Impact Factor : 0.548
- NLM ID: 101723284
- OCoLC: 999826537
- LCCN: 2017202541
Salvador Pérez-Mora1, Carlos Pérez de la Mora2, Tania Domínguez-Fernández1, María del Consuelo Gómez-García1, Sandra Perez-González2 and David Guillermo Pérez-Ishiwara1*
Received: December 06, 2022; Published: December 15, 2022
*Corresponding author: David Guillermo Pérez-Ishiwara, Molecular Biomedicine laboratory, ENMH, Instituto Politécnico Nacional. Guillermo Massieu Helguera 239, Col. La Escalera, CDMX, México
DOI: 10.26717/BJSTR.2022.47.007534
Currently, there is a great demand for effective, low-cost, and fewer side effects medicines due to the growing number of diseases in developing countries. The identification of bioactive peptides in compounds of natural sources or those generated artificially is a viable alternative that should be systematically investigated, particularly in those products that have shown immune modulating activity. Dialyzable leukocyte extract (DLE) from human or various species, a low molecular weight peptide fraction of immune cells, has been used for several decades and has clinically demonstrated its usefulness in various inflammatory diseases and those in which immune system functions are out of balance. However, due to its complexity and lack of characterization of its components, it has been difficult to determine its mechanisms of action. In the present investigation we performed proteomic, in silico and interactomic analyses of DLE obtained from Crocodylus moreletii lymphoid tissue (DLEc). The 2D proteomic characterization showed a heterogeneity of peptides in a molecular weight range of 38 to 6 KDa with pI of 4-9; by nanoLC-MS/MS orbitrap analysis it was demonstrated that it is constituted by 302 peptides, which we grouped into 5 main clusters; I. Regulators of gene expression, II. signaling, III. involved in secretion, IV. degradation pathway, V. heat shock, and to a lesser extent, immune system proteins. In silico we identified in its sequence several bioactive peptides, whose activity has been experimentally demonstrated in diverse study models. We found peptides with immunomodulatory; anti-inflammatory; phagocytosis-inducing; antihypertensive; anticancer; antioxidant; modulators of various metabolic pathways; antimicrobial and antithrombotic activities. Our analyses suggest that DLEc is a heterogeneous mixture of peptide sequences containing bioactive peptides with enormous potential as pharmacological adjuvants in different pathologies.
Keywords: Dialyzable Leukocyte Extract; Bioactive Peptides; Health Impact; Proteomics; Bioinformatics
Abbreviations: DLE: Dialyzable Leukocyte Extract; DTT: Dithiothreitol; IEF: Isoelectric Focusing; SDS: Sodium Dodecyl Sulfate; LTQ: Linear Trap Orbitrap Velos
Immunotherapy, also called biological therapy, is a type of treatment that stimulates the body’s natural defenses to combat various complex diseases in which the immune system is compromised or in those that require the regulation and proper functioning of the immune system. Substances produced by the body or manufactured in a laboratory are used to restore, stimulate, or replenish immune function in cancer, infections, or other degenerative diseases, as well as to lessen the side effects of the treatments used. The objective can be prophylactic (prevention) or therapeutic (curative or maintenance). Immunotherapy includes the use of antibodies, vaccines, growth factors, compounds extracted from plants and of relevance the dialyzable leucocyte extract (DLE), “erroneously called transfer factor”. DLE is a complex mixture of bioactive peptides obtained from human peripheral blood leukocytes or lymphoid tissues of various species. Clinical practice and various research studies suggest that DLE is a safe and effective treatment option for a wide range of pathologies. However, due to the broad spectrum of biological activity, resulting from the complexity and diversity of peptides it contains, the study of the mechanisms of action is complex and, in many cases, its probable action is not sufficiently documented and investigated. DLEs have been developed in several countries as supplements to treat immune-related diseases (Ojeda, et al. [1-3]); Inflammatory diseases (Hernández, et al. [4]); infectious diseases (Cabezas, et al. [5-8]); in asthma (Cabezas, et al. [9]); as an adjuvant in cancer treatment (Pineda, et al. [10-14]). On the other hand, our research group, has focused on, addition to studying the inflammatory effects in different disease models, such as osteoarthritis (Acosta, et al. [15]); prostatitis (Pérez-Alvarado, et al. [16]); cervicitis in premalignant lesions associated with human papillomavirus (Acosta, et al. [17]); and influenza (Pérez, et al. [18]); in determining the mechanisms of action in cellular and animal models. Research suggests that modulation of innate immunity and regulation of the inflammatory process constitute the main mechanisms exerted. Further systematized research on i. The exact composition; ii. Gastrointestinal hydrolysis and release of specific bioactive peptides; and iii. The mechanisms of action and bioavailability of the peptides that compose it, will allow a better prescription for specific pathologies, as well as for the development of new drugs and novel and specific therapies that contribute to current treatments. In this work we performed the proteomic characterization of DLEc by 2D and nanoLC-MS/MS orbitrap analyses, we identified bioactive peptides that can interact and modulate different molecular pathways within the cell related to immunomodulatory and anti-inflammatory effects. Our analyses suggest that DLEc is a heterogeneous mixture of peptide sequences containing bioactive peptides with enormous potential as pharmacological adjuvants in different pathologies.
