Abstract
Aim: To improve the quality assurance, auditability and performance in compound management systems.
Background: Managing hundreds of thousands of compounds in a drug screening facility requires the use of comprehensive data management software that can track and keep track of compound usage and movement such as duplicating multiple copies of mother-daughter plates and generating of intermediate and assay plates. Commercial software for Compound Management (CM) is usually expensive, requires an annual subscription fee for continuous support and updates. Such software is usually beyond the means of academic or publicly funded drug screening facilities.
Objective: We report here the development of an in-house web-based CM System (CMS), PharmApps (short for Pharmaceutical Applications), which tracks and logs compounds dispensed for primary High Throughput Screens (HTS), compounds hitpicked for retest and dose response, plate management and administrative functions.
Results: The program has security measures to allow only assigned bioassayist to access plate information particular to his/her assigned screening campaigns while inventory functions are assigned to CM personnel only. Unique features include visual plate and well dispensation status, plate formatting record, barcode information, inventory update and query functions. Lessons drawn are applicable to both Commercial Off-The-Shelf (COTS) and in-house CMS in areas such as quality control and quality assurance, auditability /traceability and system performance.
Method: A system design and development methodology that identifies and addresses the gaps in both workflow and information processes.
Conclusion: We have successfully created and deployed a functional CMS in our Centre. The conceptual approaches suggested are applicable for both commercial and customized CMS, and highly relevant to any organizations aiming to achieve positive returns on investment in such systems.
Keywords: Compound Management; High-Throughput Screening; Quality Control; Quality Assurance; Auditability; Traceability; System Performance; Compound Management System; Laboratory Information Management System; Application Development; Software Customizations
Abbreviations: CM: Compound Management; CMS: CM System; HTS: High Throughput Screens; COTS: Commercial Off-The-Shelf; QC: Quality Control; QA: Quality Assurance; LIMS: Lab Information Management System; LCMS: Liquid Chromatography-Mass Spectrometry; IDBS: ID Business Solutions; AD: Active Directory; AJAX: Asynchronous Javascript and XML; DOM: Document Object Modeling; JSR: Java Specification Requests; DMSO: Dimethyl Sulfoxide; SMTP: Simple Mail Transfer Protocol
Introduction
One of the major challenges in supporting High-Throughput Screening (HTS) is the need for a high-performance CMS that can perform accurate tracking, close monitoring, error reporting, and doing so while dispensing hundreds of thousands of compounds over a short period. Thus, a CMS must meet high standards of Quality Control (QC), Quality Assurance (QA), auditability, traceability and system performance to ensure operational efficiency and productivity. Satisfying all these require identifying and addressing weaknesses and gaps in the entire CMS. In this report, we detail how gaps in our CM process were uncovered and surmounted. We categorized gaps as either knowledge or information, with knowledge gap defined as information needed but the process of acquiring it is unclear. In contrast, information gap refers to missing information that can be gather through known processes. Identifying these gaps pose the first hurdle. For knowledge gaps, there is the additional challenge of finding or creating techniques to overcome them. There were two knowledge gaps. The first was how to verify a successful dispensing from a specific source to destination well occurred. The second was how to ascertain a destination well contains a target compound.
Along the way, we addressed information gaps within our CM process. We also described challenges faced with COTS software (used previously to support CM operations) and how an inhouse CMS was developed to surmount the above. Although the solution appears unique, we believed the approaches taken to be generalizable and can be applied in either CMS or Lab Information Management System (LIMS) implementation. We reviewed the literature relating to QA, QC, system costs and performance. A study of existing CMS/LIMS [1,2] painted a diverse picture. In systems such as MScreen and SAVANAH [3], there were no explicit QC measures in the software to verify that samples had been dispensed properly. Hints of dispensing problems would only surface when test results are inconsistent. For example, initial screening results might be positive, but retest results turned out negative either due to wrong samples dispensed or target samples not being dispensed. Such incidents pointed to issues upstream but do little for problem monitoring, root cause identification or even aid in the troubleshooting process. In Charles et al. [4], improvements in QA and QC were gained through capturing information from Liquid Chromatography-Mass Spectrometry (LCMS) equipment into a centralized system.
