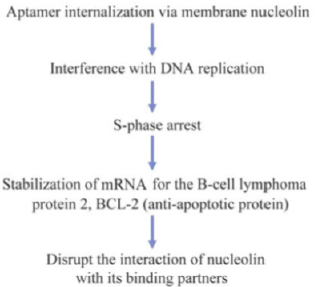
The anticancer aptamer AGRO100, now known as AS1411, belongs to the class of molecules referred to as guanosine rich oligonucleotides (GRO) [85,86]. Some of these form stable G-quadruplex structures, which allow them to exert nonantisense effects on cell proliferation in vitro. The inhibition mechanism is postulated to be via GRO-mediated arrest of DNA replication in the S-phase [87,88]. This recognition is mediated by shape complementarity between the cellular target and the oligonucleotide and to that extent, GROs bind to their targets as aptamers. A GRO100 is an unmodified 26 nucleotide long sequence, which dimerizes and self-anneals to form a quartet structure [89]. The authors commenced the selection process with four GROs, one of which exhibited clear G-quartet formation (as seen by UV-melting studies). The first step was to monitor the antiproliferation activity of the four GROs in cell cultures. One of them (which contained the G-quartet structure) exhibited enhanced inhibition of cell growth. Radioactive labeling isolated the target site as nucleolin, a multifunctional protein that is normally intracellular but is over expressed on the surface of cancer cells [90].
This oligonucleotide, after suitable truncation, was labeled as AGRO100. It was also found to recognize the NF-κ B essential modulator (NEMO), an inhibitor of κ B kinase γ. Members of the NF-κ B family function as dimeric transcription factors regulating the expression of genes involved in many areas including cell growth, differentiation, and apoptosis, and hence are good targets for anticancer therapy [91]. Binding of AGRO100 led to internalization of the complex formed between the aptamer, nucleolin, and NEMO, which probably inhibits NF-κ B activation and induces cell death. Administration of AGRO100 to various human-derived cancer cell lines inhibited NF-κ B transcriptional activity, both constitutive as well as that induced by TNF-α, in a dose-dependent manner [86]. This kind of behavior is not limited by cancer type. AGRO100 is the first aptamer to be tested in humans for the treatment of cancer. In total, 67% of the patients exhibited a stable disease profile after the Phase I trial [92,93]. The drug was administered intravenously and toxicity was assessed after regular intervals. Of the 17 patient’s understudy who were suffering from advanced cancer, more than half exhibited a stable disease profile two months after treatment commenced. The absence of side effects is one crucial feature that distinguishes aptamers from other drugs that target malignant growth [14,48] (Figure 4).
Antithrombotic Aptamers
Coagulation and Thrombosis
The delicate balance between hemostasis and hemorrhage is maintained by a complex system of plasma, cellular, and endothelial factors. Coagulation, the normal process by which a fibrin clot is generated in response to a vascular injury, is to be distinguished from thrombosis, the pathological formation of a clot in response to injury, stasis, or hyper coagulability. The latter is widespread in conditions such as acute coronary syndrome, stroke, peripheral vascular disease, and deep vein thrombosis, and can also occur in response to iatrogenic vascular injury. The clinical demand for more and better antithrombotic, which inhibit the initial formation of the platelet plug, and anticoagulants, which inhibit the cascade of reactions leading to the cross linking of fibrin, is testimony to the importance of this process. Also, the variety of patients and scenarios in which such agents are utilized requires an array of inhibitors with different mechanisms, properties, and toxicity profiles [3,5]. Thrombin is the most obvious target for the generation of both anticoagulant and antithrombotic compounds. Bock et al., [94] generated a 15 nucleotide DNA based thrombin aptamer that binds thrombin with a moderate affinity (apparent Kd ∼10-7 M) and can prolong the clotting time of human plasma. To take advantage of its rapid clearance (in vivo half-life of approximately 1-2 minutes), the thrombin DNA aptamer was developed largely as an anticoagulant for use in surgical indications requiring regional anticoagulation of an extracorporeal circuit. When administered by constant infusion, this molecule was successfully used to maintain the patency of an extracorporeal circuit in sheep and was substituted for heparin in a canine cardiopulmonary bypass model [95,96].