Protein Preparation
Pellets of trichloroacetic acid-precipitated proteins were washed four times in a large volume of 96% ethanol and then resuspended by scraping and extensive pipetting in 5 ml of an isoelectric focusing (IEF) sample solution composed of 8 M urea, 4% (wt/vol) 3-[(3-cholamidopropyl) dimethyl ammonio]-1-propanesulfonate (CHAPS), 40 mM Tris, 2% dithiothreitol (DTT), and 0.2 (wt/vol) Bio-Lyte 3/10 (Bio-Rad). The clear solution contained approximately 0.3 mg/ml protein.
2-D Electrophoresis Separation of Proteins and Spot Quantitation
DLE protein mixture was resolved first by IEF on pH 3 to 10 (nonlinear) ready-made, 13-cm, immobilized pH gradient strips (Immobiline DryStrips; GE Health care), applied to an Ettan IPGphor III (AmershamBiosciences). IEF was carried out at 500 V for 1 h, followed by two gradients of 1000 V; for 1 h and 2.5 h; a final step at 8000 V for 25 and 55 min. Strips were then processed for the second-dimension separation by a 10-min incubation in 6 M urea, 2% sodium dodecyl sulfate (SDS), 0.375 M Tris-HCl (pH 8.8), 20% glycerol, 2% (wt/vol) DTT, followed by a 10-min incubation in a similar solution in which the DTT was replaced by 2% iodoacetamide. Strips were applied to 12.5% SDS-polyacrylamide gels, and electrophoresis was conducted at 100 V for 7 h. Gels were stained with Coomassie blue G-250 (Bio-safe Coomassie; Bio-Rad) and spots detected and analyzed. At least three independently obtained DEL biological sample were evaluated by 2-D electrophoresis, at least three gels were used. The gels from which the spots were excised for sequencing were stained with colloidal Coomassie blue for 1 h, after fixation with 30% ethanol, 5 % glacial acetic acid for 1 h. Finally, the gels were washed three times with water and then washed with water. Finally, three washes were made with milliQ water to distain and visualize the spots.
Sample Preparation for Malditof Sequencing
Precipitation: The lyophilized sample of approximately 50 mg of DLEc was resuspended in 100 μl of 8 M urea, precipitated with chloroform-methanol and resuspended again in 50 μl of 8M urea for digestion. Digestion with trypsin: 50 μg of the precipitated sample was taken. Added 10 mM DTT (final concentration) and incubated for 1 h at 37ºC. Iodoacetamide was added to a final concentration of 55mM, incubated at RT 30 min in the dark. Diluted with ammonium bicarbonate to 2M urea (final concentration). Recombinant trypsin (Roche) was added at a ratio of 1:20 with the protein and incubated O/N at 37 C.
Desalting and Quantification of Protein Concentration: Sample clean-up was performed using a PorosR2-type resin@ and peptides were eluted with 80% acetonitrile (ACN) in 0.1% TFA. Samples were lyophilized in Speed-vac and dissolved in 50 μl of 2% formic acid.
Analysis by NanoLC-MS/MS (Orbitrap): Peptides were separated on-line with an Easy-nLC (Proxeon) using a 160 x 75 μm NS-AC-11-dp3 C18 column (BioSphere) and a Pre-Column Easy-column C18-A1 (Proxeon) at a flow rate of 250 nL/min using a 245 min gradient: 190 min 2-30% of B phase, 40 min 30-40% of B phase 10 min 40-90% of B phase and 4 min 99% of B phase. The composition of solvent A was formic acid 0.1%, acetonitrile 2%, water 98% and solvent B was 99.9% acetonitrile with 0.1% formic acid. Peptides were analyzed using a linear trap Orbitrap Velos (LTQ Orbitrap Velos) hybrid mass spectrometer (Thermo Electron Corp., Bremen, Germany). The electrospray voltage of 1.7 kV versus the inlet of the mass spectrometer was used. The mass spectrometer was operated in the non-data-dependent mode. The parameters for ion scanning were the following: Full-scan MS (400-1800 m/z) plus top 15 peaks MS2.The scanning was performed using a dynamic exclusion list (30s exclusion list size of 500).