The accumulated records made it possible to troubleshoot low quality samples or anomalies in biological testing. COTS software offers great cost and time savings when a lab workflow requirement can be met either directly or through one-off software configurations [5]. In cases where extensive customizations are needed or where customizations affect system performance, their value propositions are questionable. Various factors like cost of customizations, time and effort required and degree of performance degradation need to be taken into account. One way in tackling the cost of customizing COTS software is to use configurable, open source software to support lab workflows [6]. However, capturing information from lab instruments used is necessary to validate and verify that processes are working properly in the overall CMS/LIMS. Data generated by one instrument when processing samples should be compared with information generated by the next instrument, when feasible, to ensure that there are no discrepancies. Underlying this validation approach is the observation that the output of one instrument often forms the input for the next.
Although, this approach appears onerous, the successful examples [7] of existing LIMS that have used rigorous software validation and verification to achieve national accreditation highlight its importance for successful implementations. Lastly, HTS operation generate huge amount of data for analysis. We therefore expected that reviews of CMS system performance would be common in the literature. Unfortunately, most literature on CMS do not detail performance capabilities or how performance improvements are achieved; perhaps due to commercial considerations. In the next section, we will describe in details the process of developing an in-house CMS known as PharmApps. As we did not patented our system and due to its heavy customizations and uniqueness to our environment, we did not make the source code freely available. However, we are willing to work with collaborators who are interested to implement a similar system within their organizations.
Materials and Methods
Requirements
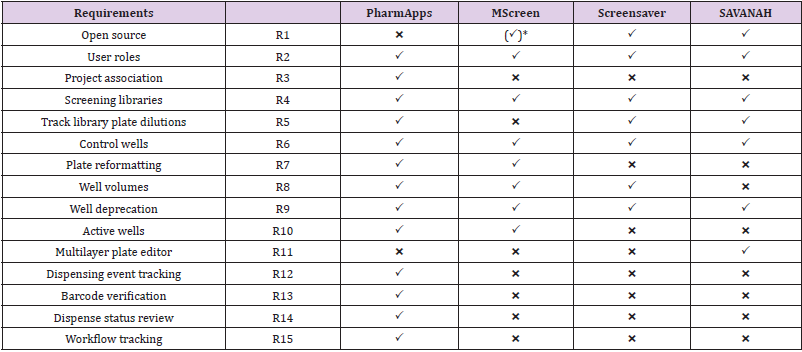
PharmApps was developed as a replacement for a COTS CMS. There were no requirements for adding analytical capabilities as those features were available in another software. Instead, information of samples was transferred from PharmApps to the analytic software as necessary. Therefore, we will only focus on listing CMS requirements in Table 1. PharmApps is not open source (R1) as the system was developed solely for internal usage. It is designed for restricted access based on different roles (R2) and projects association (R3). The system manages different screening libraries by separating them into categories (R4). The system keeps plate layout, well dilution factor and well type (control vs sample) in pre-defined templates. This allows maximum flexibility in setting plate dilutions (R5), types of control wells (R6) and plate layout formats. As the source to destination well transfers are defined during dispensing, information on plate reformatting (R7) from 96- to 384-well plates or from 384- to 1536- well plates are derived from dispensing logs.
Table 1: Feature Comparison between Pharm Apps and Comparable Solutions for CMS.
Note: *Requires licensing (freely available for academic / non-profit).
The system keeps track of the logical well volumes (R8) in addition to different well statuses such as deprecated (R9) or active (R10). There are no multilayer capturing of attributes (R11) in PharmApps. As all dispensing events are captured in log files, we are able to verify the source of each destination well (R12) in each dispensed plate. Both the source and destination plates are barcoded, and their barcodes are verified during dispensing (R13). A liquid handler measures and reports the dispensing status (R14) of each plate prepared for biological testing. The system tracked the entire process (R15) from the time of request until the handover of dispensed plates to bioassayists, so if there is an issue, we know exactly which stage it occurred.
