Abstract
Aim: A biomedical hyperspectral imaging system is proposed to facilitate visualization of biological tissues.
Methods: Images simultaneously measured with the CASSI system and the RGB camera are stacked and feed into OMP algorithm to reconstruct the hyperspectral images with a pre-trained dictionary.
Results: High reconstruction quality of hyperspectral images is demonstrated by SSIM, SAM and PSNR values, and the instance shows that with the enhancement of hyperspectral image the visualization of tissue image is improved.
Keywords: Biomedical Hyperspectral Imaging; Coded Aperture Snapshot Spectral Imager; Dictionary Learning
Introduction
Hyperspectral Imaging (HSI) captures Two-Dimensional (2D) spatial and one-dimensional spectral information of the target. With abundant material information, HSI offers great potential for medical applications [1]. For instance, as biochemical process of disease development can be monitored with spectral variation, HSI can be applied in noninvasive disease diagnosis [2]. And due to high contrast imaging for object with similar color but different spectra, HSI can be employed in visual aid during surgery [3]. However, conventional HSI systems require time-consuming scanning process in spatial or spectral dimension [4]. Snapshot Hyperspectral Imaging (SHSI) systems solve this problem by capturing Three-Dimensional (3D) data by 2D projections, without sacrificing the temporal dimension [5]. Among SHSI systems, Coded Aperture Snapshot Spectral Imager (CASSI) is one of the most promising solutions as it naturally embodies the principles of compressive sensing [6].
We aim to propose a biomedical imaging system, in which hyperspectral images are reconstructed from simultaneously measured CASSI images and RGB images with a pre-trained overcompleted dictionary. An instance shows how hyperspectral images facilitate visualization of biological tissues.
Methods
System Configuration
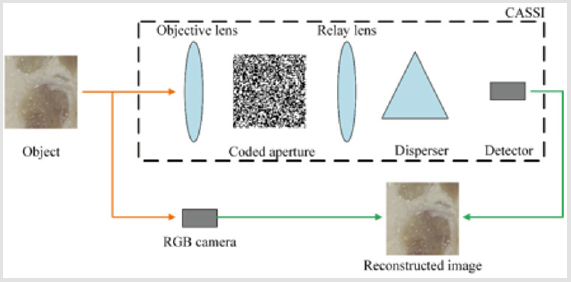
The schematic diagram of the combined system is shown in Figure 1. In addition to the commonly used RGB camera, a CASSI system simultaneously records the light from the object. In the CASSI system, the incident light is firstly spatially modulated by a binary-coded mask, which can be considered as a pinhole mask with a designed pattern. Then the light is spectrally dispersed by a dispersive element along certain spatial dimension, and finally integrated on the detector. The coded measurements are mathematically equivalent to compressive projections in compressive sensing, which enables the 3D data to be reconstructed from 2D projections by solving an inverse problem.
Reconstruction Algorithm
Supposing the input signal f is sparse in some domain by the transform f = Ψθ , where Ψ is the inverse sparse transform and θ is the vector composed of coefficients after the sparse transform, the reconstruction of CASSI can be described by the optimization problem
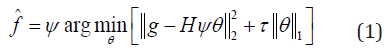
where g is the compressive sensed data, H is the system matrix denoting the sensing process of the CASSI system, and τ is the tuning parameter.
A dictionary-based method is adopted in the reconstruction of hyperspectral images from CASSI projections and RGB measurements. To keep the consistency of the reconstruction, images from the CASSI system and the RGB camera are stacked and feed into the reconstruction algorithm. An over-completed hyperspectral dictionary is trained with a hyperspectral image database by KSVD in prior to reconstruction [7]. By simply combining the system matrices of the CASSI system and the RGB camera, the secondary dictionary is obtained. Then Orthogonal Matching Pursuit (OMP) algorithm is applied to solve the optimization problem [8].
Results and Discussion
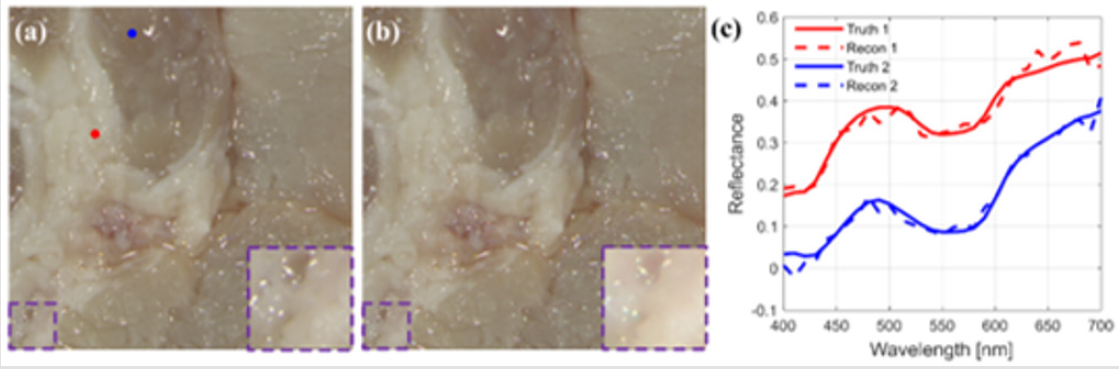
To investigate the performance of the proposed system and algorithm, the reconstruction result of the hyperspectral image from the CASSI projection and the RGB measurement, which is undertaken on a piece of pork with lean meat and fat, is shown in Figure 2b. For clear comparison, the details at the left bottom corner for both the ground truth and the reconstructed hyperspectral image are magnified and shown in Figure 2a the purple dashed boxes. As the RGB measurements are included in the compressive sensed measurements, there is only slight difference in the intuitive RGB expression for hyperspectral images of the ground truth and the reconstruction, which also indicates high spatial resolution of the reconstructed hyperspectral image. High spatial resolution can also be proved by the high structural similarity index (SSIM = 0.9670), which is usually employed to evaluate the similarity between two images in the spatial dimension. As for the spectral performance, two spatial points at lean meat area and fat area are chosen separately to show the point spectra difference, as shown in Figure 2c. The point spectra of these two points manifest that the reconstruction is less smooth than the ground truth. But the fidelity of the spectrum is still high enough for application, which well conforms to the pixel-wise low spectral angle mapper (SAM = 0.0963) value [9]. For overall performance, high peak signal-tonoise ratio (PSNR = 33.58) is realized, corresponding to relatively low reconstruction error compared with maximum intensity of the hyperspectral image.