Protein Identity: For protein identification, the non-redundant Swissprot (20233 sequences) database was searched using a licensed version of MASCOT 2.3.0 using the Proteome Discoverer software version 1.2.0.208 (Thermo). Search parameters were oxidized methionine as variable modification, carbamidomethyl cysteine as fixed modification, peptide mass tolerance 10 ppm, 1 missed trypsin cleavage site, MS/MS fragments tolerance 0.8 Da. In all protein identification, the FDR was fixed at 1%.
In Silico Analyses: Using STRING version 10.5 (https://version-10-5.string-db.org/) we obtained a general interactome of the identified DLEc proteins, which were grouped according to their biological functions. Similarly, proteins from DLEc with immune functions were challenged against the human proteome in STRING to identify putative interactions to date possible molecular mechanism. Predictions were made with values equal to or up to 70% probability in STRING. To evaluate the immunomodulatory effect of DLEc targeted in silico analysis was performed making an interactome using the DLEc proteins and IL2, IL18, IL1A, IL12A, IL12B, IFNG, CSF2, TNF, IL12RB1, IL12RB2, IL1A, IL1B, IL2, IL4, IL6, IL23A, IL23B, XBP1 and IRF-4. Similarly, modulatory effect of inflammation was evaluated by determining the interaction of DLEc proteins with proteins of the TNFR-NFkB pathway, using TRADD, TRAF2, IKBKB, NFKB1, TNF human proteins. To identify peptide sequences with beneficial biological activity for health, we searched the literature for reports of bioactive peptides with biological effects proven experimentally in cell lines, murine models or in human clinical studies. The selected bioactive peptides with anti-inflammatory; phagocytosis-inducing; antihypertensive; anticancer; antioxidant; modulators of various metabolic pathways; antimicrobial and antithrombotic activities were identified in the amino acid sequences of DLEc proteins (Uniports, https://www.uniprot.org/).
DLEc Proteomics
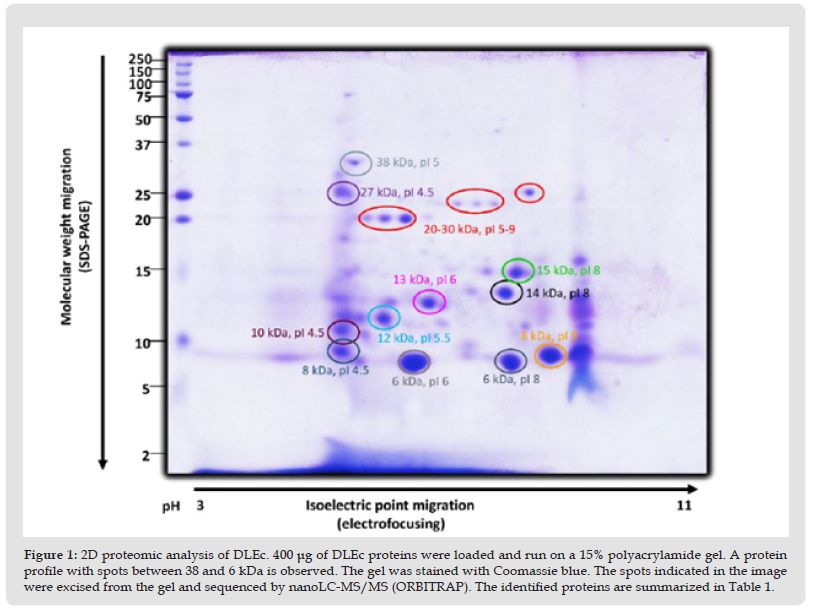
It has been postulated since its discovery that DLE, a leucocyte diafiltrate that passes through an exclusion pore of 5 KDa, is a mixture of small peptides bellow similar molecular weight. In this work, to define the identity of the constituent peptides of DLEc,2D electrophoresis analysis of the final product commercially called “Factran”, was conducted in the Ethan II isoelectrofocusing equipment. (Figure 1) shows a Coomassie/silver-stained gel representative of the peptide components of the mixture. About 80 peptides were identified, mostly between 6 and 15 KDa, although there is also a group of proteins between 20 and 38 KDa, with a pI range from 4 to 9. The most evident spots were cut out of the gel and sequenced by nanoLC MS/MS. The identity of the peptides is presented in (Table 1). Cytoskeleton proteins, ribonucleoproteins, constituent peptides of energy metabolism enzymes, heat shock proteins, and interestingly, several ubiquitin constituent peptides were identified.
