Abstract
Biotinylated-antibodies are one type of conjugated critical reagent commonly used in Ligand Binding Assays (LBAs). The antibody degree of biotinylation affects the outcomes in assay functional performance. Usually, the amount of antibodies available for preparation of critical reagents is very limited and therefore alternative purification processes of smaller batches became more attractive. Immobilized Metal Affinity Chromatography (IMAC), is a well-known method for purification of proteins with histidine-clusters, such as the Fc domain of immunoglobulin-type antibodies. Here we discuss the feasibility to conjugate and characterize smaller amounts of humanized monoclonal antibodies for screening purposes in LBAs. Our data shows that as little as 0.25mg of protein are efficiently purified using IMAC in a time effective manner. Initial steps of LBAs development should consider the screening of optimal degree of labelling and therefore the testing of batches conjugated at different degrees of labelling purified by IMAC is a straightforward approach.
Keywords: Conjugated Critical Reagents; Biotin; Immobilized Metal Affinity Chromatography
Introduction
Critical reagents are essential tools in Ligand Binding Assays
(LBAs). They can range from peptides to larger molecules such
as antibodies, which may be conjugated [1]. Several types of
conjugation tags are available depending on the assay purpose and
chemistry of conjugation. Among them, biotin is a versatile and
popular label that binds streptavidin (or avidin) with high affinity
[2]. For instance, biotinylated Monoclonal Antibodies (mAbs) are
widely used for the development of LBAs [3]. Characterization
of conjugated critical reagents is a hot topic in the bioanalytical
industry [4,5]. Bioanalytical industry leaders have recently released
guidance on this subject and agree that thorough characterization
of critical reagents at the beginning of a drug candidate’s life cycle
is vital for the long-term success of that program [6]. The expected
level of characterization includes, but is not limited to, analysis of
biophysical properties, including the protein concentration, purity,
DoL, and analysis of functional properties, such as orthogonal
methods that relate to the LBA purpose.
Recently, more attention has been drawn to what should be
the optimal Degree of Labelling (DoL), i.e., the average number
of labels coupled to a protein molecule, for each assay. As an
example, in the context of an anti-drug antibody LBA, by optimizing
the biotin DoL, the authors reported an improvement on the
functional assay performance of ~60% [7]. Another interesting
finding was that lower and higher DoLs presented a decrease
in functional activity, suggesting that a balance between under
and over labelling, must exist. In a different study (LBA to detect
neutralizing antibodies), the authors observed a significant degree
of biotin-drug conjugate leaching was affecting the report of false
positive and false negative results [8]. Furthermore, optimizing the
antibody DoL mitigated that effect. After the antibody biotinylation process, a buffer exchange step is generally required to remove
the excess of unconjugated biotin, which might affect the assay
performance [7]. Common techniques include desalting columns,
size exclusion chromatography, ultra-filtration, tangential flow
filtration or dialysis, however larger amounts of proteins are
generally required. Taking into account that these antibodies might
be scarce and expensive, alternative purification techniques that
allow preparation of smaller amounts should be considered.
Immobilized Metal Affinity Chromatography (IMAC) is widely
used for purification of proteins with histidine tags or clusters [9].
Among the later, several classes of antibodies have been purified
using IMAC, including IgG1 type mAbs [10]. IMAC exploits the
binding affinity of his-tagged proteins to positively charged metal
ions (Zn, Cu, Ni, Co) [11]. The histidine rich region of both the
conjugated and unconjugated proteins binds to a volume of metal
beads in a positively charged resin and can be eluted by excess
of imidazole [12]. Here we extensively purified smaller batches
of conjugated mAbs by IMAC. These molecules include three
humanized monoclonal antibodies utilized in immunotherapy:
Trastuzumab, Rituxumab and Ofatumumab [13]. Each mAb was
biotinylated at different molar ratios and further characterized for
protein concentration, purity and degree of labelling. This strategy
allows easy generation of a toolset of conjugated critical reagents
with small amount of starting material, allowing the scale up of
conjugations and lower processing times.