Furthermore, because of its rapid clearance, once the infusion of the aptamer was stopped, no reversal of the anticoagulant activity of this molecule was required. Based on its ability to inhibit clot-bound thrombin and platelet thrombus formation in an ex vivo whole artery angioplasty model, this aptamer also exhibited potential as a novel antithrombotic [97]. However, for this thrombin aptamer to be successfully tested in animal models of arterial thrombosis, it would probably be necessary to modify it to improve its circulating half-life. In addition to thrombin, most of the serine proteases and their cofactors in the coagulation pathway are suitable targets for the generation of experimental anticoagulants or antithrombotic [1,11]. Novel inhibitors of these proteins may also prove useful for the experimental dissection of thrombus formation in different pathologic states. Physiologic coagulation is initiated by the enzymatic activity of the protease FVIIa in complex with its requisite cofactor, tissue factor. Recently, a 2′-amino modified-RNA aptamer has been generated to coagulation FVIIa. This aptamer binds FVIIa with high affinity and specificity and has been shown to block the activity of FVIIa by preventing the formation of an active FVIIa-tissue factor complex. This aptamer is stable in plasma, with a half-life of approximately 15 hours, and can prolong the clotting time of human plasma. While this aptamer has not been studied in animals, it highlights the potential use of this technology to generate a panel of antagonists to the factors that compose the coagulation pathway [40] (Figure 5).
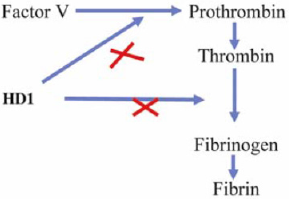
Figure 5: HD1 block fibrinogen and thrombin interaction and compete for pro-thrombin binding (Nahar et al; 2017).
Future Perspectives of Aptamers in Diagnostic Procedures
Aptamers are a promising class of compounds, both for target validation and therapy. As designer drugs, they exhibit high specificity, high affinity, and modifiable bioavailability. The ability to generate inhibitors with such properties against a variety of target proteins will be invaluable as the human genome and proteome are deciphered. Further animal and, in turn, human data are now necessary to facilitate the transition of these molecules from lab reagents to pharmaceuticals. Safety and efficacy in appropriate disease models must be established. Such development has been slowed by medium- and large-scale synthesis costs; however, these have been decreasing as a function of improved production capability. As has been witnessed with other new technologies, production capability will continue to improve if demand exists. Meanwhile, progress toward developing therapeutic aptamers will be expedited through collaborations between industry and academic researchers [5,15].
Due to the rapid growth of population and different varieties of viruses that are frequently resistant to standard treatment, there is an urgent need to develop the new diagnostic methods, characterized by high sensitivity and specificity, allowing for early and rapid pathogen detection. Biosensor technology is probably the most rapidly growing area of the current diagnosis of viral diseases. Aptamers are a perfect example of a molecular recognition biosensor element. The production cost is low, in comparison to monoclonal antibodies. They also provide sensitivity and specificity of the constructed biosensors. It is believed that aptamer-based biosensors could have been applied as a promising approach to some specific issues. They could be used for cheap diagnosis at an early stage of the disease, i.e., immediately after exposure to the pathogen, as well as to monitor the treatment process. SELEX versatility and its susceptibility to modifications, as enhancement of the selective pressure, could allow obtaining aptamers detecting precisely the level of viremia, which is below the threshold of the currently used diagnostic methods. Moreover, an aptamer-based approach ensures a more rapid and cheaper diagnosis [10]. The aptamer structure might be a major constraint in their future application as a diagnostic tool. Many existing aptamers are RNA molecules that are highly susceptible to degradation by nucleases. Consequently, their use as a molecular-recognition element of the biosensors may be limited. This problem can be solved by synthesizing a “mirror” analog of these particles that retains their original properties but are not cleaved by nucleases [98]. The alternative solution includes local modifications of the ribose 2’ sites in the aptamer chain [99,100]. In conclusion, aptamers are molecules potentially attractive for viral diagnostics including the recent pandemic Corona virus disease 19 (COVID-19).