Figure 2 2D proteomic analysis of DLEc. 400 μg of DLEc proteins were loaded and run on a 15% polyacrylamide gel. A protein profile with spots between 38 and 6 kDa is observed. The gel was stained with Coomassie blue. The spots indicated in the image were excised from the gel and sequenced by nanoLC-MS/MS (ORBITRAP). The identified proteins are summarized in Table 1.
DLEc Nanolc-MS/MS
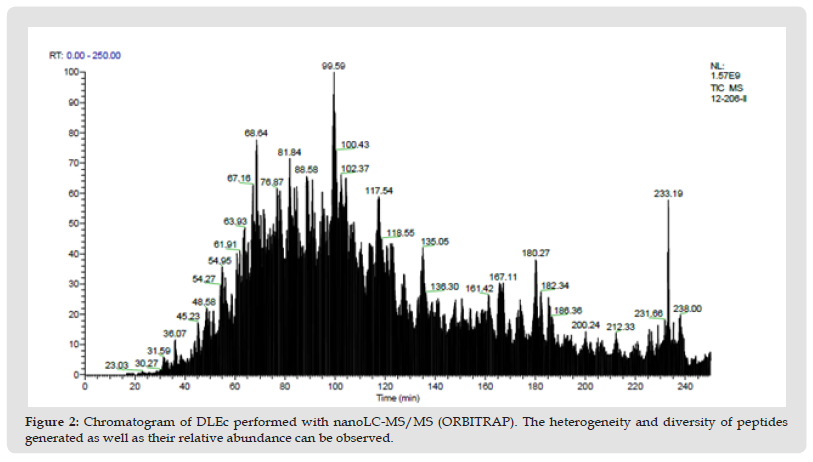
Taking into consideration the high variety of peptides whose abundance is low; in parallel we made the complete peptide sequencing of DLEc, by nanoLC-MS/MS (ORBITRAP), in collaboration with the proteomic services unit of the Complutense University of Madrid. Approximately 302 constituent peptides of the DLEc were found by ion trap sequencing. As can be seen in the chromatogram, there is a significant heterogeneity in the relative abundance of the peptides obtained (Figure 2); bioinformatic analysis of these peptides was performed in the non-redundant Swissprot (20233 sequences) database using a licensed version of MASCOT 2.3.0 and the Proteome Discoverer software version 1.2.0.208 (Thermo). The identity of each peptide is show in Supplementary Table 1.
Figure 3 Chromatogram of DLEc performed with nanoLC-MS/MS (ORBITRAP). The heterogeneity and diversity of peptides generated as well as their relative abundance can be observed.
DLEc Bioinformatics Analysis
Identification and Classification of the DLEc Proteins: The bioinformatic analysis was performed based on the results obtained in 2D and total proteomics performed by nanoLC-MS/MS of the DLEc. By means of proteomics analysis we were able to identify the different peptides belonging to the proteins that make up the DLEc. The identified proteins were used to obtain different interactomes to propose molecular pathways that could potentially modulate different cellular events deregulated in pathological states. The result obtained shows that DLEc peptides match proteins involved in crucial processes for cell survival such as signaling, gene expression regulation, secretion, heat shock or response to diverse types of stress and protein degradation (Figure 3). Proteins were distributed in the different processes as follows: gene expression (51%); regulatory proteins (11%), degradation (10%), secretion (10%), heat shock (7%) and only 4% involved in the immune system (Figure 4). Proteins involved in immune functions were ITIH4, ANXA5, SERPINB1, SERPINB3 and C3. The interactome analysis of these five proteins, suggests that ITIH4, SERPINB1 and C3 can potentially interact each other; SERPINB3 only interact with C3, while ANXA5 did not interact with the any of them. The proteins were independently challenged on the platform with the human proteome to assess whether these DLEc proteins can interact with immune modulatory proteins (Figure 5). Interactomes showed that peptides can interact with human proteins involved in the immune system response (59%); in cell cycle or cell proliferation (23%); in apoptosis (15%) and, to a lesser extent, with master regulatory proteins (3%) (Figure 6). ANXA5 is able to interact and modulate IL2, IL6, IL10, IL7, CD34, IGBP1, TNF, TNFSF10, CD19, and CSF2 proteins; SERPIND1 interacts with C4A, C3 and IL6, 8; C3 interacts with CD64, CR1, CFI, CD55, CR2,CFD, ITGAM, CFHR3, 4, 5, C5AR1, 5, IL6, VSIG4, MPO, ELANE and CD19; SERPINB3 modulates SERPINB4, ELANE, GRN, AZU1, PRTN3, MP0, C3, LYZ and GCA proteins; and ITIH4 interacts with CCR5, CD34, LGALS3BP, CTSW, CD209 and IL6. Based on these analyses, the results suggest that peptides contained in DLEc may exert a modulatory effect on the immune system and other essential biological processes in the cell.