Modular System Architecture
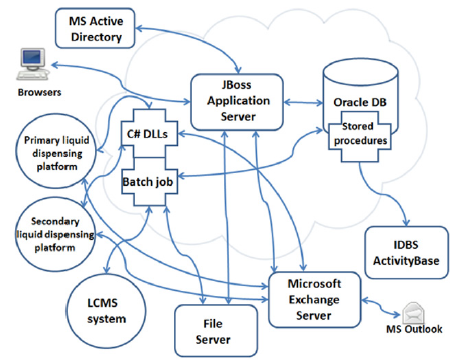
PharmApps is a modular system with a core sub-system comprised of various batch programs, an application server and a database. The core sub-system interfaces with other peripheral systems such as a Microsoft Exchange Server, ID Business Solutions (IDBS) Activity Base and Microsoft Active Directory (AD) as illustrated in Figure 1.
Figure 1: Components of an HTS workflow with the JBoss application server forming the system’s core. It integrates and coordinates the various sub-components such as batch jobs, liquid dispensing platforms, Activity Base and so on. The arrows indicate either unidirectional or bi-directional data flow between these components.
Database Server: Oracle Database 11g Enterprise Edition Release 11.2.0.4.0–64bit:
The data generated by PharmApps are stored in an Oracle database instance running on top of the Red Hat Enterprise Linux 6.3 operating system. There are other Oracle instances within the same physical machine serving other applications. The system relies heavily on customized Oracle’s PL/SQL procedures to do the heavy lifting of storing and reading large volume of data. Customized PL/SQL packages reconstruct highly normalized data into objects collections before returning to the application server or helper programs.
Web Application Server: J boss Application Server 7.1.3. Final The JBoss application server is used as the platform for publishing in-house developed Java Enterprise Edition (JEE) modules. These modules control and coordinate the CM workflow processes and ensure that the business logic is adhered to. The application server also provides other services such as rendering web pages, controlling user authentication and information access and simplifying the complexities of data manipulation, storage and extraction.
Modern Web Browsers and Javaserver Faces (JSF) Framework
Web browsers have been enhanced with the introduction of Asynchronous Javascript and XML (AJAX) technology as well as Document Object Modeling (DOM), allowing for a more immersive and interactive experience for users. JSF is a Java framework that makes it easier for developers building web applications. Leveraging on these two elements, a number of JSF frameworks had emerged to simplify the task of developing interactive and responsive web applications. PharmApps leverage on Rich Faces [8], a JSF framework, to provide a consistent and responsive user interface across different web browsers. By reusing the wide range of Javascript widgets and themes available in Rich Faces, we created a homogeneous look and feel in the web interface. A top down navigation menu mimics the experience of a desktop application; further enhancing the familiarity level for the users. Functions are grouped within menus based on an Action-Task-Goal mnemonic sequence. For example, users who needs plates dispensed for their assays, hover the mouse pointer over the Request > Screening > New submenu and click on it to display the request form. A task oriented mental model was selected as the users know what they want, and the navigational sequence mirrors the work process. By leveraging on these technologies and elements of interface design, PharmApps is able to deliver a more user-friendly and interactive experience.
Analytic Software: IDBS Activity Base
Activity Base is the analytic software used by the bioassayists to analyze their experimental results. Before doing this, the assay plate data such as well sample description, well volume and sample concentration must be transferred from PharmApps to Activity Base via Activity Base’s PL/SQL packages. Once transferred, bioassayists can then upload experimental results into Activity Base for analysis.
Emailing and Lightweight Directory Access Protocol (LDAP) System: Microsoft Exchange Server and Active Directory
To meet the workflow requirements, we identified three prerequisites.
a) The ability to send task alerts through emails. This is done through the emailing capabilities of Exchange.
b) Access to users’ information such as email addresses, departments and names. By acting as a LDAP server, the AD is the source for this.
c) Details of the user roles and their projects association. These are assigned and stored within PharmApps.
System Framework
The PharmApps system is written using JEE 6 with JBoss Seam 2.3.1-Final [9] as the foundation framework. JBoss Seam is a meta framework that amalgamates other frameworks such as JSF, Hibernate, Java Persistence API and Java Transaction API, AJAX to create a seamless platform for web application development. Together, they offer an application comprehensive capability in areas such as model-view-controller, context and dependency injection, authentication and authorization, workflow, transactions, rules engine, and web services. Java Specification Requests (JSR) 299: Contexts and Dependency Injection subsequently supersedes this framework. At the time of implementation, we selected Seam as it had a more comprehensive, mature and integrated framework compared to the relatively new JSR 299 initiative.