Challenges in Application of Aptamer
The challenges facing aptamer technology are not entirely unique. Some valuable lessons about oligonucleotide pharmacology have been learned from antisense, ribozyme (see Usman and Blatt, this Perspective series [101] and Deoxyribozyme (DNAzyme) molecules [102]. Sequence-independent, nonspecific effects of the phosphorothioate backbones commonly used for antisense molecules have been observed at the micromolar levels required for efficacy [103]. In contrast, most aptamers currently feature phosphodiester backbones with modified sugars to enhance nuclease resistance. Although comprehensive toxicity profiles for these molecules have not been published, no untoward effects have been reported in several animal studies in the literature. Furthermore, the high affinity of most aptamers should allow for sub micromolar therapeutic levels with less potential for nonspecific effects. While the plasma pharmacokinetics of aptamers is gradually being elucidated, the efficient delivery of aptamers remains a consideration for the inhibition of intracellular targets. Methods adopted from other molecular therapeutic strategies may ultimately prove useful for the delivery of aptamers into cells, such as incorporation into liposomes or generation of highly effective concentrations using various means of local delivery [3,5,14].
As one advantage over peptide-based inhibitors, aptamers are purported to be non immunogenic. Their small size and similarity to endogenous molecules theoretically make them poor antigens. Experiments designed with the intent to elicit a humoral response to modified-RNA aptamers have failed to generate a specific response, despite the generation of responses to a variety of adjuvants employed However, the presence of antibodies against nucleic acids in autoimmune diseases makes the possibility of antiaptamer antibodies one that is plausible and worthy of investigation [11]. The great specificity that aptamers can achieve represents both strength and a potential weakness of the technology. Being too specific may be problematic if species cross-reactivity is desired, as is often the case during the preclinical evaluation of a molecule in animal models. Obviously, the more conserved the protein, the less likely this situation is to occur. With knowledge of the target, subtractive selection schemes may direct the aptamer to a conserved portion of the molecule. Alternatively, this problem may be obviated by driving the selection with targets from different species either in parallel or alternating in series. Finally, the expense of synthesis is not always considered in the early developmental stages of a novel therapeutic agent. While small-scale synthesis is feasible for most in vitro applications, the scale of synthesis necessary for the transition of a candidate aptamer to animal or even human models may be cost-prohibitive for even a well-funded academic laboratory [5,14].
Conclusion
Aptamers are single-chained RNA or DNA oligonucleotides (ODNs) with a three-dimensional conformation that provides the ability to fi their targets with high affinity and specificity obtained by a method called SELEX. Cancer immunotherapy has nowadays come back to prominence due to its encouraging results in the clinic with monoclonal antibodies. Aptamers display some important advantages over antibodies at the time of translation into the clinic. They are very suitable for targeting and delivery, reducing off target side effects, and increasing the therapeutic index of a given strategy. Several properties of aptamers make them an attractive class of therapeutic compounds. Their affinity and specificity for a given protein make it possible to isolate a ligand to virtually any target, and adjusting their bioavailability expands their clinical utility. The ability to develop aptamers that retain activity in multiple organisms facilitates preclinical development. Antidote control of aptamer activity enables safe, tightly controlled therapeutics. Aptamers may prove useful in the treatment of a wide variety of human maladies, including infectious diseases, cancer, and cardiovascular disease. Hundreds of aptamers have been described for very different purposes within biomedical research. Some of the aptamers described recently have been isolated with immunotherapeutic applications to overcome current challenges in cancer immunotherapy. To elicit a specific antitumor immune response, some of these aptamers are engineered to activate costimulatory receptors or blocking immunosuppressive signals. Aptamers would hopefully gain an important niche in cancer immunotherapy due to their specific properties.
References
- Nahar N, R Bin, MS, Rahman (2017) Applications of Aptamers in Medicine: A Mini Review. Bangladesh Pharmaceutical Journal 20(1): 99-104.
- Anthony D (2010) Aptamer as therapeutics. Nat Rev Drug Discov 9: 537-530.
- Yan AC, Levy M (2009) Aptamers and aptamer targeted delivery. RNA biology 6(3): 316-320.
- Rimmele M (2003) Nucleic acid aptamers as tools and drugs: recent developments. Chembiochem 4(10): 963-971.
- White RR (2000) Developing aptamers into therapeutics. The Journal of clinical investigation 106(8): 929-934.
- Kurreck J (2003) Antisense technologies: improvement through novel chemical modifications. European Journal of Biochemistry 270(8): 1628-1644.