Figure 4 General interactome of DLEc. The clusters correspond to proteins with similar functions, arrows mark immune system related proteins.
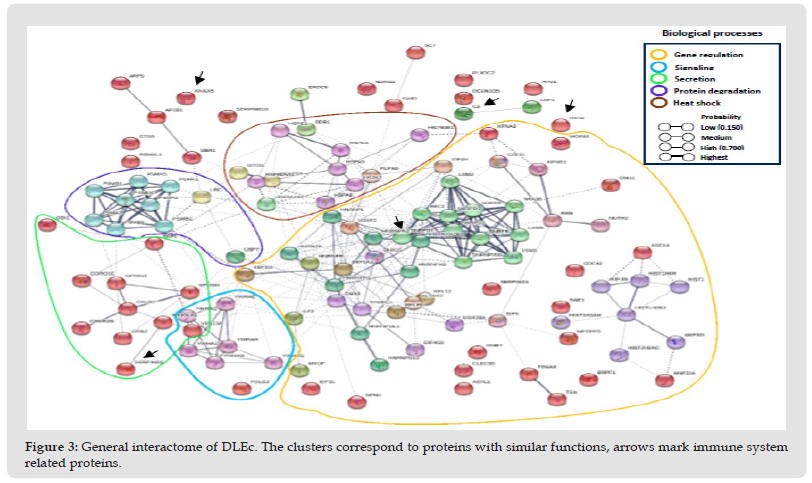
Figure 5 Distibution of DLEc identified peptides according to their function in different cellular processes.
Figure 6 Interactomes between DLEc and human proteins. A) Interaction of DLEc proteins that could modulate the immune response (ANXA5, SERPINB1, C3, SERPINB1 and ITIH4). Individual interactomes of each DLEc protein with the human proteome: B) ANXA5, C) SERPINB1, D) C3, E) SERPINB3 and F) ITIH4. Yellow circles and arrows indicate the DLEc protein.
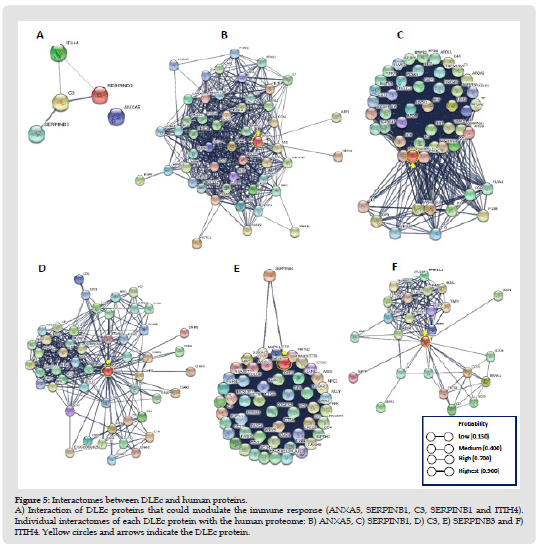
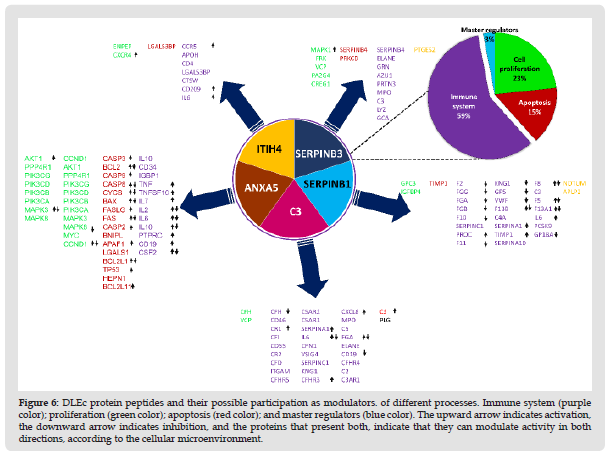
Figure 7 DLEc protein peptides and their possible participation as modulators. of different processes. Immune system (purple color); proliferation (green color); apoptosis (red color); and master regulators (blue color). The upward arrow indicates activation, the downward arrow indicates inhibition, and the proteins that present both, indicate that they can modulate activity in both directions, according to the cellular microenvironment.