System Pre-Requisites
All assay plates and tubes processed through PharmApps have to be barcoded in one- or two-dimensional format. All barcodes must be unique. One-dimensional barcodes are generated using a running number while two-dimensional barcodes are provided by a vendor (with the assurance that there are no replicates).
Integrated Workflows
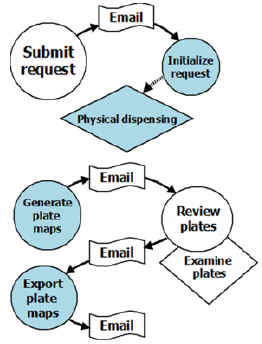
Figure 2 is an example of an integrated workflow in PharmApps. By integrating workflows into PharmApps, the interactions between users and equipment are formalized and captured. This allows active monitoring and error detection to take place. Users are also kept informed of the stage of their requests and any outstanding tasks.
Figure 2: Pharm Apps customizes and manages two main workflows. The first flow begins with sample requests and ends with dispensation tasks. The second flow generates virtual plates, reviews the dispensing status and exports the plates’ data to the analytical software. The circles indicate users’ interactions with the system while the rhombuses represent physical tasks. Email alerts inform the users of any pending tasks in each workflow.
Application Module 1: User Access and Role: Users login to PharmApps using their AD credentials and are granted the role of Guest upon login. Additional user roles were created and assigned to support the screening, CM and LCMS operations. The names of the roles are as follows: Bioassayists, Compound Administrators, LCMS Administrators and Project Administrators. The Bioassayists role is assigned to the researchers who need plates dispensed for screenings. The Compound Administrators role is given to the users who are managing the screening libraries and performing the dispensing. The LCMS Administrators role is assigned to the chemists who are performing analytical chemistry on library compounds. Lastly, the Project Administrators role is given to the head of the HTS department. This allows the designated user to assign researchers to their respective projects. Association to projects create an additional level of access control on top of user roles. Task alerts (via emails) are also directed based on role and project associations.
Application Module 2: Screening Request: The screening request module was developed to address an information gap where users’ requirements were not captured in a database. Without this information, it was difficult to validate if dispensing matches requests. This module is accessible by users with Bioassayists role. It translates the screening requirements (screening libraries needed, well volume, well concentration and so on) into the necessary dispensing activities. In the process, it validates the samples requested against what is available in the CM library. Users can select from a list of categories (i.e. screening library names) or alternatively, upload a list of source plate barcodes or unique sample IDs when preparing their request. Upon submitting of requests, the selections are broken down to the lowest denominations i.e. sample IDs. A list of available plate layouts is displayed depending on the selected experiment stage. By specifying a plate layout, the users define how dispensing will be performed. For example, depending on assay requirements, a user may choose a dose-response plate layout with either two-fold or three-fold serial dilution. Once a request is submitted, the system will notify CM administrators via emails to begin dispensing. The system keeps track of which samples are outstanding in a request and automatically closes the request once all samples are dispensed.
Application Module 3: Dispensing Process/Review: This module processes equipment logs generated by the dispensing of samples. The dispensing is choreographed by the Agilent VWorks software which coordinates various instruments such as Agilent Direct Drive Robot, Bravo and Labcyte Echo. Collectively, VWorks and the instruments it controls are known as the Biocel Integrated Systems. A set of instructions to VWorks controls the series of tasks and the mix of equipment needed to complete a dispensing. These sets of instructions can be stored as templates and these templates are called the VWorks protocols. During dispensing, both source and destination plates are barcode scanned and recorded by both the VWorks and Echo system. These two sets of barcodes are compared for discrepancies and email alerts are sent out if there are any. By placing programming hooks (java scripts, .NET dynamic linked library and executables) within the VWorks protocols, PharmApps is able to extract the dispensing logs from Bravo and Echo liquid handlers. The Echo liquid handler monitors source well depth and fluid composition with acoustic signals. The resulting measurements act as a proxy for the status of the destination wells.