Results
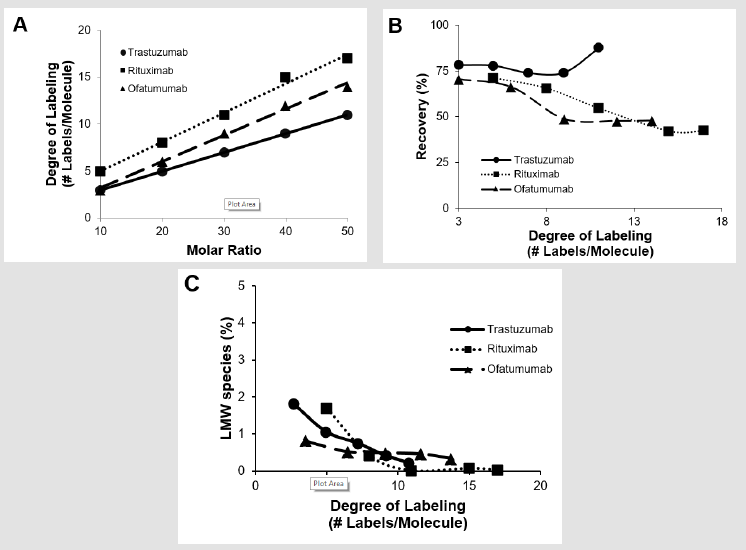
Figure 1: Biophysical properties of three biotinylated humanized mAbs (Trastuzumab, Rituximab and Ofatumumab) purified by IMAC.
a) Correlation between conjugation challenge ratio and incorporation ratio (determined by HRMS).
b) Protein recovery of the biotinylated molecules after IMAC purification.
c) Purity of LMW species determined by analytical SEC
A toolset of conjugated critical reagents was prepared to be used for method development purposes in LBAs. Here we conjugated three different humanized mAbs: Trastuzumab, Rituximab and Ofatumumab with biotin. These molecules were conjugated at distinct molar ratios of biotin/mAb (10:1, 20:1, 30:1, 40:1 and 50:1). Each reaction took place in a vial containing 0.25mg of protein with the respective amount of biotin, for 1 hour and subsequently quenched with tris buffer. After conjugation, all proteins were purified to remove the excess of unconjugated biotin using high-performance nickel-IMAC resin as described in the methods section. The DoL (number of labels per molecule) was assessed by high-resolution mass spectrometry (HRMS). As expected, increasing amounts of biotin in the reaction led to increasing amounts of biotin labels incorporated to each mAb (Figure 1A). Under the conditions used, the rate of incorporation fell between 0.2 to 0.31 labels incorporated / molar ratio / hour. Due to the versatility of the method, each set of mAbs was purified in parallel. The overall IMAC purification process took about one hour. After elution with an imidazole containing solution, the protein concentration was determined and the respective protein recovery calculated (Figure 1B).
The overall recovery of all five Trastuzumab-biotinylated molecules was around or over 75%. Both Rituximab and Ofatumumab conjugated molecules had higher recoveries (~75%) at lower degrees of labeling and lower recoveries (~50%) at lower degrees of labelling (>8 biotin labels / molecule). This data suggests that not all conjugated mAbs have the same affinity to the nickel coated resin due to factors such as accessibility of the histidine cluster or different degrees of glycosylation. The effectiveness of this method in removing the excess of non-conjugated biotin from each sample was also assessed. To that end, we tested each sample for purity using analytical Size Exclusion Chromatography (SEC). All samples had less than 2% of non-conjugated biotin (Figure 1C). Note that no high molecular weight aggregation was observed for any of the samples (data not shown). For a general method development strategy, the 2-to-3 different reagents conjugated at different degrees of labeling should be sufficient to better define the optimal critical reagent for the assay. We used 0.25mg of protein / conjugation and were able to recover sufficient reagent to perform full biophysical characterization.