- Shi H, X Chun (2007) RNA aptamers directed to discrete functional sites on a single protein structural domain. Proceedings of the National Academy of Sciences 104(10): 3742-3746.
- Zhao X, Hua S, Aarti S, Nicole L, Hillary C, et al. (2006) An RNA aptamer that interferes with the DNA binding of the HSF transcription activator. Nucleic acids research 34(13): 3755-3761.
- Hallegger M (2006) RNA aptamers binding the double-stranded RNA-binding domain. Rna 12(11): 1993-2004.
- Wandtke T (2015) Aptamers in diagnostics and treatment of viral infections. Viruses 7(2): 751-780.
- Nimjee SM (2017) Aptamers as therapeutics. Annual review of pharmacology and toxicology 57: 61-79.
- Peng L (2007) A combined atomic force/fluorescence microscopy technique to select aptamers in a single cycle from a small pool of random oligonucleotides. Microscopy research and technique 70(4): 372-381.
- Liu C, Qiong Z, Ying Z, Linda V, Steve P, et al. (2020) Research and development on therapeutic agents and vaccines for COVID-19 and related human coronavirus diseases. ACS Publications 6(3): 315-331.
- Keefe AD (2010) Aptamers as therapeutics. Nature reviews Drug discovery 9: 537-550.
- Kaur G, Roy I (2008) Therapeutic applications of aptamers. Expert opinion on investigational drugs 17(1): 43-60.
- Tuerk C, Gold L (1990) Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 249(4968): 505-510.
- Burmeister PE, Scott D, Robert F, Jeffrey R, Lilian R, et al. (2005) Direct in vitro selection of a 2′-O-methyl aptamer to VEGF. Chemistry & biology 12(1): 25-33.
- Chu TC (2006) Aptamer mediated siRNA delivery. Nucleic acids research 34(10): e73-e73.
- Sundaram P (2013) Therapeutic RNA aptamers in clinical trials. European Journal of Pharmaceutical Sciences 48(1-2): 259-271.
- Nimjee SM (2005) Aptamers: an emerging class of therapeutics. Annu. Rev. Med 56: 555-583.
- Szpechciński A, Grzanka A (2006) Aptamers in clinical diagnostics. Postepy biochemii 52(3): 260-270.
- Ng EW, David T, Perry C, Emmett T, David R, et al. (2006) Pegaptanib, a targeted anti-VEGF aptamer for ocular vascular disease. Nature reviews drug discovery 5(2): 123-132.
- Gilbert JC, Tia D, Renta M, J Horvath, G Merlino, et al. (2007) Pharmacokinetics, Pharmacodynamics, and Safety of an Anti-von Willebrand Factor Therapeutic Aptamer, ARC1779, in Healthy Volunteers. Am Heart Association 116(16).
- Jilma B, Petra P, Petra J, James C, Renta H, et al. (2010) A randomised pilot trial of the anti-von Willebrand factor aptamer ARC1779 in patients with type 2b von Willebrand disease. Thrombosis and haemostasis 104(3): 563-570.
- Laber DA, BS Taft, GH Kloecker, PJ Bates, JO Trent, et al. (2006) Extended phase I study of AS1411 in renal and non-small cell lung cancers. Journal of Clinical Oncology 24(18): 13098-13098.
- Bai H (2012) A SPR aptasensor for detection of avian influenza virus H5N1. Sensors 12(9): 12506-12518.
- Zhou J (2008) Novel dual inhibitory function aptamer-siRNA delivery system for HIV-1 therapy. Molecular Therapy 16(8): 1481-1489.
- Cullen BR, Greene WC (1989) Regulatory pathways governing HIV-1 replication. Cell 58(3): 423-426.
- Marciniak RA (1990) Identification and characterization of a HeLa nuclear protein that specifically binds to the trans-activation-response (TAR) element of human immunodeficiency virus. Proceedings of the National Academy of Sciences 87(9): 3624-3628.
- Nissim A, Chernajovsky Y (2008) Historical development of monoclonal antibody therapeutics. In Therapeutic Antibodies. Springer (181): 3-18.
- Burgert HG (2002) Subversion of host defense mechanisms by adenoviruses. In Viral Proteins Counteracting Host Defenses, Springer (269): 273-318.