Potential Health Impact of DLEc
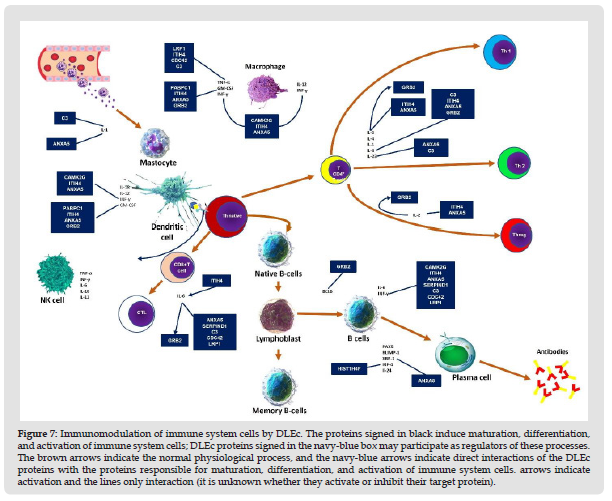
DLEc as a Potential Regulator of Immune System: Interactomes evaluation could predict biological interactions between DLEc peptides and proteins involved in immune system modulation (Szklarczyk, et al. [19]). We found eight candidate proteins: C3, ANXA5, CAMK2G, ITIH4, PABPC1, GRB2, LRP1, and SERPIND1 that most likely regulate immune system cell activity by being involved in the modulation of the interleukins IL-1 & α and β, IL- 2, IL-4, IL-6, IL-12 & α and β, IL-18, IL-23, and the INF-γ factors GM-CSF or CSF2 2, TNF, XBP-1 and IRF-4 in mast and dendritic cells (Nicholas y Lesinski, et al. [20]); T cells and their differentiation to CD4+ and CD8+ T, macrophage activation, as well as in maturation and differentiation of B cells to plasma cells (Figure 7) (Kim, et al. [21]). The results obtained strongly suggest the function of DLEc as a potent immunomodulator of the immune system under pathological conditions, such as viral, bacterial, antifungal, or parasitic infections, cancer and immunological disorders in which bioactive peptides from different species has been evaluated in several studies (Fernández-Ortega, et al. [1,22-29]).
DLEc As a Potential Regulator of Inflammation: We also evaluated the possible interactions of DLEc proteins on NF-κB-mediated regulation of inflammation. The results obtained show that annexin 5, ITIH4, C3, CDC42 and LRP1 proteins from DLEc interact with TNF-α factor, likewise, CALM3 and YWHAB, from DLEc showed interaction with IKB1, whereas DHX9, ILF2, YWHAE, ILF2, and RAN from DLEc bind directly with NF-κβ factor, as well as the activation of NF-κB via DHX9 (Figure 8). The results obtained suggest that DLEc could modulate the inflammation process at three important levels of the molecular pathway, since different DLEc proteins interact with IKB1 and TNF-α and NF-κβ In parallel, we searched the DLEc for bioactive peptides that have experimentally shown a biological effect in vitro or in vivo. We found peptides with immunomodulatory; anti-inflammatory; phagocytosis-inducing; antihypertensive; anticancer; antioxidant; modulators of various metabolic pathways; antimicrobial and antithrombotic activities (Table 2).
Figure 8 Immunomodulation of immune system cells by DLEc. The proteins signed in black induce maturation, differentiation, and activation of immune system cells; DLEc proteins signed in the navy-blue box may participate as regulators of these processes. The brown arrows indicate the normal physiological process, and the navy-blue arrows indicate direct interactions of the DLEc proteins with the proteins responsible for maturation, differentiation, and activation of immune system cells. arrows indicate activation and the lines only interaction (it is unknown whether they activate or inhibit their target protein).