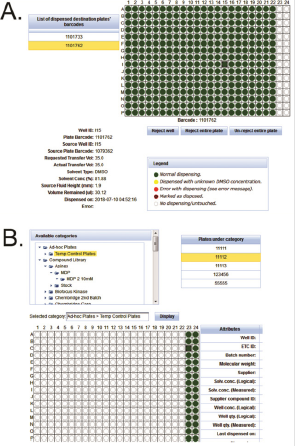
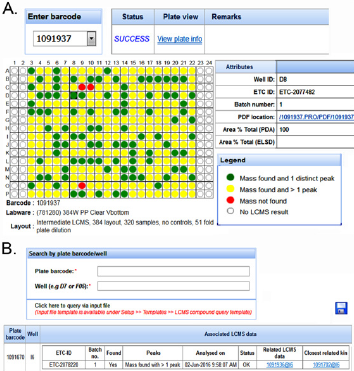
By combining the dispensing logs with instrument measurements, PharmApps reports the resulting dispensing status visually in the form of plate maps as shown in Figure 3A. Wells that are not dispensed are displayed as red circles while dispensed wells are shown as green or yellow (dispensed with unknown DMSO concentration). The internal QC standards are as follows: not more than 1.5% of test wells or not more than 3% of control wells in a plate should be red. Failing these QC standards, the CM administrators will have to re-dispense. However, individual bioassayists might be stricter in their acceptance criteria and reject a plate if there is even one non-dispensed well. Occasionally, errors do occur due to either human or machine problems or a combination of both. An example of machine problem is the robotic arm crashing which causes a dispensing session to fail halfway. In such cases, manual intervention by the IT support is needed. The system will be instructed to only extract the successful portion of the dispensing session. The failed portion would have to be re-run. An example of human error would be using the wrong source plates. Invalid plate barcodes would then be detected which prevent further system processing. In addition, CM administrators will be alerted if the wrong sample volume or concentration is dispensed. As there are many possible operational issues, it will be impossible to cover all of them. However, by requiring both the CM administrators and bioassayists (who are affected) to review the dispensing plate maps, many operational issues can be detected and addressed. This prevents the wastage of time and resources downstream.
Application Module 4: Query and Tracing: Since all plates are barcoded, scanned and tracked, the tracing module displays the lineage of each plate. This module allows the bioassayists to verify the source of their assay plates visually. The system also displays the remaining volume as measured by the instrument, giving a close approximation of the actual volume in source plates.
Application Module 5: Categories: This module allows the samples to be categorized into different groupings see Figure 3B. The names and levels of categories and sub-categories are totally flexible and configurable by the users. The samples are represented as files in a folder tree schema familiar to users, making it easier for selection and viewing. This also makes it convenient to apply certain rules or operations on a particular category. For example, multiple screening libraries can be selected for sample requests, queries or disposals.
Figure 3: After dispensing, CM administrators and bioassays can review and reject plates based on their statuses.
a) A green or yellow circle denote a dispensed well while a red circle means no dispensing occurred. We use various categories to group the plates.
b) Each category can contain further sub-groups in a format emulating folder, sub-folders and so on with plates resembling files.
Application Module 6: Lcms Module: This module was developed to partially address the question of whether a library compound is present in a destination well. The bioassayists can recommend library compounds to chemists for analysis in the LCMS module. This process is tracked as a separate workflow. Once the compounds are analyzed, the LCMS results can be downloaded and displayed in a highly customized manner see Figure 4A. Users can also query whether the compounds in their assay plates have related LCMS results see Figure 4B.
Figure 4: The LCMS module in PharmApps allow users to check for the presence of specific compound.
a) PDFs files of the results are downloadable by clicking on the links. Through the LCMS results.
b) Users can identify assay plates/wells that are closely related. The association arises because the LCMS plates and the assay plates are both prepared from the same source or ancestor.
Results and Discussion
Work on the new system began in October 2013. PharmApps was running in parallel with its predecessor system (a customized COTS software) in December 2014 and officially took over one month later. In total, the project took one year and three months. During this period, one system analyst was assigned solely to the project. One month was spent initially on user requirements followed by three months of prototyping. The prototype was developed to address the first knowledge gap. It eventually formed part of the dispensing module. Due to the novelty of the system, users were initially apprehensive about its ability to improve operational effectiveness. A series of hands-on sessions and presentations covering the modules and system features were conducted to address this. Users’ confidence level had increased with usage and the feedback had been consistently positive since rollout. The LCMS module was introduced much later in March 2016.