Methods
Biotinylation – The mAbs were conjugated with EZ-Link™ NHSLC- LC-Biotin (Thermo Scientific) as per vendor instructions. Briefly, 0.25 mg was incubated with the respective amounts of biotin at the different molar ratios. The reactions were carried out for one hour at room temperature and stopped with Tris-HCl (7.4). IMACConjugated proteins (0.25 mg) were allowed to bind HisPur™ Ni-NTA Superflow Agarose (Thermo Scientific). Briefly, the Ni-NTA agarose was equilibrated with Phosphate Buffered Saline (PBS) in mini spin columns. The conjugated proteins were then incubated with the agarose at RT. After three washing steps with PBS, the conjugated critical reagent was eluted with imidazole buffer in PBS (7.4). and stored at 4ºC until further biophysical characterization. Protein concentration- Total protein concentration was assessed using the colorimetric bicinchoninic acid assay (Thermo Scientific). Bovine Gamma Globulin was used as standard protein. Protein Purity- Purity was assessed by analytical size exclusion chromatography as described in Rocha et al [7]. Degree of Labelling- The degree of labelling (or incorporation ratio) was determined by HRMS as previously described [7].
Conclusion
Development and optimization of bioanalytical methods is an essential phase of any clinical sample analysis program. One of the initial steps involves identifying, titrating and optimizing the critical reagent conditions in the assay. Since biophysical and functional aspects can dictate the performance of a given conjugated critical reagent, it is important to perform systematic upfront characterization. Therefore, preparing a toolset of critical reagents conjugated at different DoLs might be beneficial for proper method development. Here we demonstrate that IMAC is a powerful protein purification technique that allows purification of minimal biotinylated mAb amount. The amount of protein recovered was sufficient for downstream characterization. It is also important to highlight that this is a very time and cost effective workflow.
References
- King LE, Farley E, Imazato M, Keefe J, Khan M (2014) Ligand binding assay critical reagents and their stability: recommendations and best practices from the Global Bioanalysis Consortium Harmonization Team. 16 (3): 504-515.
- Weber PC, Ohlendorf DH, Wendoloski J, Salemme FJS (1989) Structural origins of high-affinity biotin binding to streptavidin 243(4887): 85-88.
- Diamandis EP,Christopoulos TKJC (1991) The biotin-(strept) avidin system: principles and applications in biotechnology 37(5): 625-636.
- O Hara DM, Theobald V, Egan AC, Usansky J, Krishna M (2012) Ligand binding assays in the 21st century laboratory: recommendations for characterization and supply of critical reagents 14(2): 316-328.
- Pihl S, van der Strate BW, Golob M, Vermet L, Jaitner B (2018) EBF recommendation on practical management of critical reagents for PK ligand-binding assays 10 (19): 1557-1565.
- Piccoli S, Mehta D, Vitaliti A, Allinson J, Amur S (2019) White Paper on Recent Issues in Bioanalysis: FDA Immunogenicity Guidance, Gene Therapy, Critical Reagents, Biomarkers and Flow Cytometry Validation (Part 3–Recommendations on 2019 FDA Immunogenicity Guidance, Gene Therapy Bioanalytical Challenges, Strategies for Critical Reagent Management, Biomarker Assay Validation, Flow Cytometry Validation & CLSI H62) 11(24): 2207-2244.
- Rocha AG, Krynski K, Mancino A, Sciscione M, Beaver CJJB (2019) New insights on critical reagent optimization for antidrug antibody assays. 11(9): 815-823.
- Xiang Y, Kamerud J, Donley J, Olson K, Caiazzo T (2019) Approaches to Resolve False Reporting in Neutralizing Antibody Assays Caused by Reagent Leaching from Affinity Capture Elution Solid Phase. 21(1): 4.
- Gaberc-Porekar V, Menart VJJ (2001) Perspectives of immobilized-metal affinity chromatography. 49(1-3): 335-360.
- Bresolin I, Borsoi Ribeiro M, Tamashiro W, Augusto E, Vijayalakshmi M(2010) Biotechnology, Evaluation of immobilized metal-ion affinity chromatography (IMAC) as a technique for IgG 1 monoclonal antibodies purification: the effect of chelating ligand and support. 160(7): 2148-2165.
- Arnold FHJB (2009) Metal-affinity separations: a new dimension in protein processing. 9(2): 151-156.
- Block H, Maertens B, Spriestersbach A, Brinker N, Kubicek J(2009) Immobilized-metal affinity chromatography (IMAC): a review. In Methods in enzymology, Elsevier463: 439-473.
- Ecker DM, Jones SD, Levine HL(2015) In The therapeutic monoclonal antibody market, MAbs, Taylor & Francis 7(1): 9-14.

 Short Communication
Short Communication