The 2D proteomic characterization and the study of structure-function relationship of peptides contained in DLEc showed at least eighty major spots in molecular weight ranges between 38 and 6 KDa. The first interpretation of this result refutes at a stroke the long-standing assertion that transfer factor is a mixture of a small number of short amino acid sequences smaller than 5 KDa (Kirkpatrick, et al. [30]). Each spot corresponds to a protein or peptide with a characteristic molecular weight and isoelectric point, which indicated that the number of peptides far exceeded the hypothesis that remained valid for decades. For the sequencing analysis by mass spectrometry, we selected only the most abundant spots. Results showed the presence of structural proteins; enzymes ribonucleoproteins; constituent peptides of energy metabolism enzymes; heat shock proteins; interestingly, several ubiquitin constituent peptides, and various unidentified proteins. Recently, Vallejo-Castillo et al 2022, reported the proteome of DLE from human blood peripheral leucocytes, showing the presence of several peptides of at least from twenty-two human proteins, including actin, ankyrin-1, hemoglobin b-subunit, C3 complement protein, a-synuclein and as principal component, polyubiquitin-C that have immune regulatory effects. We found here that DLEc contains actins 1, 2, 4; C3 complement protein, and similar to previous work, ubiquitin protein peptides as well as some others related to the protein degradation process, such as polyubiquitin-C, ubiquitin carboxyl-terminal hydrolase 7, ubiquitin fusion degradation protein 1 homolog, ubiquitin-like modifier-activating enzyme 1, proteasome subunit alpha type-1, 2, 5, 7, proteasome subunit beta type-1, 2, 3, 26S proteasome non-ATPase regulatory subunit 2, and SUMO-activating enzyme subunit 1.
Taking into consideration that due the diversity, number, and abundance of peptides obtained by two-dimensional proteomic analysis exceeds eighty peptides, characterizing the DLEc proteome by this route will produce the loss of those peptides poorly represented, which could have relevant immune modulatory effects. Therefore, the analysis was performed using nanoLC-MS/MS orbitrap. After analyzed the entire proteome by bioinformatic analysis, about 302 peptides were identified, from those mostly represented and others with minimal abundance. Human DLE peptides are derived from 539 proteins grouped into eight clusters: I. Ankyrin-1, II. α-subunit of hemoglobin, III. Subunit β of hemoglobin, IV. Calpastatin, V. α-synuclein, VI. Cytoplasmic actin, VII. Polyubiquitin C and VIII. thymosin, as reported by Vallejo-Castillo et al. 2022. Although the polyubiquitin C cluster is one of the least abundant, they found by mass spectrometry that this protein contributes a greater number of peptides to the total DLE content. This work group highlighted the importance of this monomeric protein in a murine model (BAL/C mice) with Herpes Simplex Virus type 1 infection. Administration of DLE enriched with the monomeric ubiquitin increased the survival of mice compared to unenriched DLE, and even survival decreased further when ubiquitin is depleted from DLE. The authors continue that monomeric ubiquitin is the major bioactive component of the complex mixture of human DLE.
In our study we found that DLEc is an even more complex mixture than human DEL because it is derived from lymphoid tissue, which is rich in multiple cell subpopulations, compared to human DLE which is derived solely from blood leukocytes. Peptides correspond to a wide variety of proteins with structural, metabolic, and regulatory functions in various cellular processes. Interestingly, in DLEc we identified peptides possessed by human DLE, such as polyubiquitin C, complement protein C3 and cytoplasmic actin. Its knowledge lacks practical and medical importance if we are not able to define and associate the molecular pathways in which it participates, with special emphasis on those pathways that some groups in the world have experimentally demonstrated that different DLE are able to modulate physiological functions and favor the adjuvant or resolution of specific disorders. Immunomodulatory molecules can stimulate or suppress the immune response acting at various levels to selectively inhibit or activate populations or subpopulations of immune cells, such as lymphocytes, macrophages, neutrophils, NK killer cells and cytotoxic cells; or to modulate the production of soluble mediators such as cytokines. Among the immunomodulatory molecules are cytokines such as IL-1, INF g, IL-2, IL-5, IL-12, TNFa, IL-18 and GM-CSF, which can act directly or indirectly, being able to modulate the humoral and cellular immune response. Immunomodulators are frequently used as therapy for immunodeficiencies, cancer, hepatitis and in hematopoietic recovery (Nicholas, et al. [20]).