- Sullenger BA (1990) Overexpression of TAR sequences renders cells resistant to human immunodeficiency virus replication. Cell 63(6): 601-608.
- Sullenger BA (1991) Analysis of trans-acting response decoy RNA-mediated inhibition of human immunodeficiency virus type 1 transactivation. Journal of virology 65(12): 6811-6816.
- James W (2007) Aptamers in the virologists' toolkit. Journal of General Virology 88(2): 351-364.
- Joshi PJ (2005) Aptamers directed to HIV-1 reverse transcriptase display greater efficacy over small hairpin RNAs targeted to viral RNA in blocking HIV-1 replication. Molecular Therapy 11(5): 677-686.
- Ellington AD Szostak JW (1990) In vitro selection of RNA molecules that bind specific ligands. nature 346: 818-822.
- Gold L (1995) Diversity of oligonucleotide functions. Annual review of biochemistry 64: 763-797.
- Conrad R (1994) Isozyme-specific inhibition of protein kinase C by RNA aptamers. Journal of Biological Chemistry 269(51): 32051-32054.
- Romig TS (1999) Aptamer affinity chromatography: combinatorial chemistry applied to protein purification. Journal of Chromatography B: Biomedical Sciences and Applications 731: 275-284.
- Rusconi CP (2000) Blocking the initiation of coagulation by RNA aptamers to factor VIIa. Thrombosis and haemostasis 84(5): 841-848.
- Doudna JA (1995) Selection of an RNA molecule that mimics a major autoantigenic epitope of human insulin receptor. Proceedings of the National Academy of Sciences 92(6): 2355-2359.
- Lee JF (2006) Aptamer therapeutics advance. Current opinion in chemical biology 10(3): 282-289.
- Fletcher SJ (2010) Toward specific detection of Dengue virus serotypes using a novel modular biosensor. Biosensors and Bioelectronics 26: 1696-1700.
- Keefe AD, Schaub RG (2008) Aptamers as candidate therapeutics for cardiovascular indications. Current opinion in pharmacology 8: 147-152.
- Chi-hong BC (2003) Inhibition of heregulin signaling by an aptamer that preferentially binds to the oligomeric form of human epidermal growth factor receptor-3. Proceedings of the National Academy of Sciences 100(16): 9226-9231.
- Hale SP, Schimmel P (1996) Protein synthesis editing by a DNA aptamer. Proceedings of the National Academy of Sciences 93(7): 2755-2758.
- Diener J (2009) Inhibition of von Willebrand factor‐mediated platelet activation and thrombosis by the anti‐von Willebrand factor A1‐domain aptamer ARC1779. Journal of Thrombosis and Haemostasis 7(7): 1155-1162.
- Thiel KW, Giangrande PH (2009) Therapeutic applications of DNA and RNA aptamers. Oligonucleotides 19: 209-222.
- White R (2001) Generation of species cross-reactive aptamers using “toggle” SELEX. Molecular Therapy 4(6): 567-573.
- White RR (2003) Inhibition of rat corneal angiogenesis by a nuclease-resistant RNA aptamer specific for angiopoietin-2. Proceedings of the National Academy of Sciences 100(9): 5028-5033.
- Dunning J (2014) Antiviral combinations for severe influenza. The Lancet infectious diseases 14(12): 1259-1270.
- Nicholson KG (2003) Influenza. Lancet 362: 1733-1745.
- Sahu GK (2015) Potential implication of residual viremia in patients on effective antiretroviral therapy. AIDS research and human retroviruses 31(1): 25-35.
- Soriano V (2015) Drug interactions with new hepatitis C oral drugs. Expert Opinion on Drug Metabolism & Toxicology 11(3): 333-341.
- Zornitzki T, Stephen M (2015) Interferon therapy in hepatitis C leading to chronic type 1 diabetes. World Journal of Gastroenterology: WJG 21(1): 233-239.
- Manns MP, SC Gordan (2001) Peginterferon alfa-2b plus ribavirin compared with interferon alfa-2b plus ribavirin for initial treatment of chronic hepatitis C: a randomised trial. The Lancet 358(9286): 958-965.
- Lozano R (2012) Global and regional mortality from 235 causes of death for 20 age groups in 1990 and 2010: a systematic analysis for the Global Burden of Disease Study 2010. The lancet 380: 2095-2128.