Significant changes were observed following the implementation of PharmApps. There was a review of the sample storage protocol after PharmApps reported many failed dispenses due to invalid source well conditions. Investigations showed that this was due to source well volumes exceeding the limits of the dispensing machine. This was caused by the hydration effect of Dimethyl Sulfoxide (DMSO). The review combined with a few studies led to changes in the standard operating procedures. The changes covered the preparation of source plates, setting of solvent concentration, the well volume limit for active dispensing and the duration period before discarding old stocks. In particular, it became mandatory for all plates or tubes to be barcoded. Dispensing results are available only if tracked via the PharmApps ecosystem. This resulted in a vast improvement in the success and quality of dispensing as well as the discarding of certain non-dispensing compound libraries. Both CM administrators and bioassayists started to discard plates that failed the QC standards. Lastly, improved consistency in experimental results were reported.
Challenges Revisited
The predecessor of PharmApps recorded dispensing information through a series of user interactions. Initial source plate information was uploaded from manually created files. Subsequent information on compound transfer was generated via user instructions. For example, a user would instruct the system to create a destination plate and transfer a certain volume from a specific source plate to it. The whole process was done in silico and not verified by other sources. Thus, actual events could differ from recorded events such as the wrong source plate used for dispensing. Such errors were hard to trace and the only clue was the huge deviations uncovered in experimental results. It was hard to reconstruct what happened from the users’ recollections, as it was difficult to recall specific details against a background of voluminous activities and fading memories. All these factors placed severe strains on the QA and QC process. We realized the need to improve QA and QC through traceability and auditability in the whole workflow process. Gaps in the CM process had created blind spots that weakened the entire QC chain.
System Design and Development
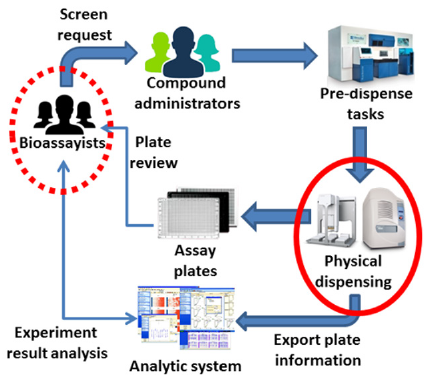
A review of the previous CMS was conducted. This was followed by a series of interviews with end-users (mainly CM administrators and bioassayists) and a study of the HTS operations. Both explicit and implicit concerns were documented. In the process, an information gap (Figure 5, dotted red circle) was identified that arose from the explicit need to match dispensing with requests. As a result, sample requests from bioassayists were captured in a new module. Implicit concerns were given special attention as they indicate the existence of knowledge gaps in the current environment. Two knowledge gaps were deduced:
a) Whether a source to destination well transfer was successful,
b) Whether a library compound is present in a well.
Figure 5: An analysis of a typical HTS operation revealed both information (dotted red circle) and knowledge (normal red circle) gaps. We overcome the information gap by recording the information generated the users. The knowledge gap required developing of new techniques for capturing inputs from dispensing instruments. By overcoming these gaps, we created an integrated workflow that enhances the overall operational quality.
Addressing the first knowledge gap required developing new tracking capabilities in the dispensing process (Figure 5, normal red circle). In PharmApps, the dispensing tasks were tracked by recording their interactions with lab equipment. Through the event logs, there is a real-time, accurate and reliable way of knowing how and when the dispensing was done as well as the equipment used (see Application module 3 in Materials and Methods section). The later addition of a LCMS equipment and its corresponding module addressed the second knowledge gap in PharmApps. PharmApps was designed for massive data processing from plate generation to queries. Generating two hundred 384-wells plates took less than fifteen minutes. These outcomes were achieved by combining large number of transactions into one or more batch processing. Compared to processing one well at a time, both throughput and performance were vastly improved. The use of asynchronous batch jobs also created positive improvements in user experience.