Figure 2: (a) Ground truth. (b) Reconstruction result of the proposed system. (c) Point spectra of points randomly selected in (a).
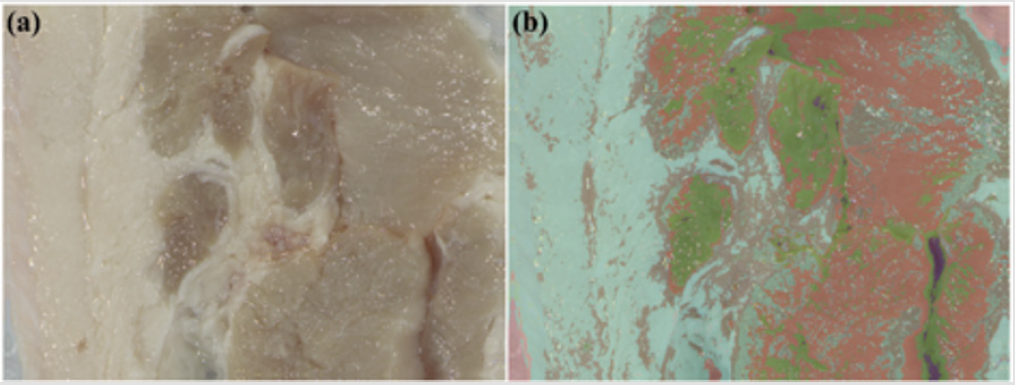
The hyperspectral imaging can be employed in biomedical imaging to enhance the contrast at the tissue level. For instance, the RGB image for a piece of pork with lean meat and fat is shown in Figure 3a, and the corresponding image enhanced by hyperspectral image is shown in Figure 3b. The enhancement is realized with the pixel-wise SAM calculated with the average spectrum of the whole hyperspectral image, and different colors correspond to different levels of SAM value. The cyan part indicates the fat area, while the green part and the red part correspond to lean meat with different density, and the magenta part at the bottom-left and the upper-right corner is the connective tissue. The colors defined with information from the hyperspectral image divide the RGB image into more detailed areas, which can function in visual enhancement in biomedical imaging.
Figure 3: (a) RGB image of the ground truth. (b) Corresponding image enhanced by hyperspectral image.
Conclusion
In summary, we propose a biomedical hyperspectral imaging system to facilitate visualization of biological tissues. In this system, hyperspectral images are reconstructed from simultaneously measured CASSI images and RGB images with a pre-trained overcompleted dictionary. High reconstruction quality is demonstrated by SSIM, SAM and PSNR values. And an instance shows that with the enhancement of hyperspectral image, more distinct image of biological tissue is obtained.s
References
- Lu G, Fei B (2014) Medical hyperspectral imaging: A review. J Biomed Opt 19(1): 10901.
- Joseph AO N (2012) Hyperspectral optical imaging for detection, diagnosis and staging of cancer. University of Southern California.
- Monteiro S T, Uto K, Kosugi Y, Watanabe E (2004) Towards a surgical tool using hyperspectral imagery as visual aid. Proceedings of the International Workshop on Augmented Environments for Medical Imaging and Computer-aided Surgery pp. 97-103.
- Garini Y, Young I T, McNamara G (2006) Spectral imaging: Principles and applications. Cytometry A 69(8): 735-747.
- Hagen NA, Kudenov MW (2013) Review of snapshot spectral imaging technologies. Opti Eng 52: 090901.
- Arce GR, Brady DJ, Carin L, Arguello H, Kittle DS, et al. (2014) Compressive coded aperture spectral imaging: An introduction. IEEE Signal Proces Magazine 31(1): 105-115.
- Aharon M, Elad M, Bruckstein A (2006) K-SVD: An algorithm for designing overcomplete dictionaries for sparse representation. IEEE Trans signal proces 54: 4311.
- Tropp JA, Gilbert AC (2007) Signal recovery from random measurements via orthogonal matching pursuit. IEEE Trans infor theory 53(12): 4655- 4666.
- Kruse FA, Lefkoff AB, Boardman J W, et al. (1993) The spectral image processing system (SIPS)—interactive visualization and analysis of imaging spectrometer data. Remote sensing of environment 44(2-3): 145-163.

 Research Article
Research Article