- Mufhandu HT (2012) UCLA1, a synthetic derivative of a gp120 RNA aptamer, inhibits entry of human immunodeficiency virus type 1 subtype C. Journal of virology 86(9): 4989-4999.
- Bellecave P, et al. (2008) Inhibition of hepatitis C virus (HCV) RNA polymerase by DNA aptamers: mechanism of inhibition of in vitro RNA synthesis and effect on HCV-infected cells. Antimicrobial agents and chemotherapy 52: 2097-2110.
- Gopinath SC, Kyoko H (2012) Aptamer that binds to the gD protein of herpes simplex virus 1 and efficiently inhibits viral entry. Journal of virology 86(12): 6732-6744.
- Hwang SY, H Young (2012) 5′-Triphosphate-RNA-independent activation of RIG-I via RNA aptamer with enhanced antiviral activity. Nucleic acids research 40(6): 2724-2733.
- Tuerk C, L Gold (1992) RNA pseudoknots that inhibit human immunodeficiency virus type 1 reverse transcriptase. Proceedings of the National Academy of Sciences 89(15): 6988-6992.
- Levine BL, Jean B, R Roy, Tessio R (2006) Gene transfer in humans using a conditionally replicating lentiviral vector. Proceedings of the National Academy of Sciences 103(46): 17372-17377.
- Weerasinghe M, SE Lim(1991) Resistance to human immunodeficiency virus type 1 (HIV-1) infection in human CD4+ lymphocyte-derived cell lines conferred by using retroviral vectors expressing an HIV-1 RNA-specific ribozyme. Journal of virology 65(10): 5531-5534.
- Fisher TS (2002) Mutations that confer resistance to template-analog inhibitors of human immunodeficiency virus (HIV) type 1 reverse transcriptase lead to severe defects in HIV replication. Journal of virology 76(8): 4068-4072.
- Khati M (2003) Neutralization of infectivity of diverse R5 clinical isolates of human immunodeficiency virus type 1 by gp120-binding 2′ F-RNA aptamers. Journal of virology 77(23): 12692-12698.
- Dey AK, Carla G (2005) Structural characterization of an anti-gp120 RNA aptamer that neutralizes R5 strains of HIV-1. Rna 11(6): 873-884.
- Dey AK (2005) An aptamer that neutralizes R5 strains of human immunodeficiency virus type 1 blocks gp120-CCR5 interaction. Journal of virology 79(21): 13806-13810.
- Hicke BJ, Stephens AW (2000) Escort aptamers: a delivery service for diagnosis and therapy. The Journal of clinical investigation 106(8): 923-928.
- Chang SS (2004) Overview of prostate-specific membrane antigen. Reviews in urology 6: S13.
- Chang SS, Heston WD (2002) The clinical role of prostate-specific membrane antigen (PSMA). In Urologic Oncology: Seminars and Original Investigations, Elsevier 7(1): 7-12.
- Lupold SE, Brian J (2002) Identification and characterization of nuclease-stabilized RNA molecules that bind human prostate cancer cells via the prostate-specific membrane antigen. Cancer research 62(14): 4029-4033.
- Bagalkot V (2006) An aptamer–doxorubicin physical conjugate as a novel targeted drug‐delivery platform. Angewandte chemie international edition 45(48): 8149-8152.
- Huang YF (2009) Molecular assembly of an aptamer–drug conjugate for targeted drug delivery to tumor cells. ChemBioChem 10(5): 862-868.
- Ferreira CS, Melissa C (2009) Phototoxic aptamers selectively enter and kill epithelial cancer cells. Nucleic acids research 37(3): 866-876.
- Better M (1994) Gelonin analogs with engineered cysteine residues form antibody immunoconjugates with unique properties. Journal of Biological Chemistry 269(13): 9644-9650.
- Chu TC, John, Laura (2006) Aptamer: toxin conjugates that specifically target prostate tumor cells. Cancer research 66(12): 5989-5992.
- Cullen BR (2002) RNA interference: antiviral defense and genetic tool. Nature immunology 3(7): 597-599.
- Sioud M (2005) On the delivery of small interfering RNAs into mammalian cells. Expert Opinion on Drug Delivery 2(4): 639-651.
- McNamara JO (2006) Cell type-specific delivery of siRNAs with aptamer-siRNA chimeras. Nature biotechnology 24(8): 1005-1015.