Lessons Learned
Our experience with a COTS and a subsequent in-house CMS highlights the importance of considering the type of research as well as the mix and match of instruments in a lab during implementation. It is also important to alert key decision makers whenever discrepancies are detected and allow reviews at crucial stages. Doing so will improve productivity and operational effectiveness. These will require an in-depth understanding of the CM operation in a lab and raise other challenges. Such as understanding the flow of information, overcoming information silos while maintaining rigid access control. Potential impacts on system performance and reliability should be addressed early. For COTS CMS, this is when customizations are made to the software. For in-house CMS, this is during system design. The design should accommodate the huge volume of data generated by HTS operations. Failure to do so can cause poor reliability and performance leading to user frustrations and lost productivity.
The following steps were taken during the implementation process:
a) Identify explicit and implicit user concerns and their corresponding information and knowledge gaps. This should be performed during the user requirements stage.
b) Inventory lab equipment used in the CM process.
c) Understand the possibilities, benefits and risks of integrating the lab equipment.
d) Perform the integration of workflows and lab equipment.
Note that there is a possibility that step 2, 3 and 4 may not address the knowledge gaps uncovered in step 1. For example, PharmApps was unable to determine the presence of a library compound in a well until the LCMS module was added.
A three-step approach was taken to integrate the workflows:
a) Identify the information that will impact the users and their operational effectiveness,
b) Extract, gather and disseminate this information to the affected users, and
c) Develop the workflow process that will automate this information exchange.
The degree of integration to lab equipment varies from low coupling to tight coupling. An example of low coupling will be copying and sanitizing the equipment log files manually before feeding them into the CMS/LIMS. With tight coupling, the system uses the API of the lab equipment software to actively interrogate and influence the operations of the lab equipment. An example would be to check if the correct source and destination plates were used and to halt operation if discrepancies are detected. The extent will depend on users’ requirements and the technical feasibility.
Although integrating the workflow processes and lab equipment within a CMS offers many benefits, there are dangers involved. These dangers occur when the constraints of integration are not well understood. These constraints are described below.
Constraints of Integration
Although improvements in information quality is observed when incorporating equipment logs and measurements, there are limitations arising from the technology used. The first constraint is the need to understand the operating specifications and limits of the equipment involved. An example is the acoustic technology used in Echo liquid handler. It cannot correctly handle a source plate or a dispensing session that has different solvent types. If the samples were dissolved in water but the dispensing was set for DMSO then a successful transfer would be reported although no dispense occurred. Similarly, the Bravo liquid handler will simply suck up air if there are no liquid in the source well, the logs will still indicate a transfer had occurred. The second limitation arises when the readings are not a direct measure of the targets of interest. An example is the Echo’s acoustic measurements of the source well prior to a transfer. The measurement taken belongs to the source well and not the destination well. There are situations where this fact is important. For example, if there are library compounds that have poor solubility and precipitate, the Echo may report a successful drop transfer but the compound may not be present in the dispensed well. Similarly, the presence of a compound in a LCMS plate does not guarantee the same in a sister plate even though both shared the same parent. The sister plate could have suffered other intermediate issues such as errors in handling or dispensing.
The third constraint arises from the integration of lab equipment and workflow processes in the CMS. This process requires project and technical expertise more commonly associated with large system integrators. Sufficient time and resources are needed for a careful study of the information flow, stakeholders and equipment used in the lab. Project management skills will be needed to handle extensive user interviews, resource planning and getting the requirements right. Strong technical expertise is needed to achieve successful integration with the lab equipment and to handle potential breakage when the lab equipment software is upgraded. These factors will impact the success of both in-house and COTS CMS implementation.
Working with Open Source Software
Unlike open source LIMS such as M Screen, Screensaver and SAVANAH, PharmApps is primarily a CMS. It offloads the analytics to specialized software such as Activity Base. Although it is possible for PharmApps to leverage on the analytic functionalities of open source software such as Screensaver, it will require populating sample information into the relevant modules of the open source software. We will need to study if code changes are required and the legal implications if copyleft licensing is involved. We used Microsoft AD as the LDAP server but open source alternatives such as Open LDAP [10] can also be used as the source of user information. The changes required would be limited to the LDAP module in PharmApps. The Microsoft Exchange Server is used for its Simple Mail Transfer Protocol (SMTP) capability and close integration with Outlook. But there are open source alternatives such as Kolab [11] which have both SMTP feature and Outlook integration. However, integrating open source software like Open LDAP and Kolab in organizations that use Microsoft AD for single sign on will be technically challenging. It will require greater investment in technical expertise, efforts and time that are hard to justify when there are ready solutions available.