Interestingly, in DLEc we found peptides derived from five proteins that can potentially interact with human proteins involved in the immune system response. Annexin A5, SERPIND1, C3, SERPINB3 and ITIH4 can interact with different proteins related to the modulation of pro- and anti-inflammatory interleukins; activation of immune cells; chemotaxis, as well as activation of the complement system. We also identified that annexin 5, ITIH4, C3, CDC42, LRP1, CALM3, YWHAB, DHX9, ILF2, YWHAE, ILF2, and RAN can interact with TNFR, IKB1, NF-κB factor. Nuclear factor enhancer of activated B-cell kappa light chains (NF-κβ) is a protein complex that controls DNA transcription. This protein plays a key role in the regulation of the immune response elicited by different stimulus. Defective regulation of NF-kB is associated with cancer; inflammatory and autoimmune diseases; septic shock; viral infections or inappropriate immune development (Echeverri et al. [31]). NF-κβ can be activated by toxins, bacteria, viruses, parasites, or oxidative stress, however, the main molecular pathway for its activation is through the TNFR receptor, which can be activated by different cytokines or by tumor necrosis factor (TNF). In turn, when activated, it passes the signal downstream activating the adaptor proteins TRAF2-IKK, this allows inducing the phosphorylation of IKB1 (under normal conditions it is heterodimerized with NF-κβ) and, therefore, the release and activation of NF-κβ (Christian, et al. [32,33]). Once NF-κB is activated, it is translocated to the nucleus and transcribes various genes that protect and induce proliferation of cells that should otherwise die by apoptosis to prevent damage from spreading. In addition, it also regulates the transcription of genes involved in different cellular processes such as the maturation of different immune cells, the regulation of the cell cycle, the biosynthesis of inflammatory and proinflammatory cytokines, among others (Chen, et al. [34]). Our previous studies did in inflammatory diseases (Acosta et al. [16-18]) suggest that DLEc treatment prevent uncoupling of IκB from NFkB and, therefore, preventing it from translocating to the nucleus to carry out transcription of proinflammatory genes.
Using the proteome obtained with nanoLC-MS/MS, we perform interactomes to identify putative molecular pathways modulated by the DLEc peptides. The systematic study and comparison of the proteome in different metabolic or pathological situations allows us to identify those proteins whose presence, absence or alteration correlates with certain physiological stages (Aslam, et al. [33-59]). Taking up diverse research from different groups in the world and our own research carried out with DLEc, we focus the OMIC analysis on the capacity of DLEc to modulate different molecular pathways in the cell, with particular emphasis on those with experimental evidence and related to the immunomodulatory processes that are deregulated in the health-disease binomial. Alternatively, we searched for peptides with proven biological effect in vitro or in vivo and found that immune and ubiquitin related proteins are rich in bioactive peptides with immunomodulatory, anti-inflammatory, antihypertensive, anticancer, antioxidant, hypocholesterolemic, triglyceride synthesis inhibitor, hypoglycemic, antithrombotic, and antimicrobial functions (Table 2). Thus, in silico analyses together presented in this work strongly suggest the existence of several bioactive peptides with enormous potential as pharmacological adjuvant in different pathologies. Undoubtedly, once the proteome of DLE has been determined, future research should focus on determining its mechanisms of action, bioavailability, and pharmacological kinetics to be considered in a new, more natural, less toxic, self-sufficient, and human-friendly therapeutic that can help in the control of a wide variety of pathological conditions.
The authors declared that there is no conflict of interest.
To Instituto Politécnico Nacional and Farmainmune SA for grants and DLEc donation, respectively given to DGPI. The proteomic analysis was conducted in the Proteomics Facility UCM-PCM,’ a member of ProteoRed network.
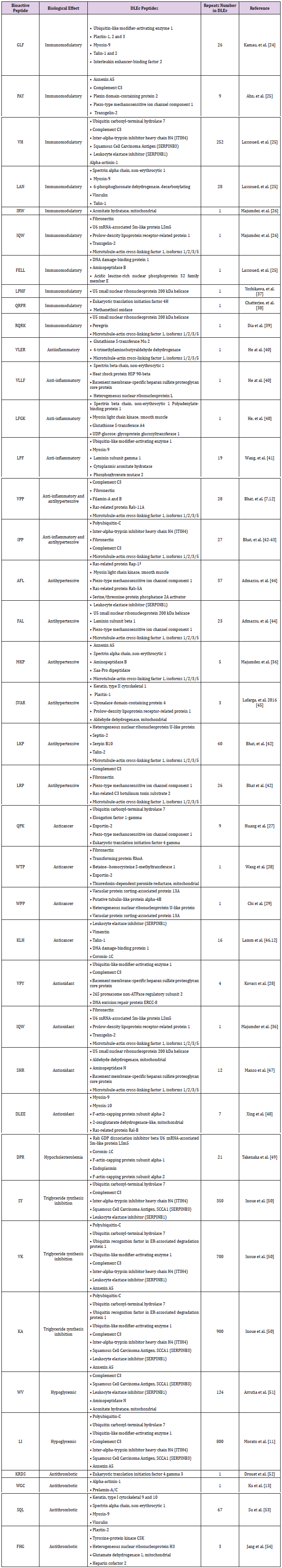
Table 2: Supplementary Table 1. PSMs, total number of identified peptide sequences; AAs, amino acids; MW (kDa), molecular weight in kilodaltons and pI, Isoelectric point.