- Dassie JP, Xiu, Gregory, Ryan (2009) Systemic administration of optimized aptamer-siRNA chimeras promotes regression of PSMA-expressing tumors. Nature biotechnology 27: 839-846.
- Farokhzad OC (2004) Nanoparticle-aptamer bioconjugates: a new approach for targeting prostate cancer cells. Cancer research 64(21): 7668-7672.
- Dhar S (2008) Targeted delivery of cisplatin to prostate cancer cells by aptamer functionalized Pt (IV) prodrug-PLGA–PEG nanoparticles. Proceedings of the National Academy of Sciences 105: 17356-17361.
- Cao Z, Rong T, A Mishra (2009) Reversible cell‐specific drug delivery with aptamer‐functionalized liposomes. Angewandte Chemie International Edition 48(35): 6494-6498.
- Bates PJ (1999) Antiproliferative activity of G-rich oligonucleotides correlates with protein binding. Journal of Biological Chemistry 274: 26369-26377.
- Girvan, AC (2006) AGRO100 inhibits activation of nuclear factor-κB (NF-κB) by forming a complex with NF-κB essential modulator (NEMO) and nucleolin. Molecular cancer therapeutics 5(7): 1790-1799.
- Lassalle HP, Sophie (2012) Aptamers as remarkable diagnostic and therapeutic agents in cancer treatment. Current drug metabolism 13(8): 1130-1144.
- Xu X (2001) Inhibition of DNA replication and induction of S phase cell cycle arrest by G-rich oligonucleotides. Journal of Biological Chemistry 276(46): 43221-43230.
- Đapić V, vedra (2003) Biophysical and biological properties of quadruplex oligodeoxyribonucleotides. Nucleic acids research 31(8): 2097-2107.
- Đapić V, Paula J (2002) Antiproliferative activity of G-quartet-forming oligonucleotides with backbone and sugar modifications. Biochemistry 41(11): 3676-3685.
- Zhou G (2016) Aptamers: A promising chemical antibody for cancer therapy. Oncotarget 7: 13446.
- Laber DA, MA Choudary (2004) A phase I study of AGRO100 in advanced cancer. Journal of Clinical Oncology 22(14): 3112-3112.
- Soldevilla MM (2017) Aptamers as a Promising Therapeutic Tool for Cancer Immunotherapy. Immunotherapy: Myths, Reality, Ideas, Future 129-150.
- Bock LC, Griffin, Latham A (1992) Selection of single-stranded DNA molecules that bind and inhibit human thrombin. nature 355(6360): 564-566.
- DeAnda Jr A (1994) Pilot study of the efficacy of a thrombin inhibitor for use during cardiopulmonary bypass. The Annals of thoracic surgery 58(2): 344-350.
- Griffin LC (1993) In vivo anticoagulant properties of a novel nucleotide-based thrombin inhibitor and demonstration of regional anticoagulation in extracorporeal circuits.
- Li WX, Kaplan V (1994) A novel nucleotide-based thrombin inhibitor inhibits clot-bound thrombin and reduces arterial platelet thrombus formation. Blood 83(3): 677-682.
- Eulberg D, Klussmann, S (2003) Spiegelmers: biostable aptamers. Chembiochem 4: 979-983.
- Mendonsa SD, Bowser MT (2004) In vitro evolution of functional DNA using capillary electrophoresis. Journal of the American Chemical Society 126(1): 20-21.
- Pagratis NC (1997) Potent 2′-amino-and 2′-fluoro-2′-deoxyribonucleotide RNA inhibitors of keratinocyte growth factor. Nature biotechnology 15: 68-73.
- Usman N, Blatt LM (2000) Nuclease-resistant synthetic ribozymes: developing a new class of therapeutics. The Journal of clinical investigation 106(10): 1197-1202.
- Khachigian LM (2000) Catalytic DNAs as potential therapeutic agents and sequence-specific molecular tools to dissect biological function. The Journal of clinical investigation 106(10): 1189-1195.
- Srinivasan SK, Iversen P (1995) Review of in vivo pharmacokinetics and toxicology of phosphorothioate oligonucleotides. Journal of clinical laboratory analysis 9(2): 129-137.

 Review Article
Review Article