It would be more challenging to switch from Oracle database to an open source alternative like PostgreSQL [12]. This would require migrating the stored procedures and functions, userdefined types as well as making Java code changes for storing data into the database. Furthermore, as Activity Base only supports Oracle database, either we use another analytic software or there is a feasible way of linking PostgreSQL to Oracle. All these would require substantial changes.
Ongoing Works
In June 2017, we began to explore ways for migrating PharmApps to newer web technologies. We decided to leverage on the official CDI feature in JEE 7 because the architect for JBoss Seam was also the specification lead for CDI. At version 1.2, CDI had matured substantially and being part of JEE 7, we do not need to download any additional libraries. We chose to upgrade the application server to WildFly [13] (at version 10.1.0. Final) as it is the successor of JBoss application server. As development work on Rich Faces had been discontinued, we decided to shift the JSF framework to Prime Faces [14] as both shared similar design approaches. The new PharmApps uses Apache Maven [15] as the build automation tool. Both the old and new PharmApps are currently in use as modules are migrated one at a time to avoid a “Big Bang” approach. This is possible because user sessions are transparently created in both systems upon login and destroyed when the user logout. When the users click on a migrated module in the old system’s menu, they are simply transfer to the new PharmApps and vice versa. Although more efforts are needed to ensure that both systems’ menus and appearance are similar, this reduces any adverse effect on the overall user experience. We created PharmApps to meet certain requirements and for operation within our unique environment. It would not be appropriate to publish its source code without giving detailed guidance on how to customize it for another lab environment. This would necessitate a process of close collaboration and knowledge transfer. As such, we are willing to offer our knowledge and expertise to any interested collaborators seeking to implement a similar system.
Acknowledgement/Conflict of Interest
There is no financial conflict of interest among the authors. Support for A.Y., C.G., M.L.C., T.S.H. and D.N.C. was from an A*STAR core funding to the Experimental Drug Discovery Centre.
References
- Jacob RT, Larsen MJ, Larsen SD, Kirchhoff PD, Sherman DH, et al. (2012) an integrated compound management and high-throughput screening data storage and analysis system. J Biomol Screen 17(8): 1080-1087.
- Al-Taee MA, Souan L, Al-Haj A, Mohsen A, Muhsin Z (2009) Quality control information system for immunology/serology tests in medical laboratories. 6th International Multi-Conference on Systems, Signals and Devices, Djerba, Tunisia, pp. 1-7.
- List M, Elnegaard MP, Schmidt S, Christiansen H, Tan Q, et al. (2017) Efficient management of high-throughput screening libraries with SAVANAH. SLAS Discov. 22(2): 196-202.
- Charles I, Sinclair I, Addison DH (2014) Capture and exploration of sample quality data to inform and improve the management of a screening collection. J Lab Autom 19(2): 198-207.
- Russom D, Ahmed A, Gonzalez N, Alvarnas J, DiGiusto D (2012) Implementation of a configurable laboratory information management system for use in cellular process development and manufacturing. Cryotherapy 14(1): 114-121.
- Tolopko AN, Sullivan JP, Erickson SD, Wrobel D, Chiang SL, et al. (2010) Screensaver: An open source Lab Information Management System (LIMS) for high throughput screening facilities. BMC Bioinformatics 11: 260.
- Taina A (2015) Software validation with respect to requirements specified by SR EN 15189: 2013 and SR EN 17025:2005. A practical approach for a medical laboratory information management system (LIMS) and a gas meter in testing laboratory, 9th International Symposium on Advanced Topics in Electrical Engineering pp. 720-724.
- Rich Faces. http://richfaces.jboss.org/ (Accessed January 10, 2019).
- JBoss Seam. http://seamframework.org/ (Accessed August 30, 2018).
- Open LDAP. https://www.openldap.org/ (Accessed January 9, 2019).
- https://kolab.org/ (Accessed January 10, 2019).
- https://www.postgresql.org/ (Accessed December 27, 2018).
- http://wildfly.org/ (Accessed January 11, 2019).
- Prime Faces. https://www.primefaces.org/ (Accessed January 11, 2019).
- Apache Maven. https://maven.apache.org/ (Accessed August 2, 2019).

 Research Article
Research Article